Week 2 HW: Dna-Read-Write-and-Edit
Part 1: Benchling & In-silico Gel Art
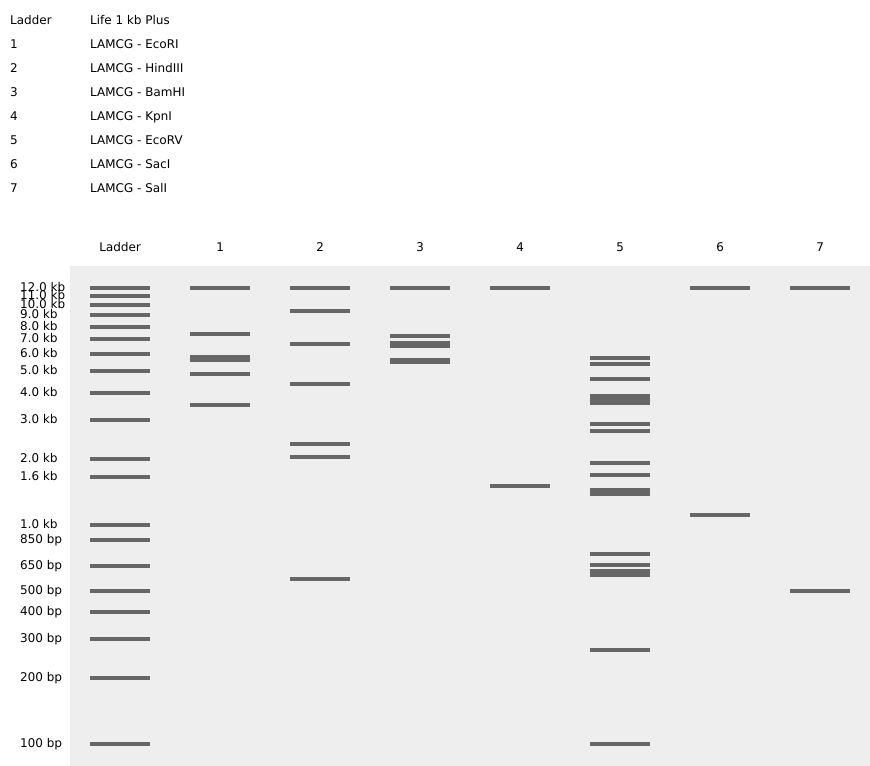
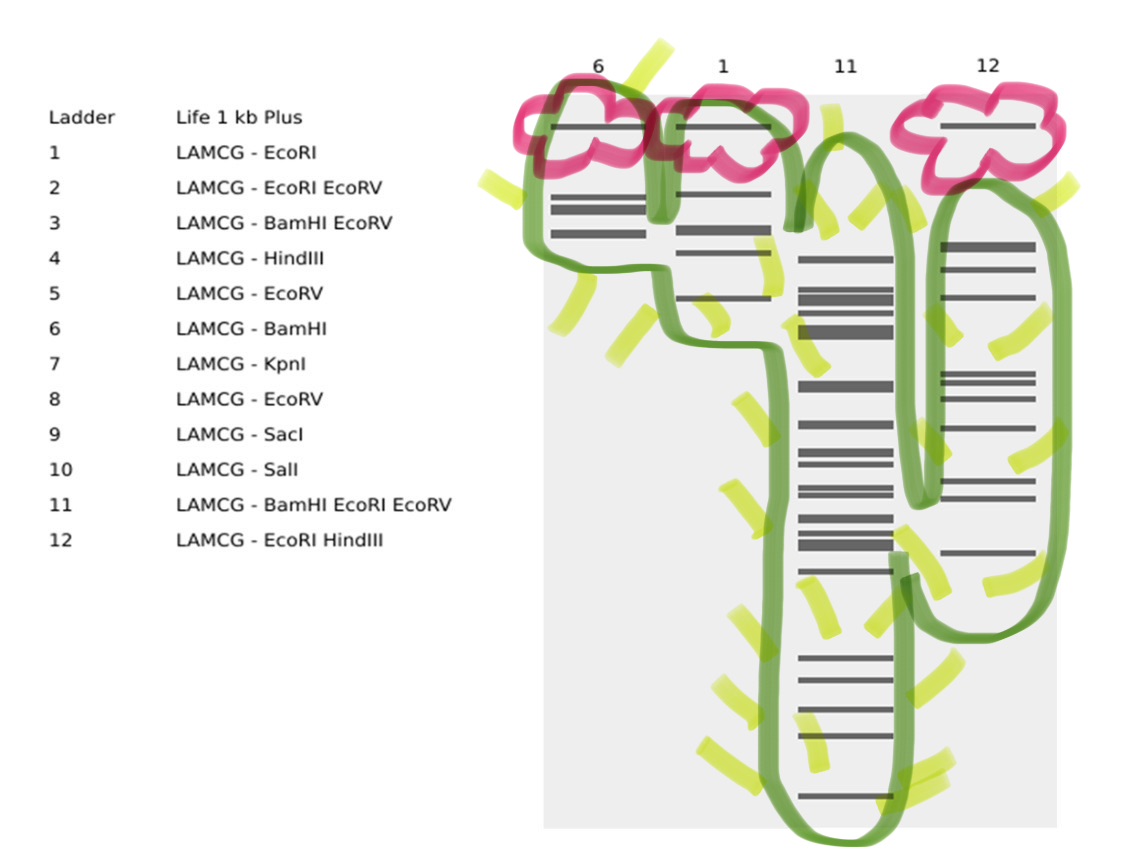
I simulated the Restriction Enzyme Digestion in Benchling to create a design. I found it initially difficult to visualise patterns or images with the 7 restriction enzymes. I therefore decided to mix certain enzymes in the same wells to generate more DNA fragments and explore shapes further.



