Week 11 HW: Bioproduction and Cloud Labs
Part A: Artwork Canvas
Contibution: I contribute with 52 dots making the “Love” Desing at the bottom left plate. I liked the collaborative working of the canvas and the constant organic evolution of the design that resembles very well with the development of synthetic biology. For a next collaborative art experiment I could propose an experiment that serves as a competence of different groups, or a collaborative project that shows thet location of the participant in the world.
Part B: Cell-Free Protein Synthesis
- Components of the cell-free reaction:
E. coli lysate
- BL21 (DE3) Star Lysate: Cell extract that contains all the components neccesary for transcription, translation, folding a nd energy regeneration of the cell-free system. E. coli BL21 strains is typically chosen for cell-free expresion because of its enhance mRNA stability and decrease protease activity (Hunt et al. 2024).
Salts/Buffer
- Potassium and Magnesium Glutamate: These salts are used to have different uses. Potassium and Magnesium ions are use to balance the charge of negatively charge biomolecules like DNA (Hunt et al., 2024). The ions are also involved in mantaining the integrity of stability of ribosomes, for example lack of magnesium is related with the loss of ribosome integrity and lack of potassium is related with the loss of ribosome activity (Nierhaus, 2014)
Glutamate acts a anionic counterion but also is used as secondary energy source. Optimizing glutamate amount is commonly done to regulate the amount of NAD and CoA cofactors because these tend to increase the cost of the expression system (Cai et al., 2015)
- HEPES-KOH and Potassium phosphate monobasic and dibasic: HEPES is traditionally used a buffer to control the pH level under acidic subproducts created from glucose metabolism (Hunt et al., 2024). Potassium phosphate can be also use as buffer and also to control ATP concentrations by supporting glycolisis and pyruvate metabolism.
Energy / Nucleotide system
Energy supplements are used for ATP regeneration through metabolic pathways. ATP is required as energy molecule for transcription and translation. Glucose is used a primary energy source for ATP generation through glycolisis but requires phosphate as source (Hunt et al., 2024)
NMPs (AMP, CMP, GMP, UMP): Are used a cost-effective supplement of nucleotides replacing NTPs. This monophosphates. Incorporation of NMPs is well stablished as the cytomin/glutamate-phosphate/NMP batch system because of its high yield and batch production (Jewett et al., 2008)
Translation Mix
17 amino acid mix: Aminoacids are neccesary for the translation process. Traditionally the 20 amino acid can be supplemented in excess to the reaction, however optimization studies have shown that several aminoacids like aspartate, lysine, and tyrosine can be produced in the cell-free reaction using the extract source materials (Jewett et al., 2008). Simmilarly other study has shown thet effect of halving every single aminoacid in the reaction showing that some aminoacids can be obtained through native metabolic process (Pedersen et al., 2010).
Cystein also plays a key role in protein synthesis, for example increasing the concentration of cystein can be related with and increase of glutathione S-transferase (Shingaki & Nimura, 2011), also cystein can cause inespecific disulfude bonds so its optimization is important.
Additives
Nicotinamide: Nicotinamide is a regulator of NAD metabolism and synthesis being an important regulate of the energy production.
Backfill
Nuclease-Free Water: Nuclease free water acts as the medium for all metabolic reactions and transcription-translation process. It is required to provide water free of nucleases and protease to avoid degradation of the desired products
- Differences between 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix
Figure below shows master mix compositions for the cell free system
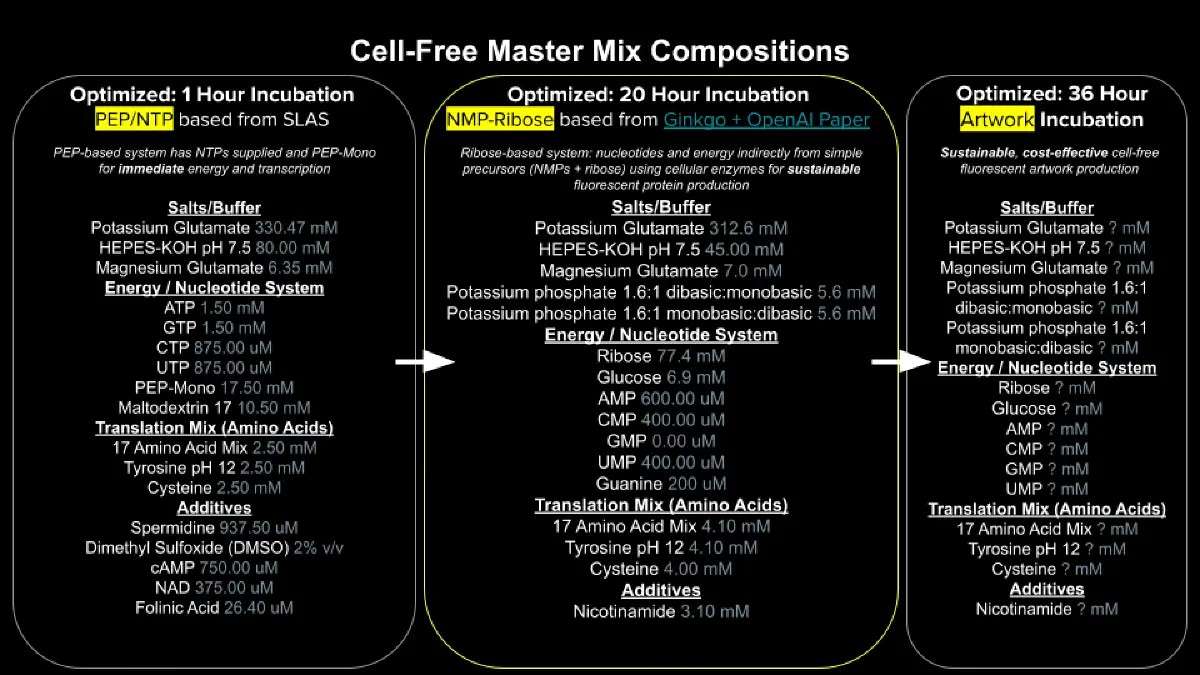

The principal difference that can be found is the source of energy and nucleotides used. For the 1 hour optimized system it can be observed that is base on Phosphoenol pyruvate (PEP), which is often used as energy soruce for the cell-free reaction. PEP is an expensive additive to the CFS reaction but initially showed better performance than glucose (Calhoun & Swartz, 2005). For the second master mix Ribose and Glucose are used a main source of energy, this could decrease the pH of the reaction and for that the addition of other buffers like Potassium phosphate are included.
