Week 10 HW: Imaging and Measurement
Homework: Final Project
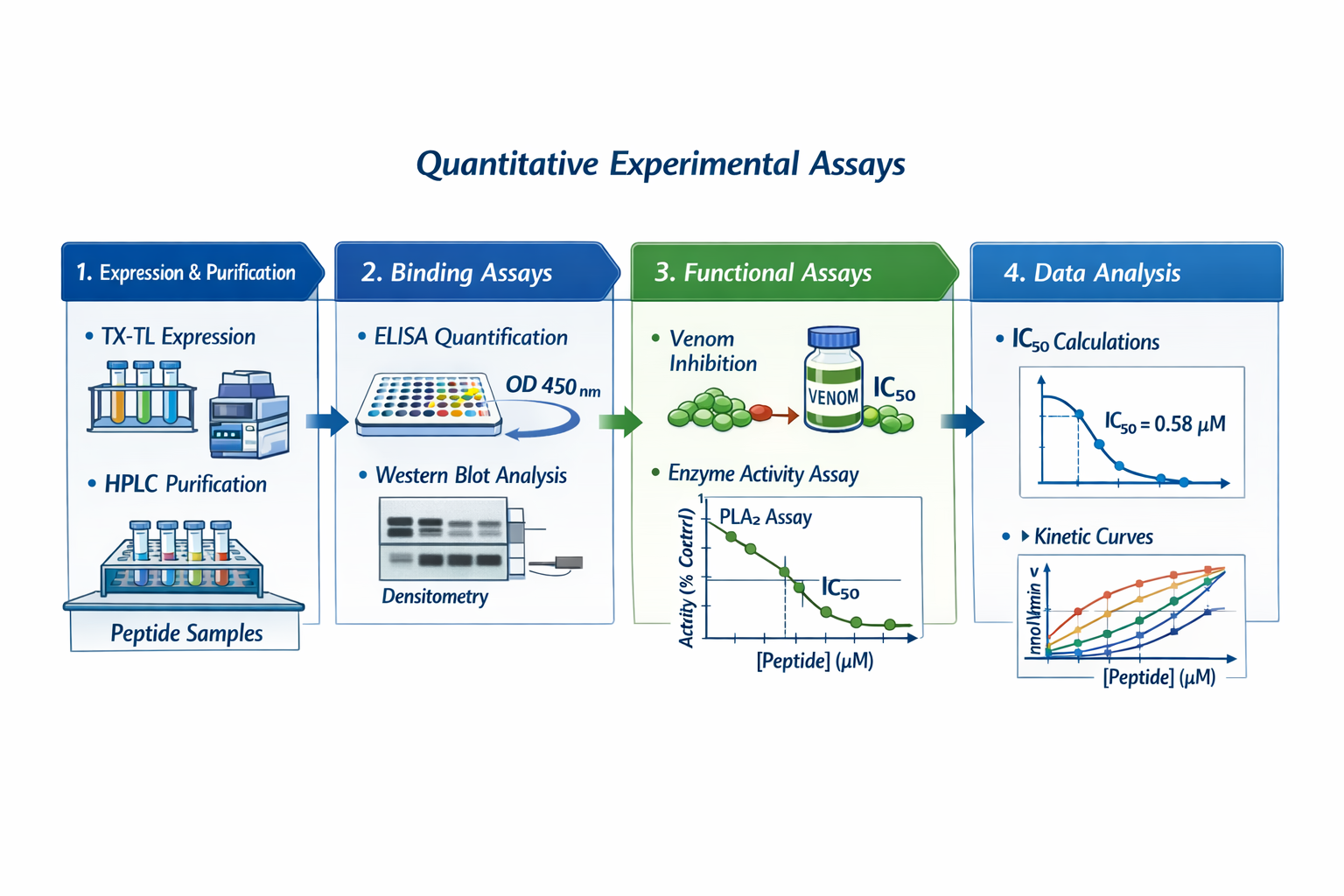
Figure 1 below presents some key aspects of my final project that require experimental testing and quantitative evaluation. These aspects refer to the expression of the protein binders generated using a Deep Learning Model and selected after in silico prediction of their therapeutic characteristics.