Part 3: DNA Design Challenge
3.1 Which protein have you chosen and why? Using one of the tools described in the recitation (NCBI, UniProt, Google), obtain the protein sequence for the protein you chose.Name of protein: psiH (tryptamine 4-monooxygenase)
I chose this specific protein as it relates to my project idea from homework 1. PsiH catalyses the 4-hydroxylation of tryptamine to 4-hydroxytryptamine, which is an essential and unique part of psilocybin biosynthesis that allows for the production of psilocin (the active therapeutic metabolite). This P450 enzyme (psiH) acts as a critical rate‐limiting step of psilocybin production. Furthermore, it needs exact heme binding and substrate fit, which is rare in nature and tough to engineer in E. coli (Huang et al., 2025). This makes PsiH the technical core of my biosynthetic pathway design from Homework 1, where engineered E. coli would produce psilocybin locally to activate gut serotonin signalling for IBD treatment (Robinson et al., 2023).
Refrences:
Amino Acid Protein Sequence:
MIAVLFSFVIAGCIYYIVSRRVRRSRLPPGPPGIPIPFIGNMFD MPEESPWLTFLQWGRDYNTDILYVDAGGTEMVILNTLETITDLLEKRGSIYSGRLEST MVNELMGWEFDLGFITYGDRWREERRMFAKEFSEKGIKQFRHAQVKAAHQLVQQLTKT PDRWAQHIRHQIAAMSLDIGYGIDLAEDDPWLEATHLANEGLAIASVPGKFWVDSFPS LKYLPAWFPGAVFKRKAKVWREAADHMVDMPYETMRKLAPQGLTRPSYASARLQAMDL NGDLEHQEHVIKNTAAEVNVGGGDTTVSAMSAFILAMVKYPEVQRKVQAELDALTNNG QIPDYDEEDDSLPYLTACIKELFRWNQIAPLAIPHKLMKDDVYRGYLIPKNTLVFANT WAVLNDPEVYPDPSVFRPERYLGPDGKPDNTVRDPRKAAFGYGRRNCPGIHLAQSTVW IAGATLLSAFNIERPVDQNGKPIDIPADFTTGFFRHPVPFQCRFVPRTEQVSQSVSGP
Source:3.2 DNA Reverse Translation:
atgattgcggtgctgtttagctttgtgattgcgggctgcatttattatattgtgagccgc cgcgtgcgccgcagccgcctgccgccgggcccgccgggcattccgattccgtttattggc aacatgtttgatatgccggaagaaagcccgtggctgacctttctgcagtggggccgcgat tataacaccgatattctgtatgtggatgcgggcggcaccgaaatggtgattctgaacacc ctggaaaccattaccgatctgctggaaaaacgcggcagcatttatagcggccgcctggaa agcaccatggtgaacgaactgatgggctgggaatttgatctgggctttattacctatggc gatcgctggcgcgaagaacgccgcatgtttgcgaaagaatttagcgaaaaaggcattaaa cagtttcgccatgcgcaggtgaaagcggcgcatcagctggtgcagcagctgaccaaaacc ccggatcgctgggcgcagcatattcgccatcagattgcggcgatgagcctggatattggc tatggcattgatctggcggaagatgatccgtggctggaagcgacccatctggcgaacgaa ggcctggcgattgcgagcgtgccgggcaaattttgggtggatagctttccgagcctgaaa tatctgccggcgtggtttccgggcgcggtgtttaaacgcaaagcgaaagtgtggcgcgaa gcggcggatcatatggtggatatgccgtatgaaaccatgcgcaaactggcgccgcagggc ctgacccgcccgagctatgcgagcgcgcgcctgcaggcgatggatctgaacggcgatctg gaacatcaggaacatgtgattaaaaacaccgcggcggaagtgaacgtgggcggcggcgat accaccgtgagcgcgatgagcgcgtttattctggcgatggtgaaatatccggaagtgcag cgcaaagtgcaggcggaactggatgcgctgaccaacaacggccagattccggattatgat gaagaagatgatagcctgccgtatctgaccgcgtgcattaaagaactgtttcgctggaac cagattgcgccgctggcgattccgcataaactgatgaaagatgatgtgtatcgcggctat ctgattccgaaaaacaccctggtgtttgcgaacacctgggcggtgctgaacgatccggaa gtgtatccggatccgagcgtgtttcgcccggaacgctatctgggcccggatggcaaaccg gataacaccgtgcgcgatccgcgcaaagcggcgtttggctatggccgccgcaactgcccg ggcattcatctggcgcagagcaccgtgtggattgcgggcgcgaccctgctgagcgcgttt aacattgaacgcccggtggatcagaacggcaaaccgattgatattccggcggattttacc accggcttttttcgccatccggtgccgtttcagtgccgctttgtgccgcgcaccgaacag gtgagccagagcgtgagcggcccg
Source
3.3 Codon optimisation
In your own words, describe why you need to optimise codon usage. Which organism have you chosen to optimise the codon sequence for and why?
