Week 2 HW: DNA Read, Write & Edit
Part 1: Benchling & In-silico Gel Art
Lambda Sequence: Sequence from E.coli I cl857 S7 lambda bateriophage (Daniels, et al., 1983) available at New England Biolabs (N3011)
A digest simulation was performed using the lambda sequence and 7 different restriction enzyme (EcoRI, HindIII, BamHI, KpnI, EcoRV, SacI, and SalI). The range of fragments obtained from this simulation varies depending on the enzyme used.
EcoRV, for example, has 21 restriction sites, giving 22 band in the simulation. On the other hand, KpnI, SacI, and SalI have a few restriction sites, showing only two bands in the simulation (Figure 2).
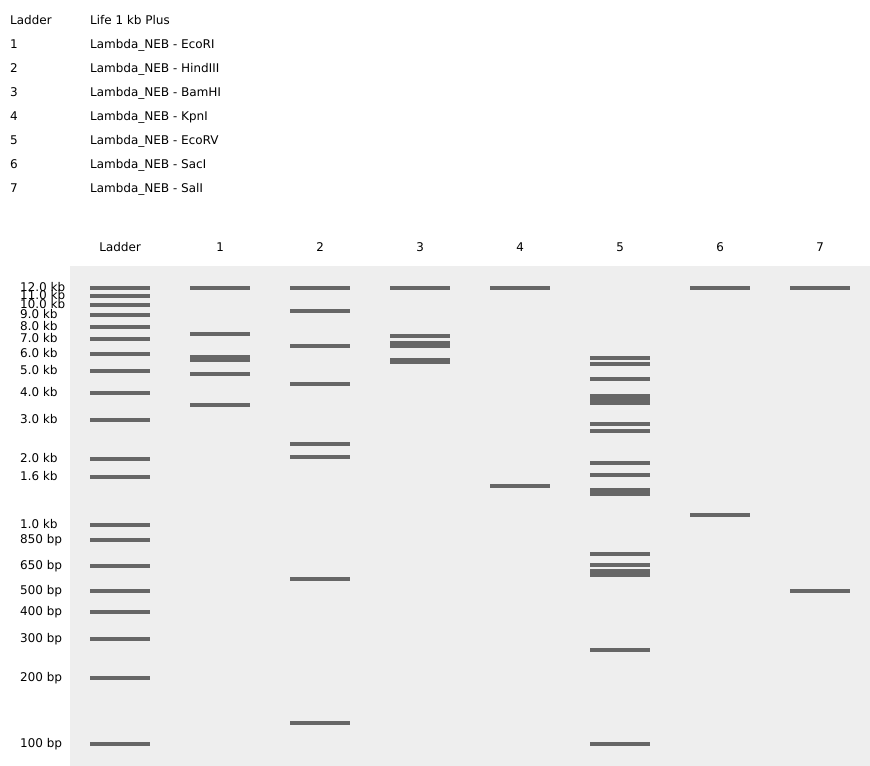


Since restriction enzymes cleave specific sequences in the genome, the difference between the number of sites for EcoRV compared to KpnI, SacI, and SalI raises the question: **Why does the lambda genome have more restriction sites for EcoRV than others?