- Aspect 1: Cell-Free Expression System (CFS) Cell-Free Expression System (CFS) enables rapid expression of multiple protein variants in parallel. Since this project aims to predict and express different protein binders, CFS provides a scalable and automatable platform that avoids cell culture and allows preliminary functional testing without full purification (Cui et al., 2022)
To ensure reproducibility, the CFS workflow must be standarized and quantitatively monitored. Expression efficiency will be measures by using detection tags like biotinylated lysine or His-tag into the peptide of interest, enabling detection through SDS-PAGE followed by Western Blotting. A colorimetric readout using biotin-binding secondary antibody and chromogenic substrate will aloow quantification of expression levels (Hunt et al., 2024)
- Quantitative outputs for this aspect
- Relative Expression Intensity (Densitometry of Wester Blot bands)
- Expression yield per reaction
- Reproducibility across replicates (CV%)
Figure 2 illustrates the Promega FluoroTect and Trascend detection systems that portrays the methodology to evaluate the expression of protein from a CFS.
- Aspect 2: Peptide Purification To evaluate neutralization capacity, the expressed peptides must be purified from the CFS reaction. PURExpress In vitro Protein Synthesis Kit (New England Biolabs) for example uses reverse purification, a method that uses magnetic affinity resing based on metals that can selective capture His-tagged peptides (Figure 3)
This approach could enable rapid purification suitable for small peptides or mini-proteins and is compatible with downstream biochemical assays.
Quantitative outputs for this aspect
- Purification yield
- Purity (Assessed by SPS-PAGE)
- Recovery efficiency to crude expression levels The metrics allow can be use later to generated an automated system for protein binders expression and purification
Aspect 3: Reactivity and Neutralization The functional evaluation of the binders focuses on their ability to interact and neutralize the target toxin.
In vitro Neutralization: Neutralization can be test in vitro through the determination of the enzymatic activity of many protein targets (e.g. PLA2 or SVMP activity) in the presence of each binder. This provides a measureable neutralization parameter such as % inhibition or IC50.
Another strategy is the use of proteomics technique that involve the formation of the toxin-binder complex and and analyzed the interaction using a liquid chromatography-ms thechnique, which enables:
- Confirmation of the complex formation
- Identification of bound toxin fragments
- Comparizon of binding efficiency acroos binder variants
Proteomics thecnique are most suitable to analyze different binder-protein interaction in an automated process
- Quantitative outputs for this aspect
- % Reduction in toxin enzymatic activity
- IC50 values for each binder
- LC-MS peak intensities corresponding to toxin-binder complexes
References:
- Cui, Y., Chen, X., Wang, Z., & Lu, Y. (2022). Cell-Free PURE System: evolution and achievements. BioDesign Research, 2022, 9847014. https://doi.org/10.34133/2022/9847014
- Hunt, A. C., Rasor, B. J., Seki, K., Ekas, H. M., Warfel, K. F., Karim, A. S., & Jewett, M. C. (2024). Cell-Free gene Expression: methods and applications. Chemical Reviews, 125(1), 91–149. https://doi.org/10.1021/acs.chemrev.4c00116
Waters Part I: Molecular Weight
- Theoretical Molecular Weight and Isoelectric Poin The molecular weight (MW) of the eGFP protein was calcultad using the Expasy tool. Prior the calcultion, the His-tag and linker sequences were removed to obtain the MW corresponding to the native eGFP amino-acid sequence:
- Native eGFP MW: 26.941 kDa
- Isoelectric point: 5.58
The isoelectric point, being below than 7, indicated that the protein behaves as a weak acid, carrying a negative charge at a physiological pH
Expasy calculates MW using the average isotopic masses of amino acid and stimates the isoelectric point using the Bjellqvist pK set.