Although different codons can code for the same amino acid, each species/organism has a bias for its codon preferences. This is done by changing/optimising the DNA codon sequence (not the amino-acid sequence) of the protein in order to match the codon preferences of the host organism (Cheema et al., 2022). If I were to take a human gene and insert it into a bacterium, it might use certain codons that the bacterium wouldn’t/would rarely use. This, in turn, makes the translation process slower or incomplete, resulting in a low protein yield (Creative BioLabs,2025). Therefore, by optimising the sequence with a specific host, I can make the translation process faster and more reliable.
For this specific exercise, I chose to optimise the psiH protein for E. Coli. I made this choice because E. coli is one of the most commonly used hosts for genetic engineering due to its rapid culture rate, simple nutritional needs and well-understood genetics (Adamczyk and Reed, 2017). Additionaly it is relatively cheap to culture (Francis and Page,2010). In relation to Homework 1, during the time of this course, E. coli is an appropriate host for prototyping the psilocybin pathway to conceptually extend toward microbiome-targeted therapies.
sources
What technologies could be used to produce this protein from your DNA? Describe in your own words how the DNA sequence can be transcribed and translated into a protein.
A variety of technologies could be used to produce the psiH protein from its DNA. One of the most suitable that we have discussed in lectures would be Gibson Assembly. It would allow us to stitch together multiple DNA fragments (promoter, Ribosome binding site, my optimised DNA sequence of psiH, terminator…) inside the plasmid of the E. Coli without the use of restriction enzymes. You may also use external platforms like Twist Bioscience to order your full plasmid by giving them the appropriate optimised sequence.
The DNA sequence may be transcribed and translated into a protein with the following steps:
Sources:
Part 4: Prepare a Twist DNA Synthesis Order
Building my DNA Insert Sequence
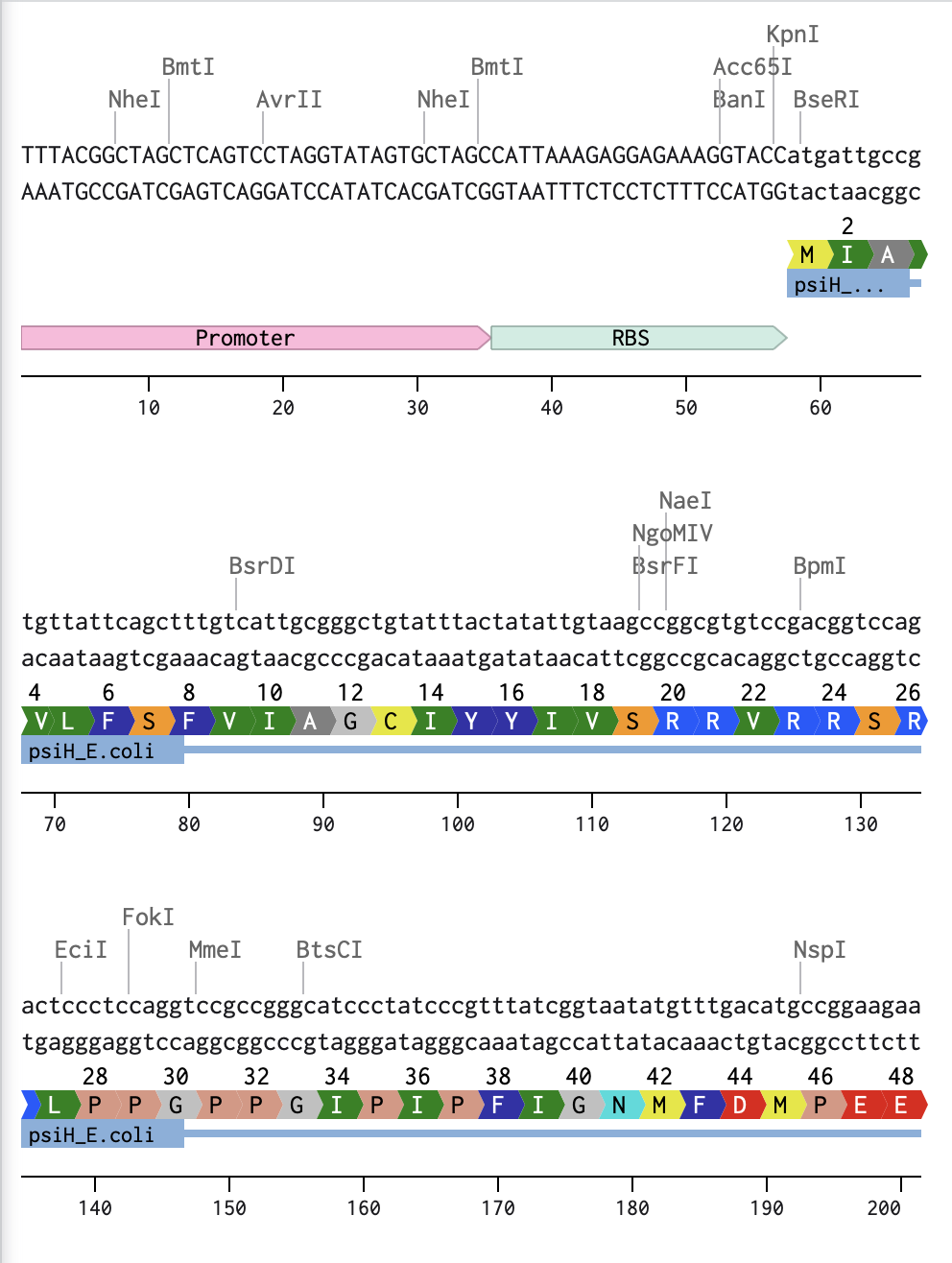
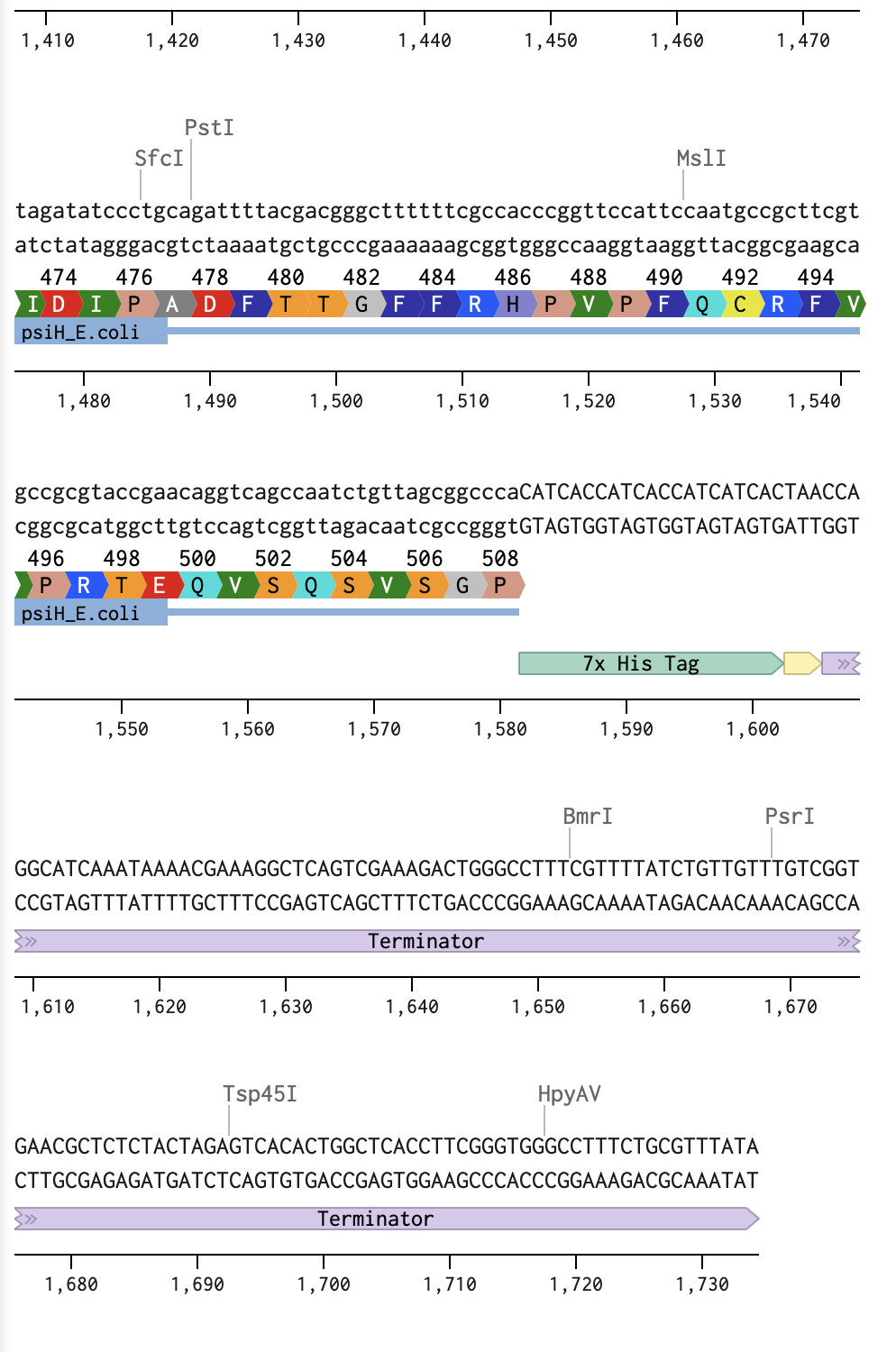
I began by optimising my psiH translated protein DNA sequence in Benchling with a linear topology and optimising it for E. coli. I then added the reading direction (forward), and the given DNA sequences highlighted in the homework (Promoter, RBS, Coding Sequence, 7x His Tag, Stop Codon, Terminator).
