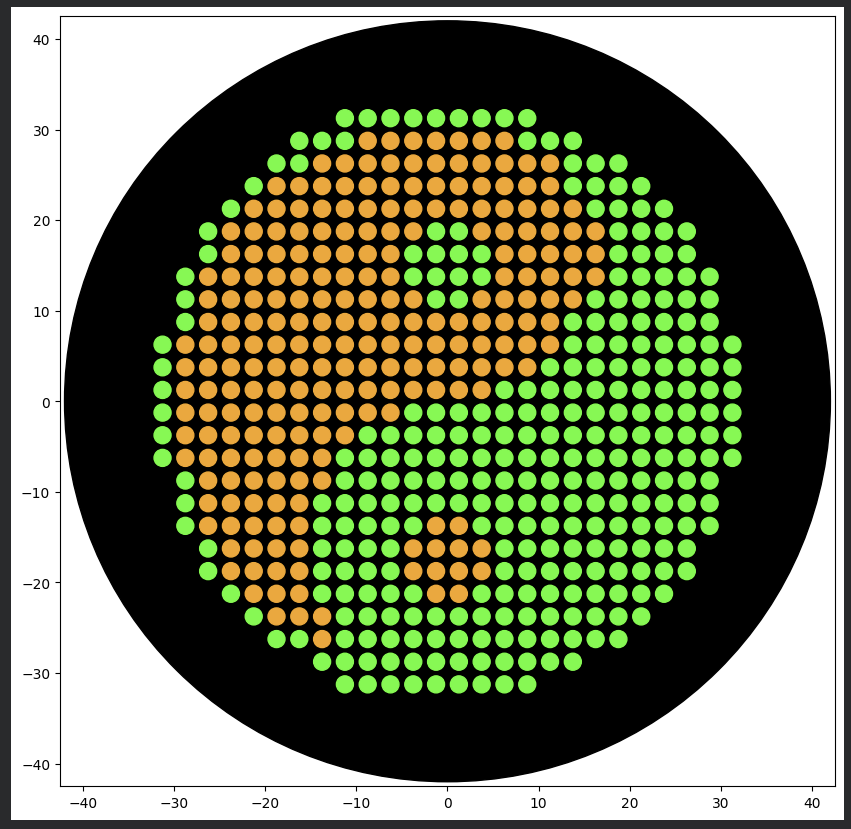
Each coordinate corresponds to a 1.5 µL droplet deposited at a specific XY location relative to the plate center.
Below is the complete script used for the Opentrons simulation and execution.
from opentrons import types
import string
metadata = {
'protocolName': 'Juan Francisco Larrea - Opentrons Art - HTGAA',
'author': 'HTGAA',
'source': 'HTGAA 2026',
'apiLevel': '2.20'
}
Z_VALUE_AGAR = 2.0
POINT_SIZE = 1.5
mclover3_points = [(-11.25,31.25), (-8.75,31.25), (-6.25,31.25), (-3.75,31.25), (-1.25,31.25), (1.25,31.25), (3.75,31.25), (6.25,31.25), (8.75,31.25), (-16.25,28.75), (-13.75,28.75), (-11.25,28.75), (8.75,28.75), (11.25,28.75), (13.75,28.75), (-18.75,26.25), (-16.25,26.25), (13.75,26.25), (16.25,26.25), (18.75,26.25), (-21.25,23.75), (13.75,23.75), (16.25,23.75), (18.75,23.75), (21.25,23.75), (-23.75,21.25), (16.25,21.25), (18.75,21.25), (21.25,21.25), (23.75,21.25), (-26.25,18.75), (-1.25,18.75), (1.25,18.75), (18.75,18.75), (21.25,18.75), (23.75,18.75), (26.25,18.75), (-26.25,16.25), (-3.75,16.25), (-1.25,16.25), (1.25,16.25), (3.75,16.25), (18.75,16.25), (21.25,16.25), (23.75,16.25), (26.25,16.25), (-28.75,13.75), (-3.75,13.75), (-1.25,13.75), (1.25,13.75), (3.75,13.75), (18.75,13.75), (21.25,13.75), (23.75,13.75), (26.25,13.75), (28.75,13.75), (-28.75,11.25), (-1.25,11.25), (1.25,11.25), (16.25,11.25), (18.75,11.25), (21.25,11.25), (23.75,11.25), (26.25,11.25), (28.75,11.25), (-28.75,8.75), (13.75,8.75), (16.25,8.75), (18.75,8.75), (21.25,8.75), (23.75,8.75), (26.25,8.75), (28.75,8.75), (-31.25,6.25), (13.75,6.25), (16.25,6.25), (18.75,6.25), (21.25,6.25), (23.75,6.25), (26.25,6.25), (28.75,6.25), (31.25,6.25), (-31.25,3.75), (11.25,3.75), (13.75,3.75), (16.25,3.75), (18.75,3.75), (21.25,3.75), (23.75,3.75), (26.25,3.75), (28.75,3.75), (31.25,3.75), (-31.25,1.25), (6.25,1.25), (8.75,1.25), (11.25,1.25), (13.75,1.25), (16.25,1.25), (18.75,1.25), (21.25,1.25), (23.75,1.25), (26.25,1.25), (28.75,1.25), (31.25,1.25), (-31.25,-1.25), (-3.75,-1.25), (-1.25,-1.25), (1.25,-1.25), (3.75,-1.25), (6.25,-1.25), (8.75,-1.25), (11.25,-1.25), (13.75,-1.25), (16.25,-1.25), (18.75,-1.25), (21.25,-1.25), (23.75,-1.25), (26.25,-1.25), (28.75,-1.25), (31.25,-1.25), (-31.25,-3.75), (-8.75,-3.75), (-6.25,-3.75), (-3.75,-3.75), (-1.25,-3.75), (1.25,-3.75), (3.75,-3.75), (6.25,-3.75), (8.75,-3.75), (11.25,-3.75), (13.75,-3.75), (16.25,-3.75), (18.75,-3.75), (21.25,-3.75), (23.75,-3.75), (26.25,-3.75), (28.75,-3.75), (31.25,-3.75), (-31.25,-6.25), (-11.25,-6.25), (-8.75,-6.25), (-6.25,-6.25), (-3.75,-6.25), (-1.25,-6.25), (1.25,-6.25), (3.75,-6.25), (6.25,-6.25), (8.75,-6.25), (11.25,-6.25), (13.75,-6.25), (16.25,-6.25), (18.75,-6.25), (21.25,-6.25), (23.75,-6.25), (26.25,-6.25), (28.75,-6.25), (31.25,-6.25), (-28.75,-8.75), (-11.25,-8.75), (-8.75,-8.75), (-6.25,-8.75), (-3.75,-8.75), (-1.25,-8.75), (1.25,-8.75), (3.75,-8.75), (6.25,-8.75), (8.75,-8.75), (11.25,-8.75), (13.75,-8.75), (16.25,-8.75), (18.75,-8.75), (21.25,-8.75), (23.75,-8.75), (26.25,-8.75), (28.75,-8.75), (-28.75,-11.25), (-13.75,-11.25), (-11.25,-11.25), (-8.75,-11.25), (-6.25,-11.25), (-3.75,-11.25), (-1.25,-11.25), (1.25,-11.25), (3.75,-11.25), (6.25,-11.25), (8.75,-11.25), (11.25,-11.25), (13.75,-11.25), (16.25,-11.25), (18.75,-11.25), (21.25,-11.25), (23.75,-11.25), (26.25,-11.25), (28.75,-11.25), (-28.75,-13.75), (-13.75,-13.75), (-11.25,-13.75), (-8.75,-13.75), (-6.25,-13.75), (-3.75,-13.75), (3.75,-13.75), (6.25,-13.75), (8.75,-13.75), (11.25,-13.75), (13.75,-13.75), (16.25,-13.75), (18.75,-13.75), (21.25,-13.75), (23.75,-13.75), (26.25,-13.75), (28.75,-13.75), (-26.25,-16.25), (-13.75,-16.25), (-11.25,-16.25), (-8.75,-16.25), (-6.25,-16.25), (6.25,-16.25), (8.75,-16.25), (11.25,-16.25), (13.75,-16.25), (16.25,-16.25), (18.75,-16.25), (21.25,-16.25), (23.75,-16.25), (26.25,-16.25), (-26.25,-18.75), (-13.75,-18.75), (-11.25,-18.75), (-8.75,-18.75), (-6.25,-18.75), (6.25,-18.75), (8.75,-18.75), (11.25,-18.75), (13.75,-18.75), (16.25,-18.75), (18.75,-18.75), (21.25,-18.75), (23.75,-18.75), (26.25,-18.75), (-23.75,-21.25), (-13.75,-21.25), (-11.25,-21.25), (-8.75,-21.25), (-6.25,-21.25), (-3.75,-21.25), (3.75,-21.25), (6.25,-21.25), (8.75,-21.25), (11.25,-21.25), (13.75,-21.25), (16.25,-21.25), (18.75,-21.25), (21.25,-21.25), (23.75,-21.25), (-21.25,-23.75), (-11.25,-23.75), (-8.75,-23.75), (-6.25,-23.75), (-3.75,-23.75), (-1.25,-23.75), (1.25,-23.75), (3.75,-23.75), (6.25,-23.75), (8.75,-23.75), (11.25,-23.75), (13.75,-23.75), (16.25,-23.75), (18.75,-23.75), (21.25,-23.75), (-18.75,-26.25), (-16.25,-26.25), (-11.25,-26.25), (-8.75,-26.25), (-6.25,-26.25), (-3.75,-26.25), (-1.25,-26.25), (1.25,-26.25), (3.75,-26.25), (6.25,-26.25), (8.75,-26.25), (11.25,-26.25), (13.75,-26.25), (16.25,-26.25), (18.75,-26.25), (-13.75,-28.75), (-11.25,-28.75), (-8.75,-28.75), (-6.25,-28.75), (-3.75,-28.75), (-1.25,-28.75), (1.25,-28.75), (3.75,-28.75), (6.25,-28.75), (8.75,-28.75), (11.25,-28.75), (13.75,-28.75), (-11.25,-31.25), (-8.75,-31.25), (-6.25,-31.25), (-3.75,-31.25), (-1.25,-31.25), (1.25,-31.25), (3.75,-31.25), (6.25,-31.25), (8.75,-31.25)]