- Adjacent Charge State Apprach for Experimental MW Determination The molecular weight of eGFP was determined using LC-MS data provide in the homework figure. Two of the most intense adjacent charge-state peaks were selected for analysis (Figure 4)
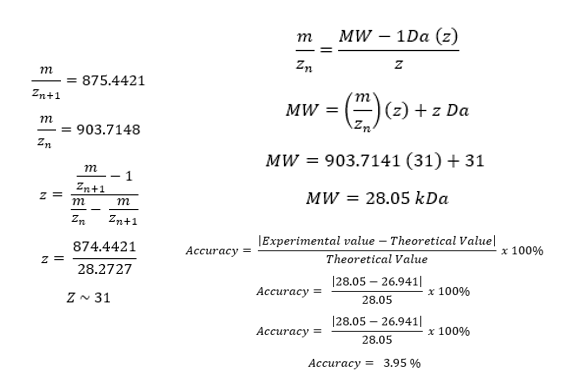

Using this approach a charge stat of 31 was stimated and the experimental MW was estimated near to 28.05 kDa
Comparison with the theoretical MW of native eGFP revealed a difference of 1.108 kDa. This deviation cannot be explained by common post‑translational modifications (PTMs), which typically contribute mass shifts in the range of +14 to +80 Da. Furthermore, eGFP expressed in E. coli does not undergo glycosylation, eliminating the possibility of large PTM‑related mass additions.
The calculated accuracy of the experimental result relative to the native theoretical MW was 3.95%, which is significantly higher than the accuracy typically reported for intact‑protein mass determination using this methodology (Durbing et al., 2025). This discrepancy strongly suggested that the analyzed protein did not correspond to the native eGFP sequence.
- Molecular Weight including His-Tag and Linker It was propose that the theoretical MW include the his-tag and linker sequences so a new theoretical MW was determine including this sequences:
- Molecular Weight (MW): 28.006 kDa
- Isoelectric point (pI): 5.90
When compared to the experimental MW (28.05 kDa), the difference was only 0.044 kDa, corresponding to an accuracy of 0.09%. This level of agreement is consistent with the expected performance of intact‑mass LC‑MS analysis and suggests that the protein analyzed retains its purification tag and linker.
Part 3: Waters Part 3
- The eGFP peptide was analyzed using the Benchling software that states the sequence contains 6 arginines and 20 lysines that compose 2.4% and 8.1% of the whole protein respectively (Figure 5)
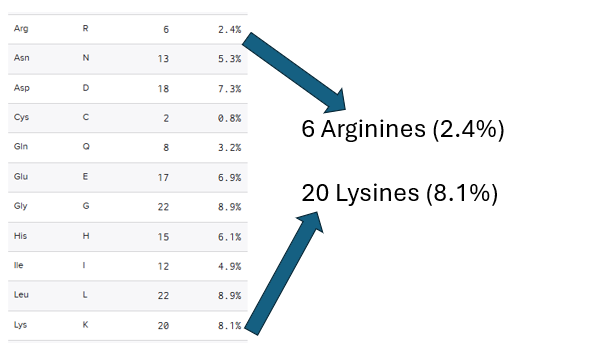

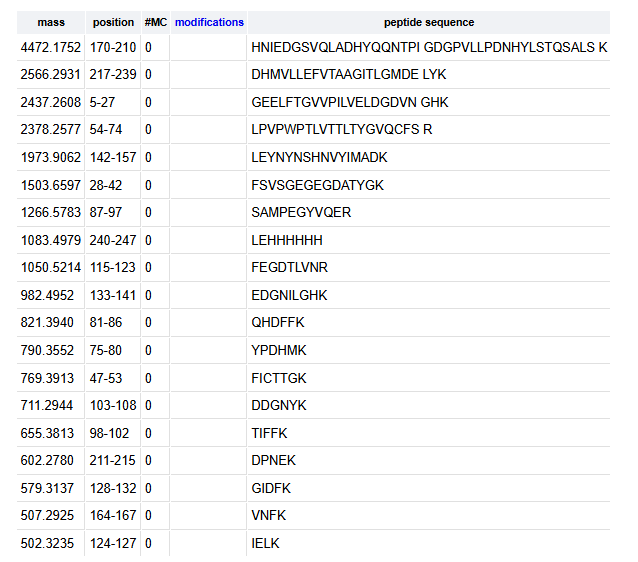
- Usin the Expasy Peptide mass tool it is predicted that the eGFP would have 19 fragments after the digestion with trypsin.