Week 4: Protein Design Part 1
Conceptual Questions
Why do humans eat beef but do not become a cow, eat fish but do not become fish?
As we eat other materials, the proteins and molecules are broken down through our digestive system, leaving us with building blocks to support our cell replication processes.
Why are there only 20 natural amino acids?
These 20 amino acids were created as life was forming, and after the ‘frozen accident’, the proteins that developed at this time seem to have standardized these 20 amino acids. There were more and are more amino acids, however this connection between protein and AA during early evolution created this set of 20.
Can you make other non-natural amino acids? Design some new amino acids.
Amino acids are primarily composed of hydrogen, oxygen, carbon, and nitrogen and as proven from the beginnings of RNA, there are many non-standard amino acids that could be made in fact many non-natural AAs already exist. In terms of function, I would design an AA that could create florescent tags for injury sites, helping identify issues that are less visible like endometreosis or even to find a bug bite from tics. It could help speed up wound identification and reduce time lost in exploritory tests.
Where did amino acids come from before enzymes that make them, and before life started?
Many natural events have created the primodial soup that led to life as we know it, and the ingredients of AAs seem to have formed in a similar way.
If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
I would expect righthandedness because ‘D’ typicially refers to right handed molecules and ‘L’ refers to left.
Can you discover additional helices in proteins?
Probably! Given that there is a high percentage of proteins that are still to be discovered and studies, there are probably proteins with additional helices.
Why are most molecular helices right-handed?
DNA is right-handed, but also many aspects of the natural world have a handedness, like the right hand test in physics as well. This seems to be connected back to thermodynamics and create helices that are more stable.
Why do β-sheets tend to aggregate? What is the driving force for β-sheet aggregation?
Their structure is flat, making it easy to pack, and the edges are designed with hydrogen backbones that help form hydrogen bonds with other beta sheets.
Why do many amyloid diseases form β-sheets? Can you use amyloid β-sheets as materials?
Many amyloid diseases happen when proteins misfold, and they form beta sheets because of their thermodynamic stability and their straight hydrogen bond structure allows for tighter folds. They can probably be used to support material tensile strength and rigidity, but I don’t think they classify as materials themselves.
Protein Analysis and Visualization
Fro this analysis, I am focusing on silaffin once again, following through from week 2. Silaffin is the protein responsible for the lacy biosilica patterning that diatom frustules form. I am interested in modifying this protein because tuning that pattern could allow for more targeted and intentional interactions between cement rubble and diatom placement for the final project.
silaffin sequence (from marine diatom: Thalassiosira pseudonana)
MKVTTSIITLLFASCGAADVQRVLEDVTEPAVTTPAATPAPITPEPATPAPTICEGRNFYRDDDTGKCSNEATGGIYGTLIECCVAISGSDSCPYVDICNTLQPSPSPETNEPSAKPITAAPISSAPVSAAPVTSAPVAAPVETTSMTGPTTIVASIVSTNAPSSTNAPSSSLEAVVTRIPVETTNTASPTTTAASIVSTNAPSSSPEAVVTPRPTFRPSPKGTESNTFPASIASDVMFDPARSEPTFTPTSSSQPSSSEPTLSPSVSKEPTRYPTSSPSHSPTKSPSKSPSSSPTTSPSASPTETPTETPTESPTELPTLSPTEFPSLSPTLSPTWSPTGYPTLAPSPSPSISSAPSVSSAPSSSPSISSAPSVSSAPSKNFGFLPGRNEMPTISPTEDHYFFGKSHKSHKSKATKTLKVSKSGKSSKSSKSSGSRPLFGVSQLSEGIAAGYAKSSGRSSQQAVGSWMPVAAACILGALSFFLN
silaffin has 485 amino acids, with S appearing most frequently occuring 99 times. It doesn’t belong to any protein families, which could be because this protein has a very low confidence score and is more rare, leading to less information on its structure and protein family.