Bacteriophages usually present fewer restriction sites as a response to this defense mechanism. This difference may change depending on the interaction between the bacteriophage and its host (Pleška & Guet, 2017)
Gel Art: Raimondi Stela Using a simulation of the digestion of lambda genome with different restriction enzymes, I tried to portray the “god of staffs”. This is a deity found in the Raimondi Stela that belonged to the Chavin culture (Figure 3)
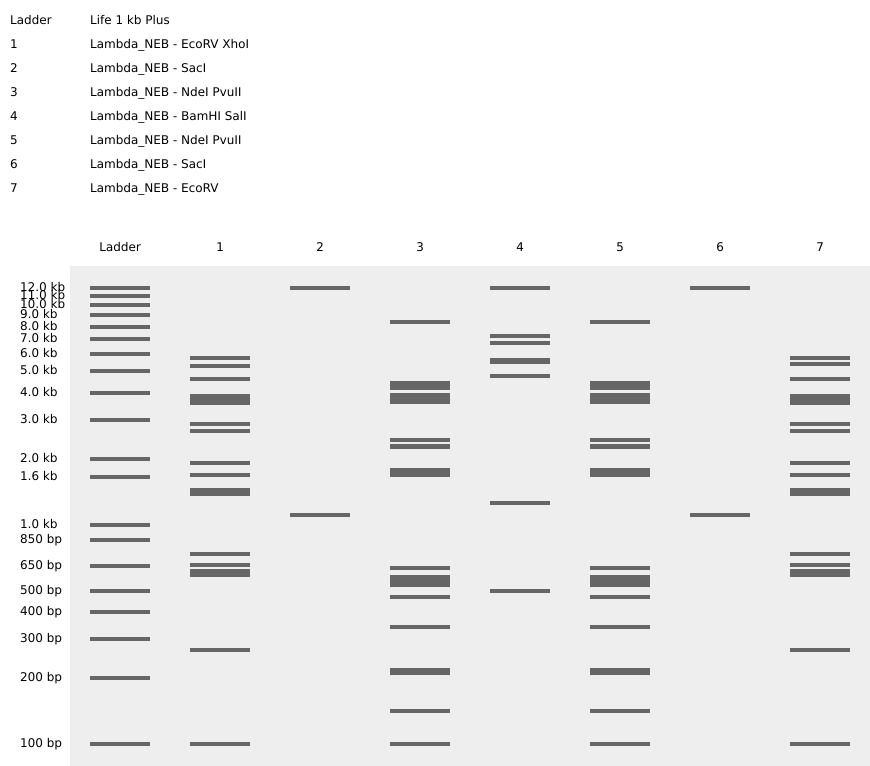

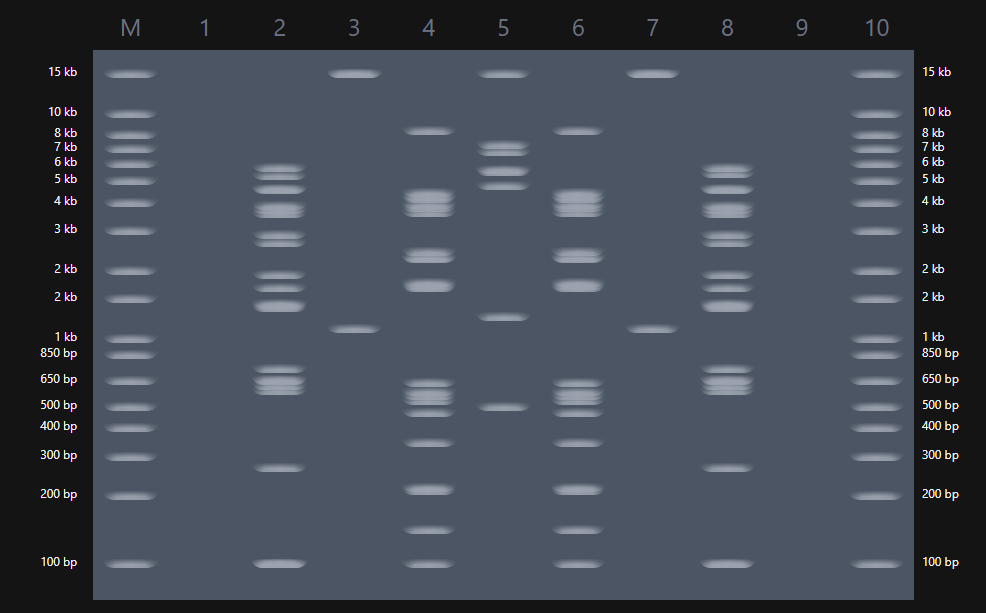

The gel created tried to be similar to the deity holding their staffs.