azurite_points = [(-8.75,28.75), (-6.25,28.75), (-3.75,28.75), (-1.25,28.75), (1.25,28.75), (3.75,28.75), (6.25,28.75), (-13.75,26.25), (-11.25,26.25), (-8.75,26.25), (-6.25,26.25), (-3.75,26.25), (-1.25,26.25), (1.25,26.25), (3.75,26.25), (6.25,26.25), (8.75,26.25), (11.25,26.25), (-18.75,23.75), (-16.25,23.75), (-13.75,23.75), (-11.25,23.75), (-8.75,23.75), (-6.25,23.75), (-3.75,23.75), (-1.25,23.75), (1.25,23.75), (3.75,23.75), (6.25,23.75), (8.75,23.75), (11.25,23.75), (-21.25,21.25), (-18.75,21.25), (-16.25,21.25), (-13.75,21.25), (-11.25,21.25), (-8.75,21.25), (-6.25,21.25), (-3.75,21.25), (-1.25,21.25), (1.25,21.25), (3.75,21.25), (6.25,21.25), (8.75,21.25), (11.25,21.25), (13.75,21.25), (-23.75,18.75), (-21.25,18.75), (-18.75,18.75), (-16.25,18.75), (-13.75,18.75), (-11.25,18.75), (-8.75,18.75), (-6.25,18.75), (-3.75,18.75), (3.75,18.75), (6.25,18.75), (8.75,18.75), (11.25,18.75), (13.75,18.75), (16.25,18.75), (-23.75,16.25), (-21.25,16.25), (-18.75,16.25), (-16.25,16.25), (-13.75,16.25), (-11.25,16.25), (-8.75,16.25), (-6.25,16.25), (6.25,16.25), (8.75,16.25), (11.25,16.25), (13.75,16.25), (16.25,16.25), (-26.25,13.75), (-23.75,13.75), (-21.25,13.75), (-18.75,13.75), (-16.25,13.75), (-13.75,13.75), (-11.25,13.75), (-8.75,13.75), (-6.25,13.75), (6.25,13.75), (8.75,13.75), (11.25,13.75), (13.75,13.75), (16.25,13.75), (-26.25,11.25), (-23.75,11.25), (-21.25,11.25), (-18.75,11.25), (-16.25,11.25), (-13.75,11.25), (-11.25,11.25), (-8.75,11.25), (-6.25,11.25), (-3.75,11.25), (3.75,11.25), (6.25,11.25), (8.75,11.25), (11.25,11.25), (13.75,11.25), (-26.25,8.75), (-23.75,8.75), (-21.25,8.75), (-18.75,8.75), (-16.25,8.75), (-13.75,8.75), (-11.25,8.75), (-8.75,8.75), (-6.25,8.75), (-3.75,8.75), (-1.25,8.75), (1.25,8.75), (3.75,8.75), (6.25,8.75), (8.75,8.75), (11.25,8.75), (-28.75,6.25), (-26.25,6.25), (-23.75,6.25), (-21.25,6.25), (-18.75,6.25), (-16.25,6.25), (-13.75,6.25), (-11.25,6.25), (-8.75,6.25), (-6.25,6.25), (-3.75,6.25), (-1.25,6.25), (1.25,6.25), (3.75,6.25), (6.25,6.25), (8.75,6.25), (11.25,6.25), (-28.75,3.75), (-26.25,3.75), (-23.75,3.75), (-21.25,3.75), (-18.75,3.75), (-16.25,3.75), (-13.75,3.75), (-11.25,3.75), (-8.75,3.75), (-6.25,3.75), (-3.75,3.75), (-1.25,3.75), (1.25,3.75), (3.75,3.75), (6.25,3.75), (8.75,3.75), (-28.75,1.25), (-26.25,1.25), (-23.75,1.25), (-21.25,1.25), (-18.75,1.25), (-16.25,1.25), (-13.75,1.25), (-11.25,1.25), (-8.75,1.25), (-6.25,1.25), (-3.75,1.25), (-1.25,1.25), (1.25,1.25), (3.75,1.25), (-28.75,-1.25), (-26.25,-1.25), (-23.75,-1.25), (-21.25,-1.25), (-18.75,-1.25), (-16.25,-1.25), (-13.75,-1.25), (-11.25,-1.25), (-8.75,-1.25), (-6.25,-1.25), (-28.75,-3.75), (-26.25,-3.75), (-23.75,-3.75), (-21.25,-3.75), (-18.75,-3.75), (-16.25,-3.75), (-13.75,-3.75), (-11.25,-3.75), (-28.75,-6.25), (-26.25,-6.25), (-23.75,-6.25), (-21.25,-6.25), (-18.75,-6.25), (-16.25,-6.25), (-13.75,-6.25), (-26.25,-8.75), (-23.75,-8.75), (-21.25,-8.75), (-18.75,-8.75), (-16.25,-8.75), (-13.75,-8.75), (-26.25,-11.25), (-23.75,-11.25), (-21.25,-11.25), (-18.75,-11.25), (-16.25,-11.25), (-26.25,-13.75), (-23.75,-13.75), (-21.25,-13.75), (-18.75,-13.75), (-16.25,-13.75), (-1.25,-13.75), (1.25,-13.75), (-23.75,-16.25), (-21.25,-16.25), (-18.75,-16.25), (-16.25,-16.25), (-3.75,-16.25), (-1.25,-16.25), (1.25,-16.25), (3.75,-16.25), (-23.75,-18.75), (-21.25,-18.75), (-18.75,-18.75), (-16.25,-18.75), (-3.75,-18.75), (-1.25,-18.75), (1.25,-18.75), (3.75,-18.75), (-21.25,-21.25), (-18.75,-21.25), (-16.25,-21.25), (-1.25,-21.25), (1.25,-21.25), (-18.75,-23.75), (-16.25,-23.75), (-13.75,-23.75), (-13.75,-26.25)]
point_name_pairing = [("Green", mclover3_points),("Orange", azurite_points)]
# Robot deck setup constants
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
# Place the PCR tubes in this order
well_colors = {
'A1' : 'Red',
'B1' : 'Green',
'C1' : 'Orange'
}
# Initialize volume_used globally
volume_used = {}
def update_volume_remaining(current_color, quantity_to_aspirate):
global well_colors
global volume_used
rows = string.ascii_uppercase
cols_str = [str(i) for i in range(1, 13)] # Columns 1 to 12
if current_color not in volume_used:
volume_used[current_color] = 0
# Find the current well for this color
current_well_for_color = None
for well, color in list(well_colors.items()):
if color == current_color:
current_well_for_color = well
break
if current_well_for_color is None:
raise ValueError(f"Color {current_color} not found in well_colors for volume update.")