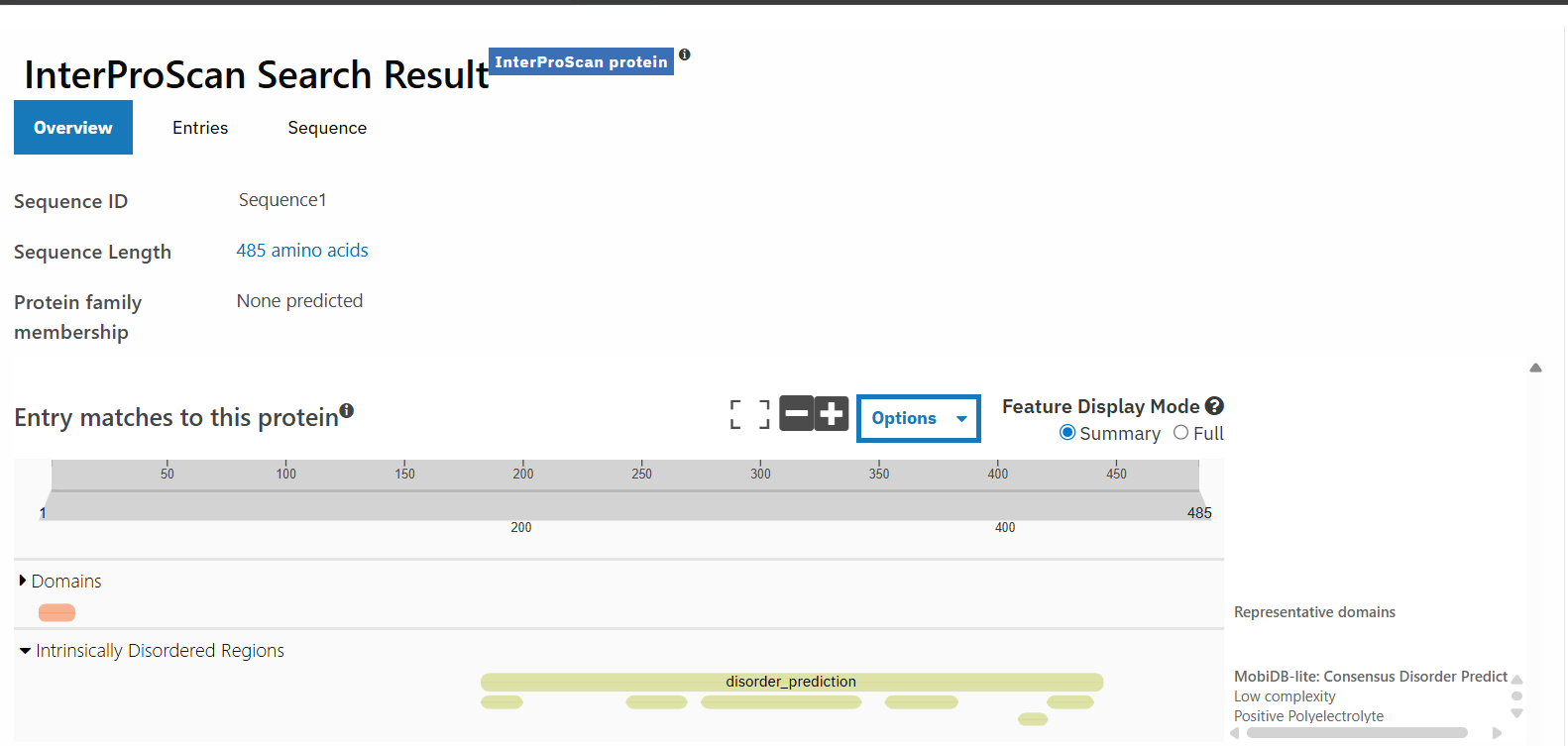

screenshot from uniprot
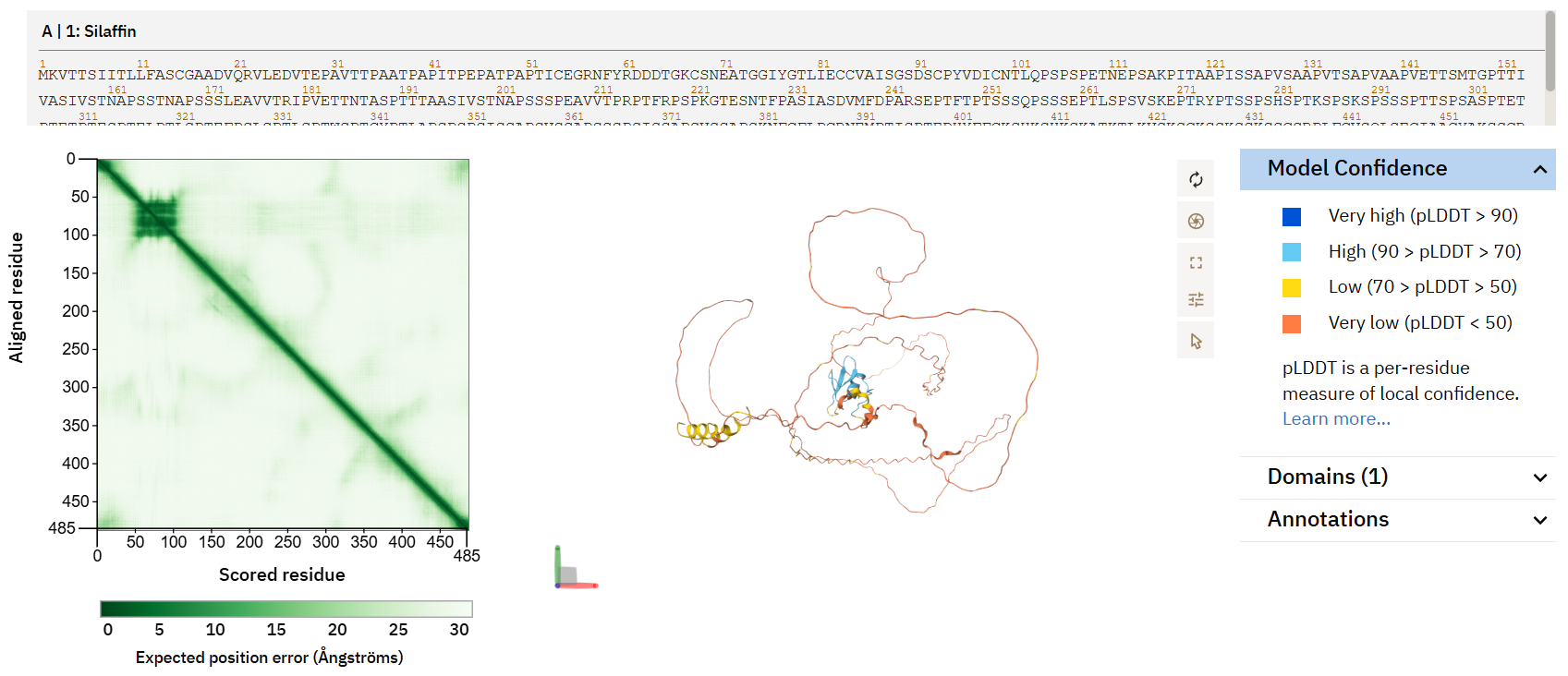

screenshot from alphafold
With a less documented protein, there is also more likelihood for error. While some areas of the protein with better confidence have a defined beta sheets and alpha helices, most