Week 2 HW: How is DNA Read Written and Edited
1. Benchling & In-silico Gel Art
Restriction enzymes used:
EcoRI HindIII BamHI KpnI EcoRV SacI SalI
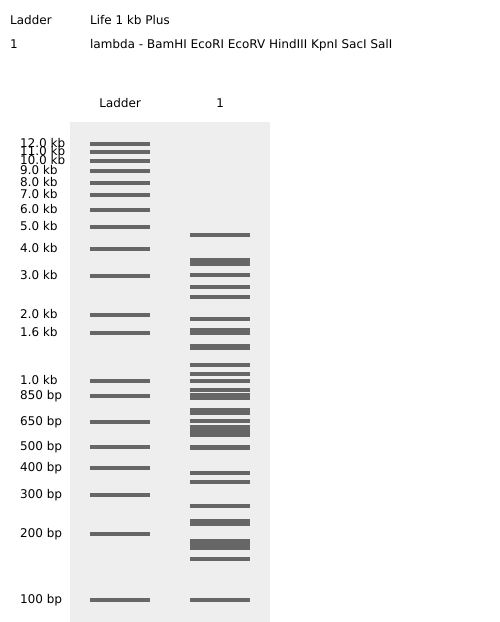

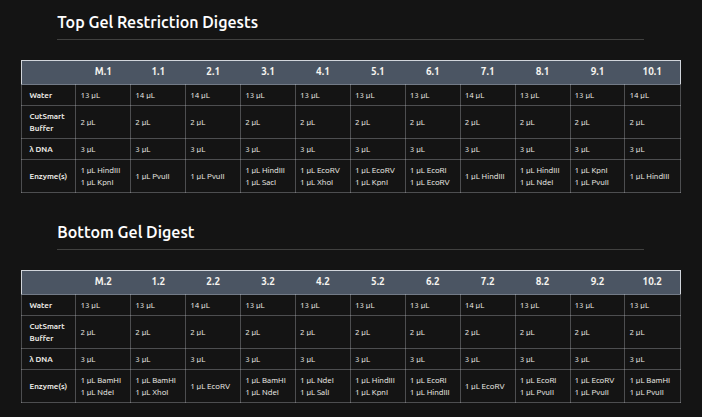
Digest In RCDONOVAN
An skull or spider
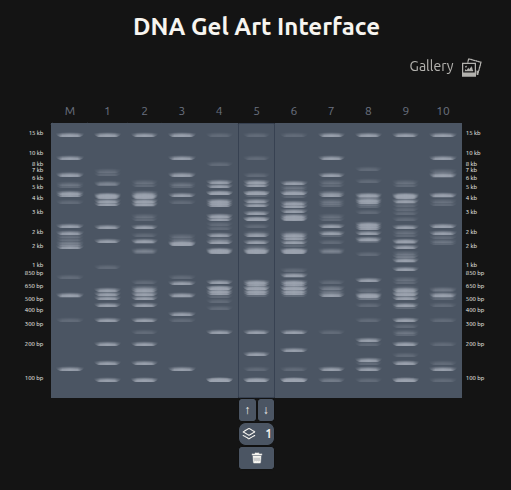