Part 3: DNA design challenge
Protein Chosen: Bothrops atrox snake venom nerve growth factor
Description: Nerve Growth Factor (NGF) is a member of the neurotrophin family that regulates the growth, differentiation, and survival of peripheral neurons during the development of the nervous system. This factor acts through two key receptors, tyrosine kinase A receptor (TrkA) and p75 neurotrophin receptor (p75NTR). The TrkA receptor activates signaling cascades that promote neuron differentiation and neurite growth.
Snake venom NGF (sNGF) is a protein that has been reported from the venom of elapid and viperid snakes. It is proposed that the presence of sNGF in the venom helps the envenomation process by causing the release of signaling chemicals that promote inflammatory reactions and increase vascular permeability, aiding the spread of other toxins and promote the apoptosis of cells (Sunagar et al., 2013).
Justification: NGFs have been proposed as promising options to treat neurodegenerative diseases and promote regenerative processes. sNGFs show high similarities to human NGFs and have been studied for many applications like chondrogenesis, neurite outgrowth, neuroprotection, tumor growth inhibition, etc. (Devi & Jayaraman, 2025).
Because sNGFs present special activities during envenomation, the study of sNGFs from other snake species may help to find new functions with a possible use in the study of regeneration and nervous development. These new functions may contribute to the design of synthetic alternatives with specific functions that can be applied for therapeutic purposes.
Protein Sequence: I have chosen the sequence of a sNGF from Bothrops atrox snake venom available at the UniProt database (ID: A0A1L8D608). The existence of this protein was proved through transcription level.
Selection of the expression system To continue with the process of reverse translation and codon optimization, I investigated which expression system would be the most suitable to produce this protein. Schütz et al. (2023) offers a concise guide for expression system selection with a decision graph depending on the characteristics of the protein to be expressed (Figure 3).
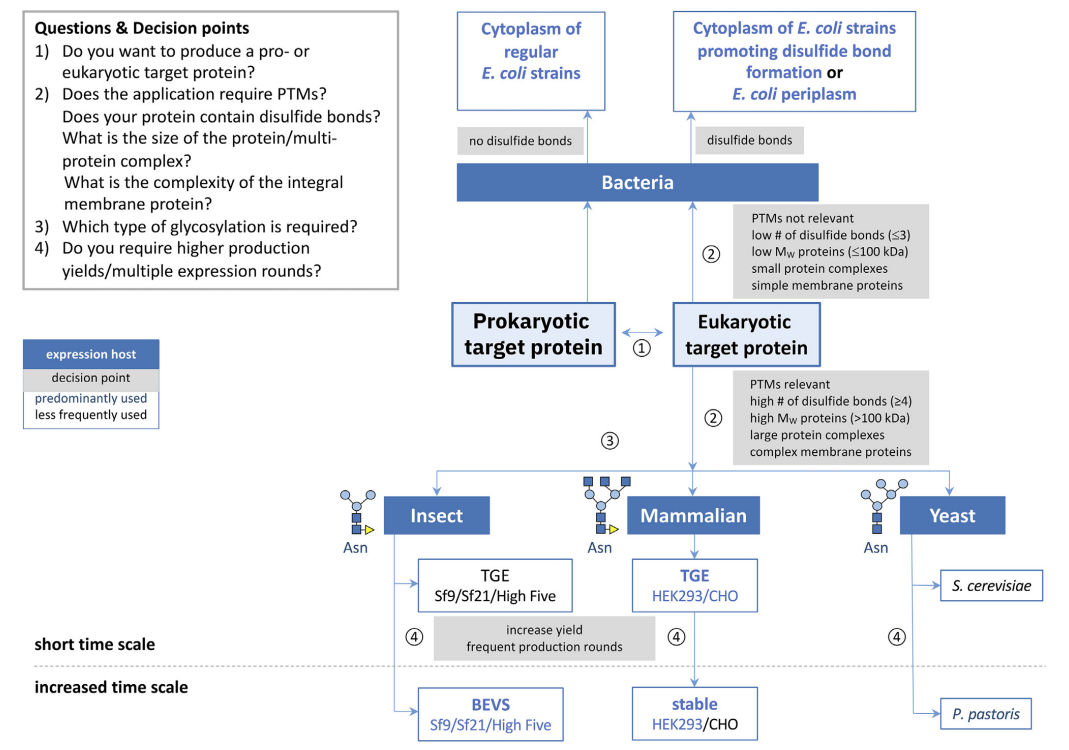
I gathered the following information of the protein based on four decision points proposed by Schütz:
- The target is eukaryotic protein
- Uniprot PTM/Processing section describes that the protein contains a signal region related to its secretion between the 1-18 amino acids and three disulfide bonds (Figure 4).