of the protein is modeled with very low confidence.
ML Design Tools
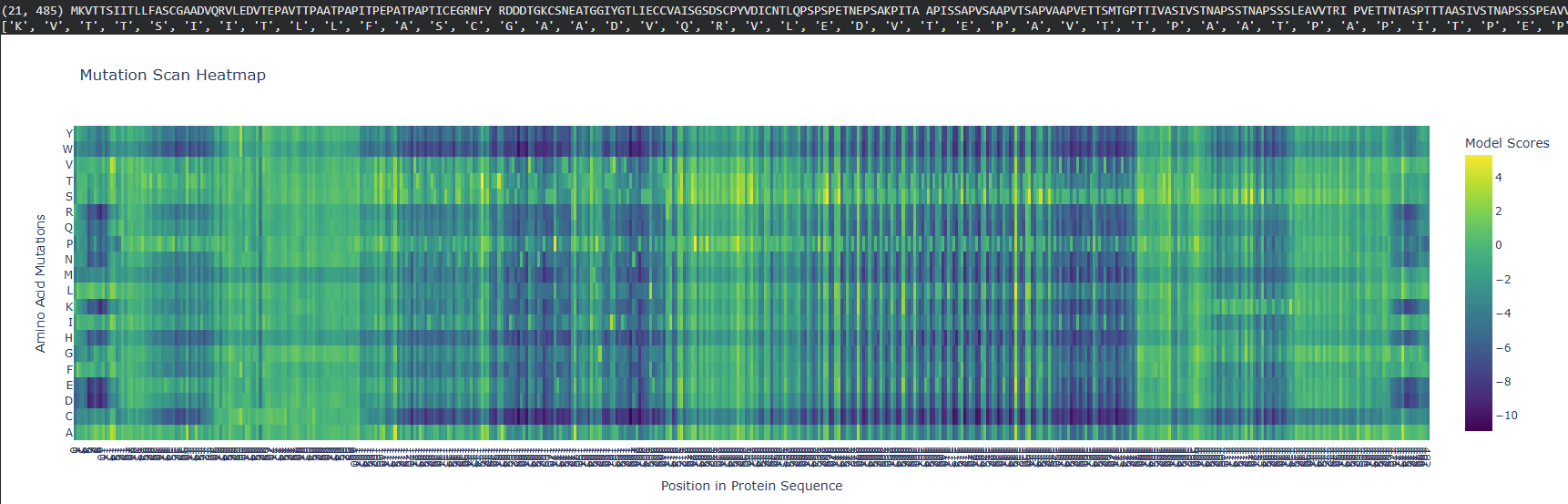
 the mutation scan shows that amino acids that are most likely to mutate (more yellow in this diagram), are the L,R, and S amino acids. Position 27 is most likely to have a mutation as well.