Linear Map of the entire sequence:
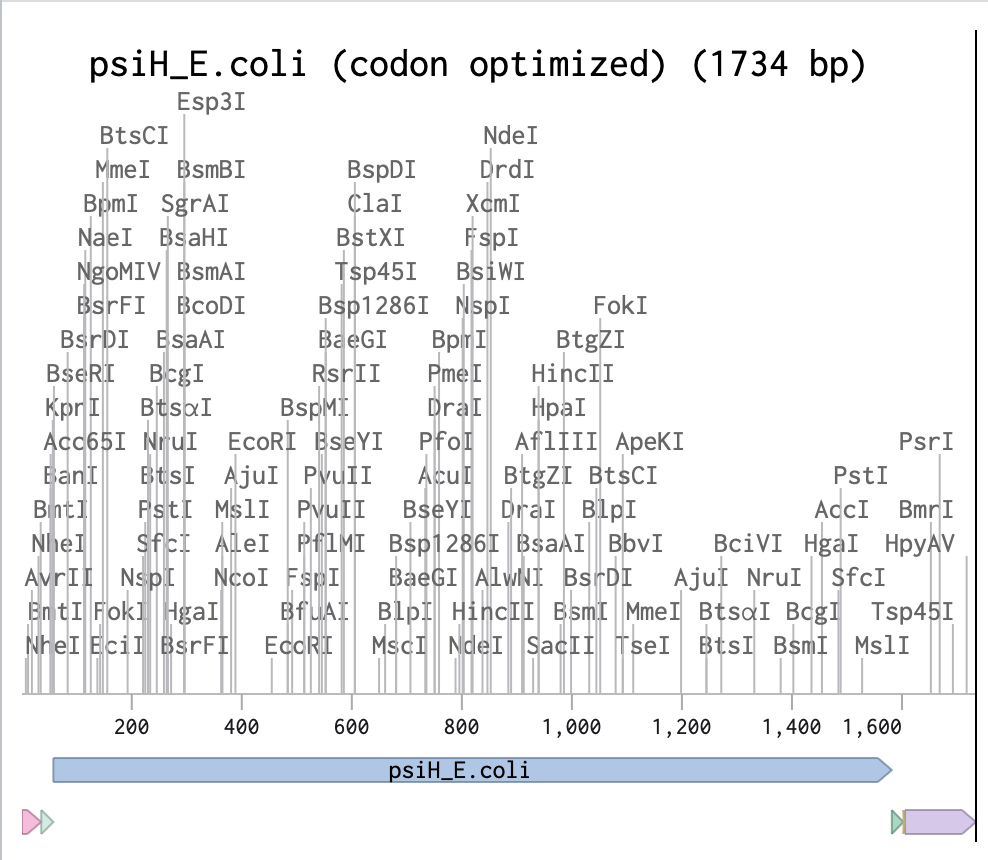

Final Sequence Benchling link!
Building my Full Plasmid Sequence
After downloading my insert sequence (expression cassette) as a FASTA file and uploading it into my Twist account, selecting an appropriate vector (pTwist Amp High Copy), I was able to download the full plasmid sequence (GenBank). I then imported the GenBank file of my plasmid back into Benchling.