- The resulting protein would have 241 amino acids in total when expressed and 233 amino acids when secreted with a molecular mass of 27.197 KDa.
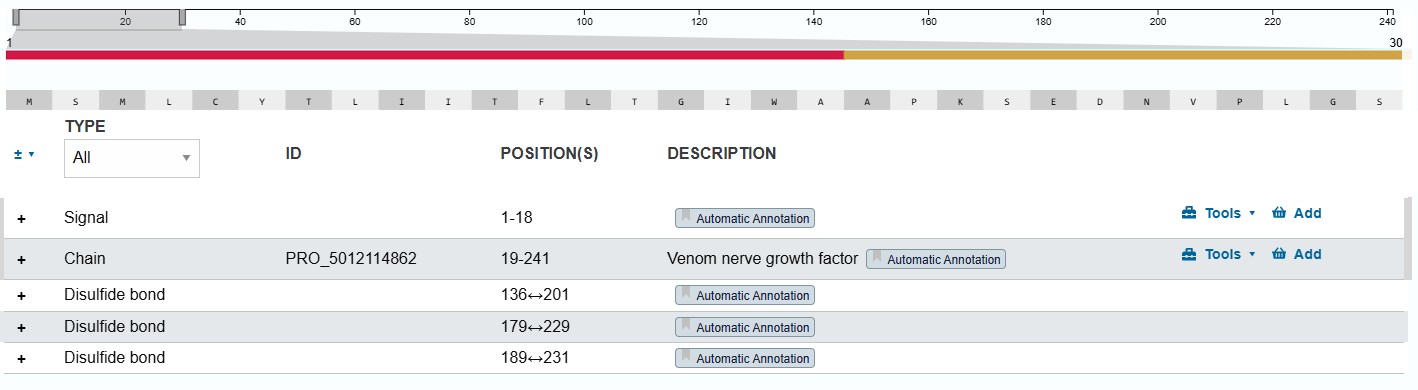

- Uniprot information from other NFGs does not show that requires glycosylation when expressed
- In the case of this design, it wouldn’t be necessary to have an expression system with higher yield
Based on this information the decision graph suggests using an expression system using a strain of E. coli that promotes disulfide bond formation. This decision is also supported by other studies that use E. coli to express human NFGs in vitro (Tilko et al., 2016; Dicou, 1992)
Reverse Translate Before performing the reverse translation of the protein, I decided to eliminate the amino acids 1-18 because they are part of the signal region of the protein and this won’t be used for this design. I included an initiator methionine at the N-terminus to allow translation initiation. The sequence modification was realized using the Benchling software.