Part 3: DNA Design Challenge
Choose your protein
Hydrophobin HFBII (2B97) I believe is an interesting protein for , it allows for the gas exchange and water flow vital for the photobiont mantaining an inner atmosphere to weaker organisms inside. Aerial hyphae of the medula and surrounding photobiont have water repellent properties, sometimes covering both photobiont and mycobiont.
Sequence
sp|P79073|HYP2_HYPJE Class II hydrophobin 2 OS=Hypocrea jecorina OX=51453 GN=hfb2 PE=1 SV=1 MQFFAVALFATSALAAVCPTGLFSNPLCCATNVLDLIGVDCKTPTIAVDTGAIFQAHCAS KGSKPLCCVAPVADQALLCQKAIGTF
Reverse translation
atgcagttttttgcggtggcgctgtttgcgaccagcgcgctggcggcggtgtgcccgacc ggcctgtttagcaacccgctgtgctgcgcgaccaacgtgctggatctgattggcgtggat tgcaaaaccccgaccattgcggtggataccggcgcgatttttcaggcgcattgcgcgagc aaaggcagcaaaccgctgtgctgcgtggcgccggtggcggatcaggcgctgctgtgccag aaagcgattggcaccttt
Improved sequence
Looking after the organism we would use to produce a protein we improve a sequence depending on how available tRNA there is. In some organisms, many codons are rare and there would be many ribosome pausing causing misfolding.
In this case, as is a protein from lichen we are using Saccharomyces cerevisae as a guide to improve the DNA sequence. Also a high GC abundance will cause hairpin formations while a low amount will cause DNA innestability
Improved DNA[1]: GC=40.31%, CAI=0.92
ATGCAATTTTTTGCTGTTGCATTGTTCGCTACTTCAGCTTTGGCTGCTGTTTGTCCTACTGGTTTGTTCTCTAACCCATTATGTTGTGCTACAAATGTTTTGGATTTAATTGGTGTTGATTGTAAAACTCCAACAATTGCTGTTGACACAGGTGCAATCTTTCAAGCTCACTGTGCATCTAAAGGTTCTAAGCCATTGTGTTGTGTCGCTCCAGTTGCTGATCAAGCTTTGTTGTGTCAAAAGGCCATTGGTACCTTT
3.4 I have a Sequence! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
Saccharomyces cerevisiae is the closest organism to a mycobiont from the list of most studied models. First we need to introduce the gene into a yeast plasmid. TO build the plasmid we need a yeast promoter GAL1 a CYC1 Terminator- We need to transform yeast introducing the plasmid via electroporation. This protein is especially easy to purifiy due to its physicochemical characteristics.
How does it work in nature/biological systems?
- Describe how a single gene codes for multiple proteins at the transcriptional level.
At transcriptional levels the initial RNA Must be processed before it becomes a mature messenger RNA (mRNA) , so , primary mRNA becomes a mature mRNA via splicing removing intervening sequences, the introns. Having the exons , diferent exons can lead to diferent proteins from a single gene. Causing the generation of different POst transcriptional procesing cuuld also alter the proteins modifying them into smaller fragments.
- Try aligning the DNA sequence, the transcribed RNA, and also the resulting translated Protein!!! See example below.
DNA ATGCAATTTTTTGCTGTTGCATTGTTCGCTACTTCAGCTTTGGCTGCTGTTTGTCCTACTGGTTTGTTCTCTAACCCATTATGTTGTGCTACAAATGTTTTGGATTTAATTGGTGTTGATTGTAAAACTCCAACAATTGCTGTTGACACAGGTGCAATCTTTCAAGCTCACTGTGCATCTAAAGGTTCTAAGCCATTGTGTTGTGTCGCTCCAGTTGCTGATCAAGCTTTGTTGTGTCAAAAGGCCATTGGTACCTTT
RNA AUGCAAUUUUUUGCUGUUGCAUUGUUCGCUACUUCAGCUUUGGCUGCUGUUUGUCCUACUGGUUUGUUCUCUAACCCAUUAUGUUGUGCUACAAAUGUUUUGGAUUUAAUUGGUGUUGAUUGUAAAACUCCAACAAUUGCUGUUGACACAGGUGCAAUCUUUCAAGCUCACUGUGCAUCUAAAGGUUCUAAGCCAUUGUGUUGUGUCGCUCCAGUUGCUGAUCAAGCUUUGUUGUGUCAAAAGGCCAUUGGUACCUUU
Aminoacid translation MQFFAVALFATSALAAVCPTGLFSNPLCCATNVLDLIGVDCKTPTIAVDTGAIFQAHCASKGSKPLCCVAPVADQALLCQKAIGTF
Part 4: Prepare a Twist DNA Synthesis Order
https://benchling.com/s/seq-bsFO3WBzxpE4UVmNA9t0?m=slm-eJZA5VO4mRD3VjZ7jpvL INtroucing a hydrophobin into a bacterial sequence
PLasmid Designed
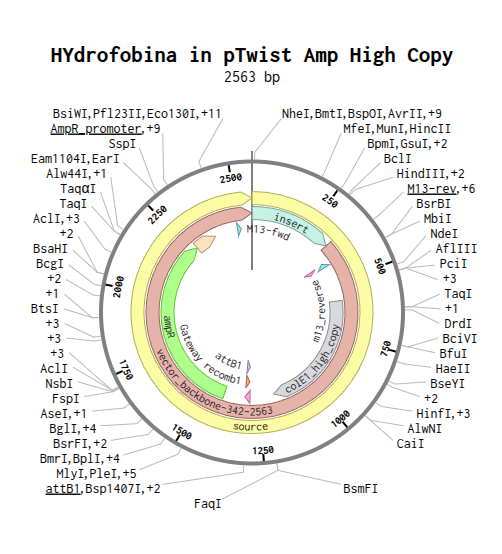

DNA READ
- What DNA would i Sequence?
For this specific project the first recoemndation is isolate and sequence components from lichen, the extraction methods many papers have used requiere a combined extraction and lysis method to destroy the hard protective layer from the mycobiont to the photobiont. It would be requiered if there is resources available to sequence de mycobiont at least.
From the engineering standpount it would enable to validate the strand integrity and benchmark how good was the extraction method for all beings inside the symbiosis .
- Sequence technology choice
Primary platform would be illumina sequencing by synthesis , a second generation technique, while not being the best for long reads, it would ensure quality due to a difficult management of DNA extraction. Lichen requieres higher accuraccy and deeper coverage of each individual sequence.
ONT also would be requiered for structural variation and unknown sequence of many different organisms in the consortia. Both are requiered for this specific project.
The input for Illumina would be a purified double stranted DNA , preparing the libraries using mechanical fragmentation, end repair, amplification of ITS region for the mycobiont , the output we would have would be FASTQ files, paired end.
- DNA write
I would sytnhesize a biosensor circuit from lichen. As they can synthesize many resistant factors and metabolites in rapidly in response to speicif stresses, lichen can survive extreme UV, desiccation, exposure to reactive oxygen species.
For environmentally dedicated bacteria we could build a sensor that may detect different type of stress and activate an specific pathway or conformation ensuring the protection of a whole system. It also may affect different systems at once.
THe synthesis technology would be phosphoramidite solid phase DNA sysnthesis, is not so hard or complex to do, the common method is enough.
DNA edit
- What DNA would I edit and Why.
I would engineer DNA repair pathways to study genotype and phenotype relationships, as lichens are exposed to many mutagenic agents it could have major applications to build climate resistent crops or even reduce the requierement for kill-switches constantly.
Reducing the mutation rate could open to more synthetic ecological applications due to lower risk in the introduction of new systems.
- The primary editing technology would be CRUSPR Cas9 .
sgRNA hybridizes and targets the DNA, Cas9 recogizes PAM, double strand break induced and the repair pathway would determine small insertion and deletions, small changes to learn how the full systems work. THe input requiered would be the Cas9 expression vector, the sgRNA cassette, the donor DNA template, selectable marker and host cell.