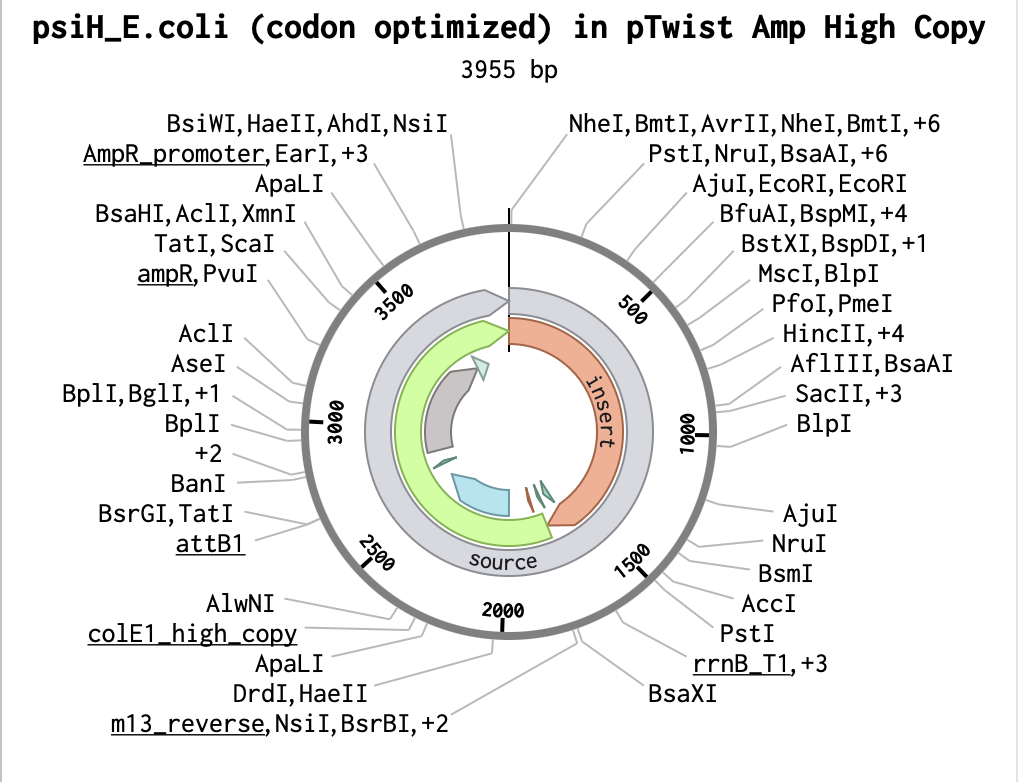

Part 5: DNA Read/Write/Edit
5.1 DNA Read
1) What DNA would you want to sequence (e.g., read) and why?
I want to sequence the 3955 bp E. coli plasmid containing the codon-optimised PsiH gene (tryptamine 4-monooxygenase from Psilocybe cubensis) that I developed with Twist Bioscience technology for exercise 4. This would allow for the verification of the construct (Gibson Assembly) of the plasmid, to verify that there are no mutations in the critical heme-binding site essential for the following steps of psilocibin synthesis. Ultimately, the goal would be to create a baseline sequence for future IBD therapy-scale engineering (Adams et al., 2019).
Sources
1) In the lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA, and why?
I initially considered Sanger sequencing due to its high accuracy (~99.9%) and effectiveness for targeted validation of small DNA regions. However, my E. coli psiH plasmid has 3955 bp, which exceeds Sanger’s read length of ~800 bp per reaction. Full coverage would require multiple reads; consequently, needing the development of multiple primers, which creates a time-consuming and costly process (AAT Bioquest, 2024).
I therefore continued my research and thought Oxford Nanopore Technologies' MinION device was a more appropriate fit (Oxford Nanopore Technologies, 2024). This technology generates long reads (>20 kb) capable of sequencing my entire 3955 bp plasmid in 1-2 reads with >99% accuracy (Brown et al., 2023). This approach would allow for the verification of the complete PsiH integration in the plasmid, promoter/RBS/terminator junctions and detect assembly errors.
Sources
1) Is your method first-, second-, or third-generation or other? How so?
Oxford Nanopore Technologies' MinION is a 3rd generaation sequencer. This means that it can read much longer sequences than any 1st or 2nd generation sequencing technologies (Hilt and Ferrieri, 2022). Additionally, it is one of the rare sequencing technologies that allows for real-time analysis. (Oxford Nanopore Technologies,2021)
Sources
2) What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
Input: Purified plasmid DNA from my engineered E. coli (3955 bp PsiH construct)
Preparation
Sources
3) What are the essential steps of your chosen sequencing technology? How does it decode the bases of your DNA sample (base calling)?
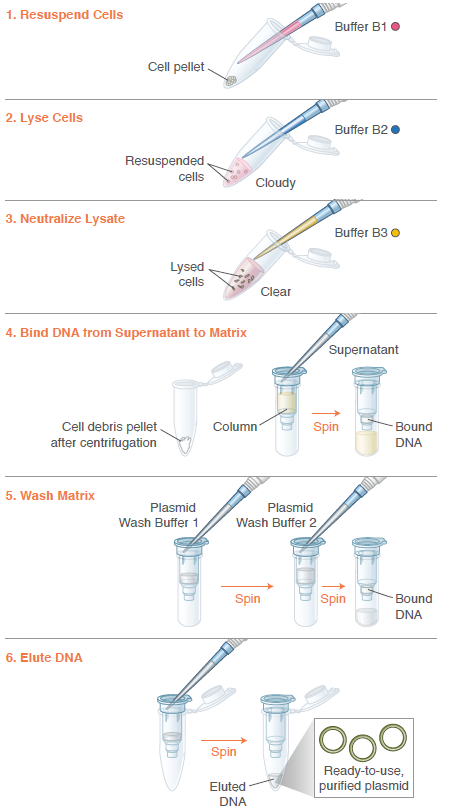
 Image: Instructions from Monarch Spin Plasmid Miniprep Kit, 2026.
Image: Instructions from Monarch Spin Plasmid Miniprep Kit, 2026.
Sources
4) What is the output of your chosen sequencing technology?
Output: A FASTQ file containing the complete 3955 bp sequence of my optimised E.coli PsiH plasmid.