Using the same software I reversed translated the protein using Escherichia coli (K12) genetic code using the method Match codon usage. The result of this process is an optimized sequence of 672 bp. This sequence was used later to perform a Blastx analysis where was found that the resulting sequence matches with other NGF from snakes (Figure 5)
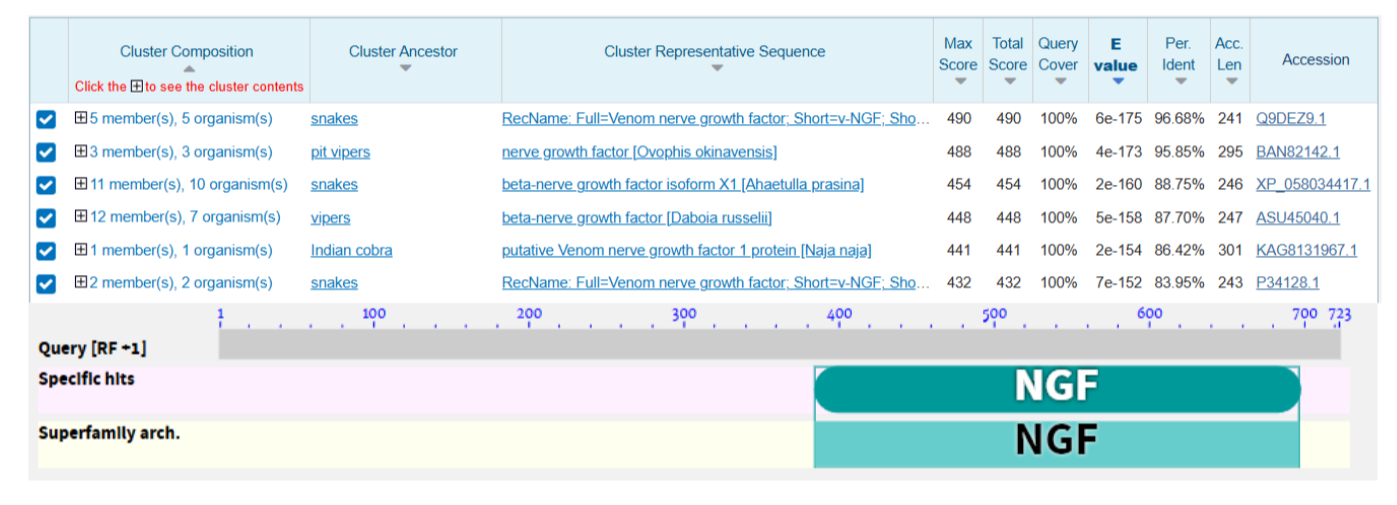

Codon optimization To simulate the creation of a clonal gene using the Twist Bioscience environment, the optimized sequence was uploaded in the software. A codon optimization was performed in the application. During the configuration of the optimization, I conserved the region 321-524 since it’s predicted as the NGF region by the Blastx result.
The resulting sequence was later labeled as optimized B.atrox NGF (BatroxNGFOptimized) and finally chosed as the sequence to be used for the creation of the expression cassette.
Expression Vector Selection To select a suitable expression vector, it is necessary to consider that the protein requires a proper environment to develop three disulfide bonds. The formation of disulfide bonds can be achieved by expressing the protein in E. coli periplasm or in the cytoplasm of engineered E coli.
A study performed by Shamriz et al. 2016 uses the pET-32a expression vector that contains the Trx-tag for increasing the solubility of the protein and its expression in E. coli Origami (DE3) to promote the correct formation of disulfide bonds in the cytoplasm of E. coli. Another strategy aims to translocate the recombinant protein into the periplasm using a signal peptide that helps the formation of disulfide bonds and increases its stability (Pouresmaeil & Azizi-Dargahlou, 2023).
Based on this information I opted for a pET-29b(+) expression vector from Twist Bioscience because it contains an N-terminal S•Tag™ sequence and may help with the protein solubility and a C-terminal His•Tag® sequence for its easy purification.
To help with the sulfide formation I selected SHuffle® strain from New England Biolabs that is engineered for the formation of disulfide bond in the cytoplasm.
Another way to express this protein is by adding signal sequence to allow the translocation of the protein to the periplasm and this could be analyzed later if possible.