Another main difference between both systems is the used of tri-phosphate nucleotides for the 1-hour master mix and monophosphate nucleotides for the 20 hours master mix, According to Jewett et al (2008), NMPs are cost-effective replacements of NTPs and can be used in batch systems with high yields.
Some additives are used in the 1 hour master mix to improve the reaction, for example Spermidine is usually used to increase transcription fidelity (Igarashi et al., 1982) and DMSO is commonly used to dissolve water-insoluble substances. In the case of the 20-hour master mix this additives can be avoided to decrease the cost of the reaction.
Part C: Planning the Global Experiment
Fluorescent Proteins
- sfGFP: Superfolded GFP is a variant of GFP from Aequorea victoria that is superfolded to avoid the effect of GFP folding in oxidizing environments (Aronson et al., 2011). One characteristic that can affect the role of GFP is the formation of inespecific agrupation because the protein tends to form weak dimers that can produce inclusion bodies.
- mRFP1: mRFP1 main disadvantages are the slow maturation and tendecy to form oligomers
- mKO2: mKO2 shows moderated acid sensitivity with a pKa of (5.5) indicating the necessity to control the pH of the medium
References
- Aronson, D. E., Costantini, L. M., & Snapp, E. L. (2011). Superfolder GFP is fluorescent in oxidizing environments when targeted via the SEC Translocon. Traffic, 12(5), 543–548. https://doi.org/10.1111/j.1600-0854.2011.01168.x
- Calhoun, K. A., & Swartz, J. R. (2005). Energizing cell‐free protein synthesis with glucose metabolism. Biotechnology and Bioengineering, 90(5), 606–613. https://doi.org/10.1002/bit.20449
- Cai, Q., Hanson, J. A., Steiner, A. R., Tran, C., Masikat, M. R., Chen, R., … & Yin, G. (2015). A simplified and robust protocol for immunoglobulin expression in E scherichia coli cell‐free protein synthesis systems. Biotechnology progress, 31(3), 823-831.
- Hunt, A. C., Rasor, B. J., Seki, K., Ekas, H. M., Warfel, K. F., Karim, A. S., & Jewett, M. C. (2024). Cell-Free gene Expression: methods and applications. Chemical Reviews, 125(1), 91–149. https://doi.org/10.1021/acs.chemrev.4c00116
- Igarashi, K., Hashimoto, S., Miyake, A., Kashiwagi, K., & Hirose, S. (1982). Increase of fidelity of polypeptide synthesis by spermidine in eukaryotic Cell‐Free systems. European Journal of Biochemistry, 128(2–3), 597–604. https://doi.org/10.1111/j.1432-1033.1982.tb07006.x
- Jewett, M. C., Calhoun, K. A., Voloshin, A., Wuu, J. J., & Swartz, J. R. (2008). An integrated cell‐free metabolic platform for protein production and synthetic biology. Molecular Systems Biology, 4(1), 220. https://doi.org/10.1038/msb.2008.57
- Nierhaus, K. H. (2014). MG 2+ , K + , and the ribosome. Journal of Bacteriology, 196(22), 3817–3819. https://doi.org/10.1128/jb.02297-14
- Pedersen, A., Hellberg, K., Enberg, J., & Karlsson, B. G. (2010). Rational improvement of cell-free protein synthesis. New Biotechnology, 28(3), 218–224. https://doi.org/10.1016/j.nbt.2010.06.015
- Shingaki, T., & Nimura, N. (2011). Improvement of translation efficiency in an Escherichia coli cell-free protein system using cysteine. Protein Expression and Purification, 77(2), 193–197. https://doi.org/10.1016/j.pep.2011.01.017