the mutation scan shows that amino acids that are most likely to mutate (more yellow in this diagram), are the L,R, and S amino acids. Position 27 is most likely to have a mutation as well.
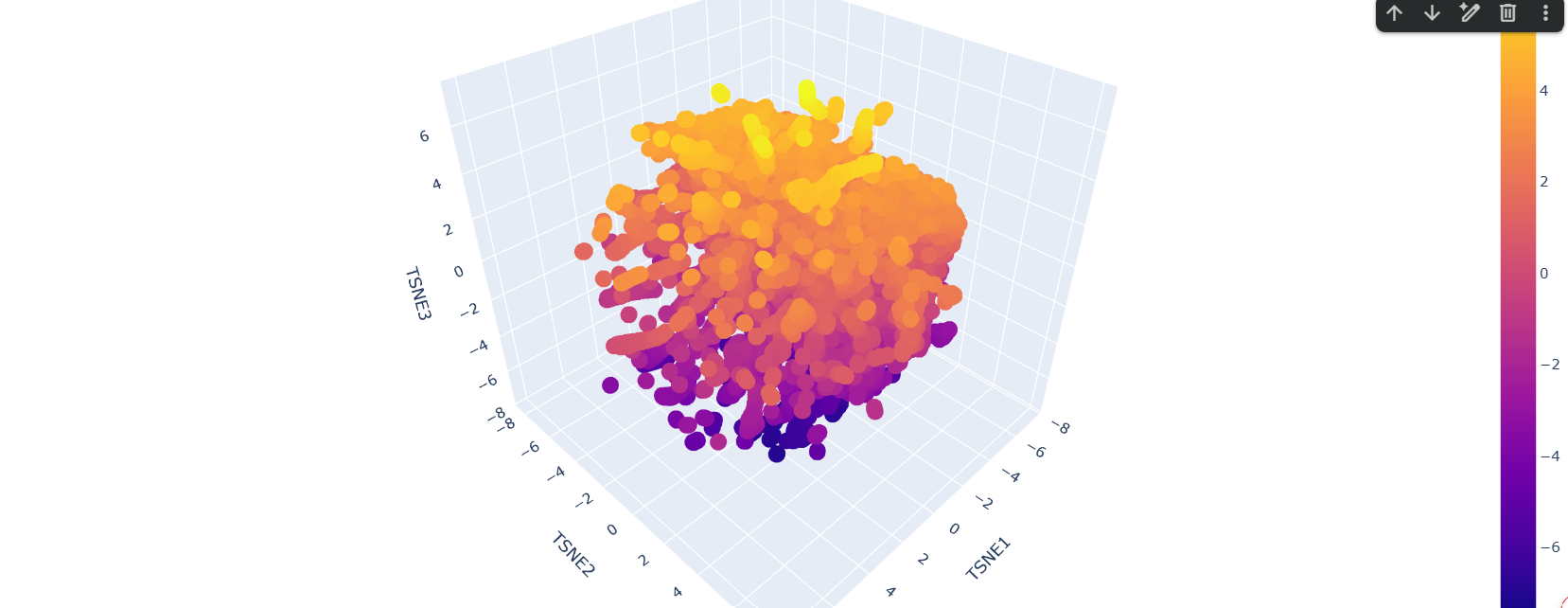
 latent space analysis shows concentrated areas. Some of the matches include proteins from organisms ranging from fungi to humans with seemingly low similarity in function. This could be due to the low confidence in the silaffin protein, so the comparative space could be drawing upon very different organisms since there is very little information to respond to.
latent space analysis shows concentrated areas. Some of the matches include proteins from organisms ranging from fungi to humans with seemingly low similarity in function. This could be due to the low confidence in the silaffin protein, so the comparative space could be drawing upon very different organisms since there is very little information to respond to.