Part 4: Preparation of Twist DNA Synthesis Order A simulation of DNA Synthesis order was generated using the optimized NGF sequence obtained from the previous part and inserted into the pET-29b (+) expression vector generating the plasmid as can be observed below (Figure 6)
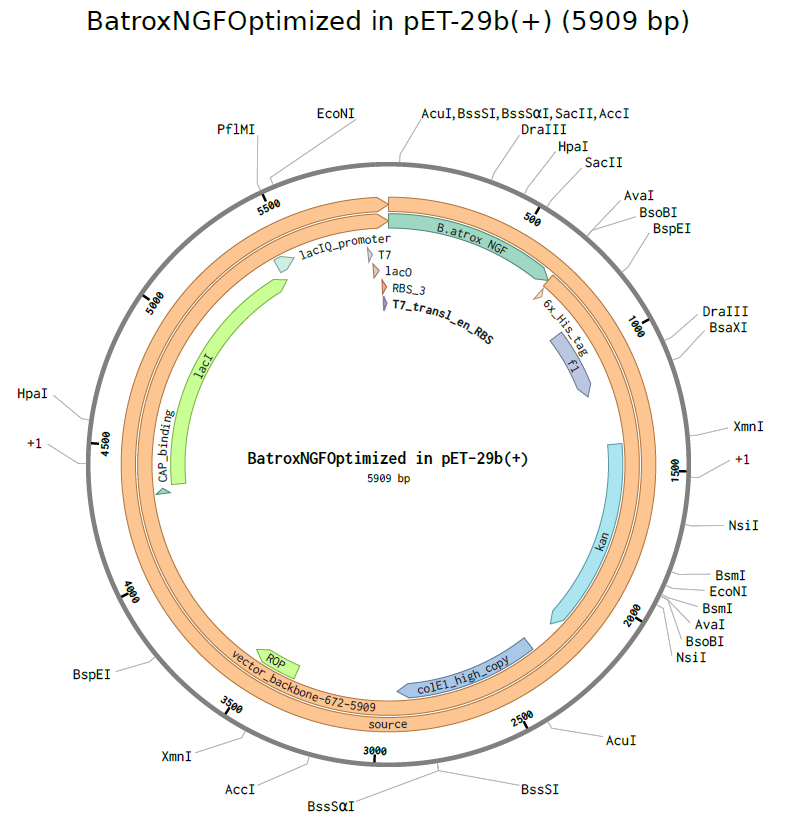
Part 5: DNA Read/Write/Edit
DNA Read Sequencing Idea: Genome-Wide Association Studies of Genetic Elements Related with Peanut Allergy Diversity in Peru Description: Allergies are misdirected immune reactions against a specific molecule (Allergens) to a previously exposed patient. These reactions are associated with an immune response mediated by a particular type of antibody called IgE. Allergies are diverse in nature and involve several ambiental and congenital factors, but also genetic factors. Several genes have been investigating for their involvement in allergic reactions, showing a complex heterogeneity that varies person to person (Falcon & Caoili, 2023) Genetic factors associated with allergies may help to elucidate the mechanisms that promote allergies predisposition. For that purpose, Genome-wide association studies (GWASs) offer a good option to study the genetic elements associated with allergies. GWAS are used to identify the association between genotypes with phenotypes. This is performed by selecting a group of individuals to obtain their phenotypic information. Using different GWAS arrays or sequencing strategies, genotypes of these individuals are obtained. Phenotypic and genotypic information is later used to conduct association tests to obtain relevant genetic elements that may be important for the phenotype studied (Uffelmann et al., 2021)
Technologies to perform GWAS genotyping
GWAS genotyping technologies are microarrays, Whole Exome Sequencing (WES) or Whole Genome Sequencing (WGS). To study the genetic component of peanut allergy in Perú we can use previously associated genes like HLA-DQ and HLA-DR o genes located in chromosome six (Allergies and Genetics | Health and Medicine | Research Starters | EBSCO Research, n.d.). The objective of genotyping these genes is to determine Single Nucleotide Polymorphisms (SNPs) that might have strong association with peanut allergy.
Whole genome sequencing technologies can be applied for SNP genotyping and involves sequencing all regions of the entire genome. On the other hand, Whole exome sequencing is a method for sequencing only the exonic region of the human genome.
- Is the method first-, second- or third- generation or other? Whole and Exome genome sequencing are part of the Next Generation Sequencing (NGS) because they are based in the massively parallel sequencing process.
- What is your input? How do you prepare your input? For this study my input is genomic DNA extracted from a representative Peruvian sample of individuals diagnosed with peanut allergy.
- What are the essential steps of your chosen technology, how does it decode the bases of your DNA sample? For WGS studies, Illumina uses a sequencing technology by synthesis, where fluorescently labeled nucleotides to sequence millions of clusters on a cell surface in parallel.