Sources
5.2 DNA Write
I will synthesise the recombinant DNA of the engineered E. Coli as it contains the introduced plasmid from the psilocybin producing fungi.
5.2 DNA Write
(i)What DNA would you want to synthesise (e.g., write) and why?
I would like to synthesise the codon-optimised PsiH gene. This is an essential step for the development of this project, as fungal codons usually don’t express well in bacteria (Naqvi et al., 2016). Synthesising it will allow for a higher level of psiH with the ultimate goal of psilocin production.
Sources
(ii)What technology or technologies would you use to perform this DNA synthesis, and why?
I would use the services of Twist Bioscience to ensure a correct synthesis with no PCR errors, and be ready for the Gibson Assembly of my plasmid.
1) What are the essential steps of your sequencing methods?
Twist uses a Phosphoramidite synthesis method. These are the four essential steps:
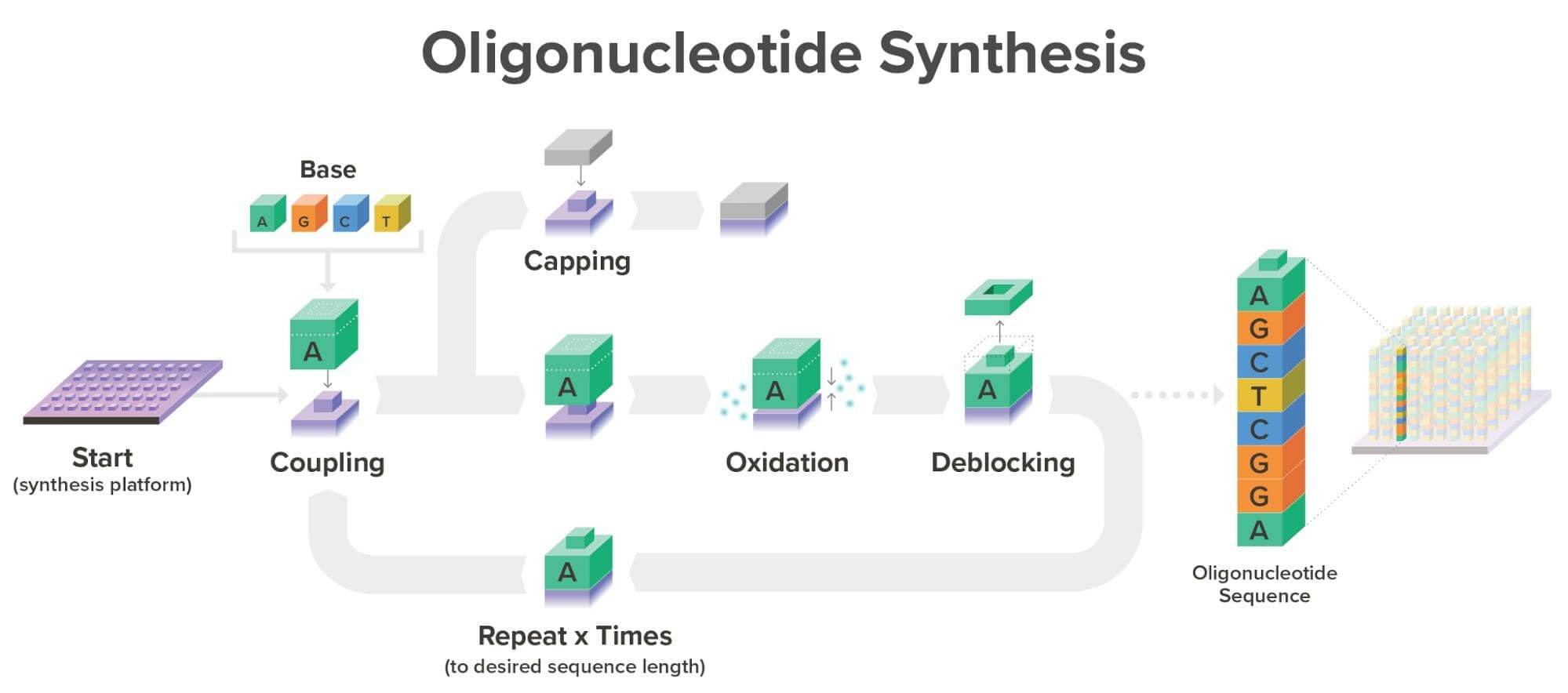
 Image: Twist Bioscience Website, 2026.
Image: Twist Bioscience Website, 2026.
Sources
2) What are the limitations of your sequencing method (if any) in terms of speed, accuracy, and scalability?
Phosphoramidite synthesis makes short DNA pieces (200-1500 bp) (Hughes and Ellington,2017), but Twist Bioscience assembles them into my full 2155 bp PsiH gene with error correction anyway, so it wouldn’t affect the actual synthesis process of my DNA. However, another issue I would like to raise is the use of hazardous reagents in reactions, washing and purification processes of this type of synthesis, which raises significant safety and environmental issues which shouldn’t be undermined (Gao et al., 2025).