if (volume_used[current_color] + quantity_to_aspirate) > 250:
row_letter = current_well_for_color[0]
col_number_str = current_well_for_color[1:]
next_col_index = cols_str.index(col_number_str) + 1
if next_col_index >= len(cols_str):
raise IndexError(f"Ran out of wells for color {current_color} in row {row_letter} (max column reached)!")
next_well = f"{row_letter}{cols_str[next_col_index]}"
# Remove the old well from well_colors map, and add the new one.
# This is safe because each color is assumed to have its own row.
del well_colors[current_well_for_color]
well_colors[next_well] = current_color
volume_used[current_color] = quantity_to_aspirate # Reset volume for new well
else:
volume_used[current_color] += quantity_to_aspirate
def run(protocol):
##############################################################################
### Load labware, modules and pipettes
##############################################################################
# Tips
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
# Pipettes
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
# Modules
temperature_module = protocol.load_module('temperature module gen2', COLORS_DECK_SLOT)
# Temperature Module Plate
temperature_plate = temperature_module.load_labware('opentrons_96_aluminumblock_generic_pcr_strip_200ul',
'Cold Plate')
# Choose where to take the colors from
color_plate = temperature_plate
# Agar Plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate') ## TA MUST CALIBRATE EACH PLATE!
# Get the top-center of the plate, make sure the plate was calibrated before running this
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
# Helper function (color location)
def location_of_color(color_string):
for well,color in well_colors.items():
if color.lower() == color_string.lower():
return temperature_plate[well]
raise ValueError(f"No well found with color {color_string}")
# For this lab, instead of calling pipette.dispense(1, loc) use this: dispense_and_detach(pipette, 1, loc)
def dispense_and_detach(pipette, volume, location):
"""
Move laterally 5mm above the plate (to avoid smearing a drop); then drop down to the plate,
dispense, move back up 5mm to detach drop, and stay high to be ready for next lateral move.
5mm because a 4uL drop is 2mm diameter; and a 2deg tilt in the agar pour is >3mm difference across a plate.