- What is the output of your sequencing technology? Illumina sequencing data is obtained through the signal intensity measurement of the labeled nucleotides that serve a terminators
DNA Write Project Idea: Snake venom NGF from B atrox The following idea aims to express a snake venom NGF from B. atrox. sNGFs have been applied in numerous studies to test their potential effect on regenerative processes because of their similarity with the human NGF and because of novel properties that may appear because of its evolution in the snake venom. For its production I propose the recombinant production of this protein, for that I realized used a sequence available at UniProt (ID: A0A1L8D608) a reverse translated to then propose it cloning using a vector in E. coli and expression in the same organism.
DNA Edit Project Idea: Using genetic engineered cells in hydrogels for cartilage regeneration Hydrogels are tridimentional networks polymers that can be used as scaffold for cartilage tissue engineering. A promising approach is to modify the genome of stem cells, creating specific gene circuits to promote cartilage regeneration. Trough gene edition, we could use steam cells to modify their proliferation capacity or control it using genetic circuits, a concept that may help with this idea is the concept of BioBricks that allows to create libraries that coul be used to modify the behavior of these stem cells (Elnaggar et al., 2025).
References
- Allergies and genetics | Health and Medicine | Research Starters | EBSCO Research. (n.d.). EBSCO. https://www.ebsco.com/research-starters/health-and-medicine/allergies-and-genetics
- Daniels, D.L. et al. (1983). Appendix II: Complete Annotated Lambda Sequence. R.W. Hendrix, J.W. Roberts, F.W. Stahl and R. A. Weisberg(Ed.), Lambda-II. 519-676. New York: Cold Spring Harbor Laboratory Press.
- Devi, S., & Jayaraman, G. (2025). Unraveling the molecular basis of snake venom nerve growth factor: human TrkA recognition through molecular dynamics simulation and comparison with human nerve growth factor. Frontiers in Bioinformatics, 5, 1674791. https://doi.org/10.3389/fbinf.2025.1674791
- Elnaggar, K. S., Gamal, O., Hesham, N., Ayman, S., Mohamed, N., Moataz, A., Elzayat, E. M., & Hassan, N. (2025). A guide in synthetic biology: Designing genetic circuits and their applications in stem cells. SynBio, 3(3), 11. https://doi.org/10.3390/synbio3030011
- Falcon, R. M. G., & Caoili, S. E. C. (2023). Immunologic, genetic, and ecological interplay of factors involved in allergic diseases. Frontiers in Allergy, 4, 1215616. https://doi.org/10.3389/falgy.2023.1215616
- Pleška, M., & Guet, C. C. (2017). Effects of mutations in phage restriction sites during escape from restriction–modification. Biology Letters, 13(12). https://doi.org/10.1098/rsbl.2017.0646
- Pouresmaeil, M., & Azizi-Dargahlou, S. (2023). Factors involved in heterologous expression of proteins in E. coli host. Archives of Microbiology, 205(5), 212. https://doi.org/10.1007/s00203-023-03541-9
- Shamriz, S., Ofoghi, H., & Amini-Bayat, Z. (2016). Soluble Expression of Recombinant Nerve Growth Factor in Cytoplasm of Escherichia coli. Iranian Journal of Biotechnology, 14(1), 16–22. https://doi.org/10.15171/ijb.1331
- Sunagar, K., Fry, B. G., Jackson, T. N. W., Casewell, N. R., Undheim, E. a. B., Vidal, N., Ali, S. A., King, G. F., Vasudevan, K., Vasconcelos, V., & Antunes, A. (2013). Molecular Evolution of Vertebrate Neurotrophins: Co-Option of the Highly Conserved Nerve Growth Factor Gene into the Advanced Snake Venom Arsenalf. PLoS ONE, 8(11), e81827. https://doi.org/10.1371/journal.pone.0081827
- Uffelmann, E., Huang, Q. Q., Munung, N. S., De Vries, J., Okada, Y., Martin, A. R., Martin, H. C., Lappalainen, T., & Posthuma, D. (2021). Genome-wide association studies. Nature Reviews Methods Primers, 1(1). https://doi.org/10.1038/s43586-021-00056-9