Sources
5.3 DNA Edit
What DNA would you want to edit and why?
I’d like to edit the PsiH gene via mutagenesis. This would allow for the optimisation of tryptamine binding, which in turn will enhance enzyme activity in E. coli without needing host genome changes (Huang et al., 2025).
Sources
(ii)What technology or technologies would you use to perform these DNA edits and why?
I would use the site-directed mutagenesis method to examine the relationship between function and structure of my selected protein (Zhang et al., 2009). To do so, I would amplify my Twist PsiH plasmid with mutant primers in order to enhance the tryptamine binding in E.coli host.
Sources
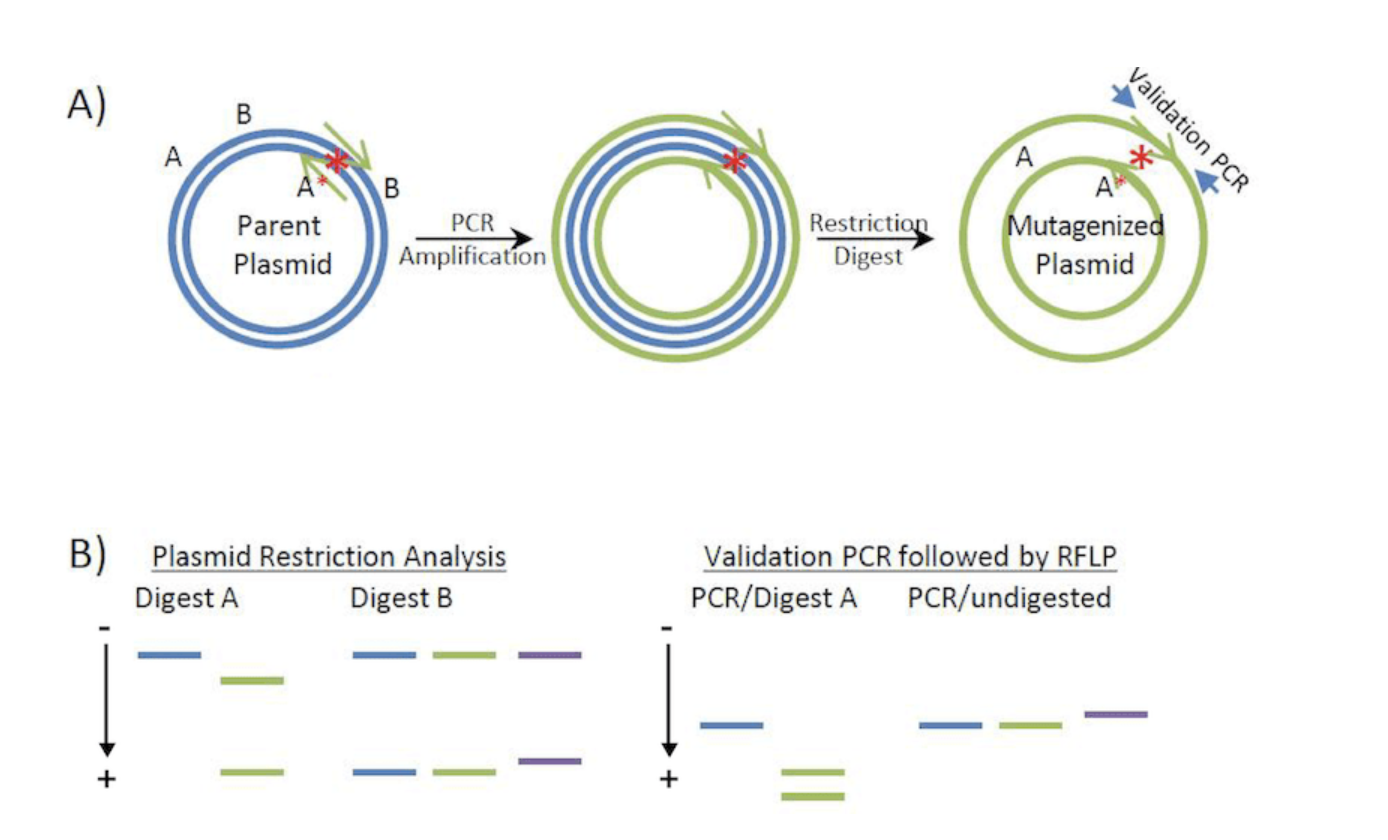
1) How does your technology of choice edit DNA? What are the essential steps?
PCR site-directed mutagenesis edits PsiH using custom primers carrying my mutation (one base change to improve tryptamine binding) to amplify the full Twist plasmid during PCR, producing nicked circular copies with the incorporated alteration.
2) What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
Sources
3) What are the limitations of your editing methods (if any) in terms of efficiency or precision?
Considering that I am only editing a small part of my DNA, using this method has a relatively high precision rate for a single alteration. Prior steps, though, like a mistake in the primer preparation, which will cause wrong mutations (Alvarez, 2024), as well as a very large plasmid, will cause a drop in accuracy (Jacobs et al., 2011).
Sources