"""
assert(isinstance(volume, (int, float)))
above_location = location.move(types.Point(z=location.point.z + 5)) # 5mm above
pipette.move_to(above_location) # Go to 5mm above the dispensing location
pipette.dispense(volume, location) # Go straight downwards and dispense
pipette.move_to(above_location) # Go straight up to detach drop and stay high
# Print pattern by iterating over lists
for idx, (current_color, point_list) in enumerate(point_name_pairing): # Renamed i to idx to avoid conflict
# Skip the rest of the loop if the list is empty
if not point_list:
continue
pipette_20ul.pick_up_tip()
max_aspirate = int(18 // POINT_SIZE) * POINT_SIZE
quantity_to_aspirate = min(len(point_list)*POINT_SIZE, max_aspirate)
# Get the initial well for this color before any volume updates
initial_aspirate_well = location_of_color(current_color)
# Update volume (this might change `well_colors` for `current_color`)
update_volume_remaining(current_color, quantity_to_aspirate)
# Aspirate from the (potentially updated) location
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Iterate over the current points list and dispense them, refilling along the way
for j in range(len(point_list)):
x, y = point_list[j]
adjusted_location = center_location.move(types.Point(x, y))
dispense_and_detach(pipette_20ul, POINT_SIZE, adjusted_location)
if pipette_20ul.current_volume == 0 and len(point_list[j+1:]) > 0:
# Need to refill
refill_quantity = min(len(point_list[j+1:])*POINT_SIZE, max_aspirate)
# Get the current source well for this color *before* updating volume, in case it changes
previous_refill_well = location_of_color(current_color)
# Update volume and potentially move the color to a new physical well
update_volume_remaining(current_color, refill_quantity)
# Get the (potentially new) source well for this color
new_refill_well = location_of_color(current_color)
if new_refill_well != previous_refill_well:
# If the source well has changed for this color, we must drop the tip and pick up a new one
pipette_20ul.drop_tip()
pipette_20ul.pick_up_tip()
# Now aspirate from the correct (potentially new) well
pipette_20ul.aspirate(refill_quantity, new_refill_well)
# Drop tip between each color
pipette_20ul.drop_tip()
Before running on the real robot, the protocol was validated using the Opentrons simulator.
The robot successfully deposited bacteria following the coordinate map.
After incubation, bacterial growth revealed the intended image on the agar plate.
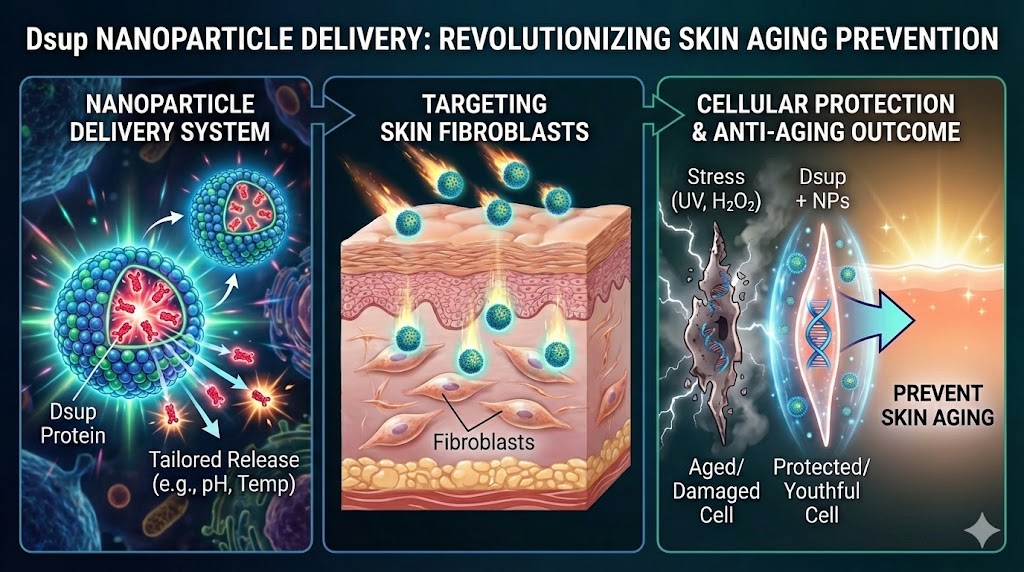
This project proposes to automate the screening and characterization of nanoparticle delivery systems for the delivery of the damage suppressor protein (Dsup) — a nucleosome-binding protein from Ramazzottius varieornatus (tardigrade). Dsup has been demonstrated to protect mammalian cells from oxidative stress and UV-induced DNA damage.
The goal is to determine which nanoparticle formulation most effectively delivers Dsup into human dermal fibroblasts, improving resistance to oxidative stress (H₂O₂ exposure). The long-term application is skin regeneration and anti-aging therapies.