Week 3 HW: Lab Automation
Assignment: Python Script for Opentrons Artwork
1) Generate an artistic design using the GUI at opentrons-art.rcdonovan.com.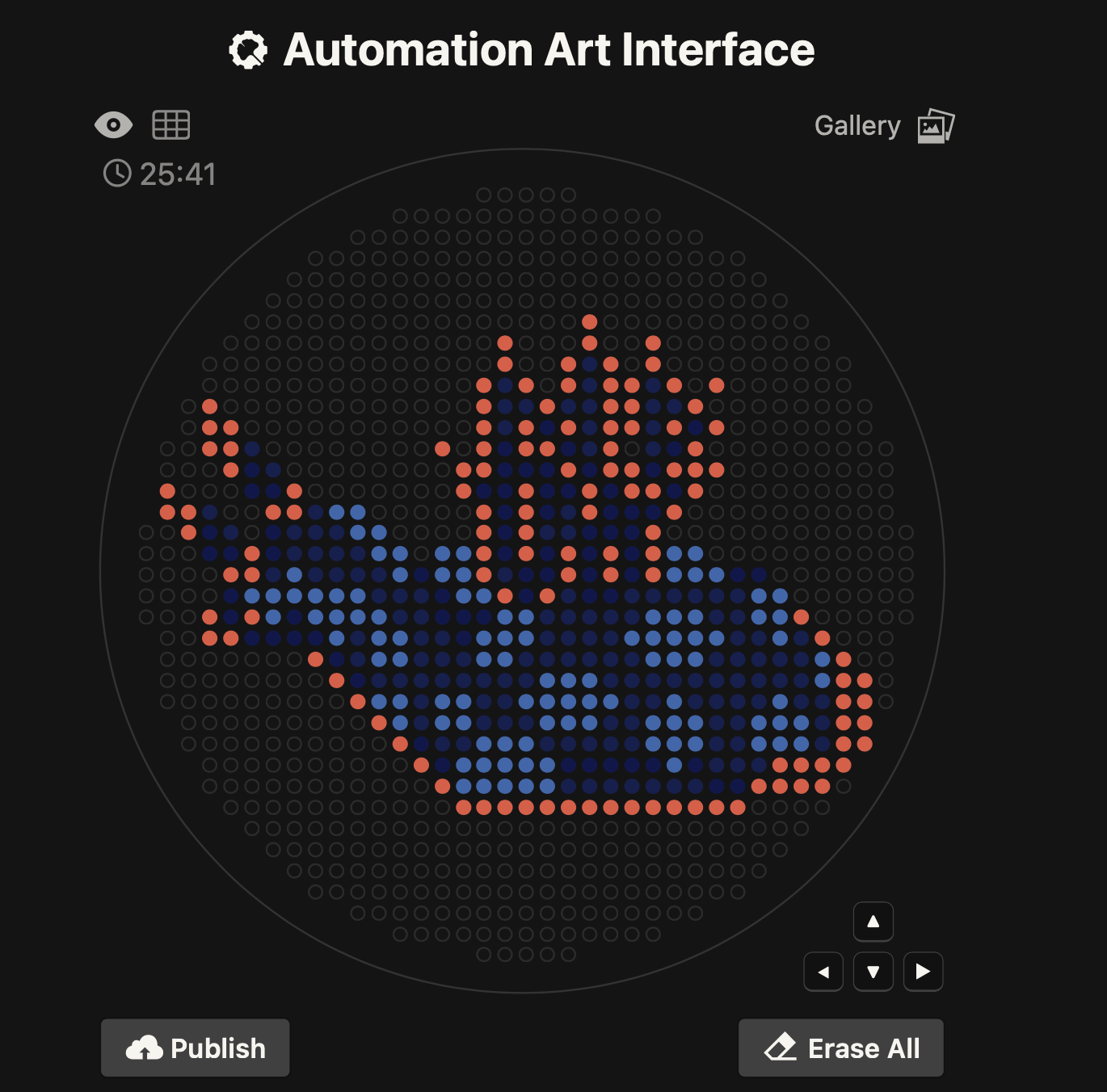

My OpenTron design is inspired by the nudibranch (sea slug) from the Mollusca phylum.

 Image source: Siewert, I. (2014). Nudibranch - Marine Life in Thailand. [online] Diving in Phuket Thailand. Available at: https://www.diving-thailand-phuket.com/nudibranch-marine-life-thailand/.
Image source: Siewert, I. (2014). Nudibranch - Marine Life in Thailand. [online] Diving in Phuket Thailand. Available at: https://www.diving-thailand-phuket.com/nudibranch-marine-life-thailand/.
Using the coordinates from the GUI, follow the instructions in the HTGAA26 Opentrons Colab to write your own Python script which draws your design using the Opentrons.
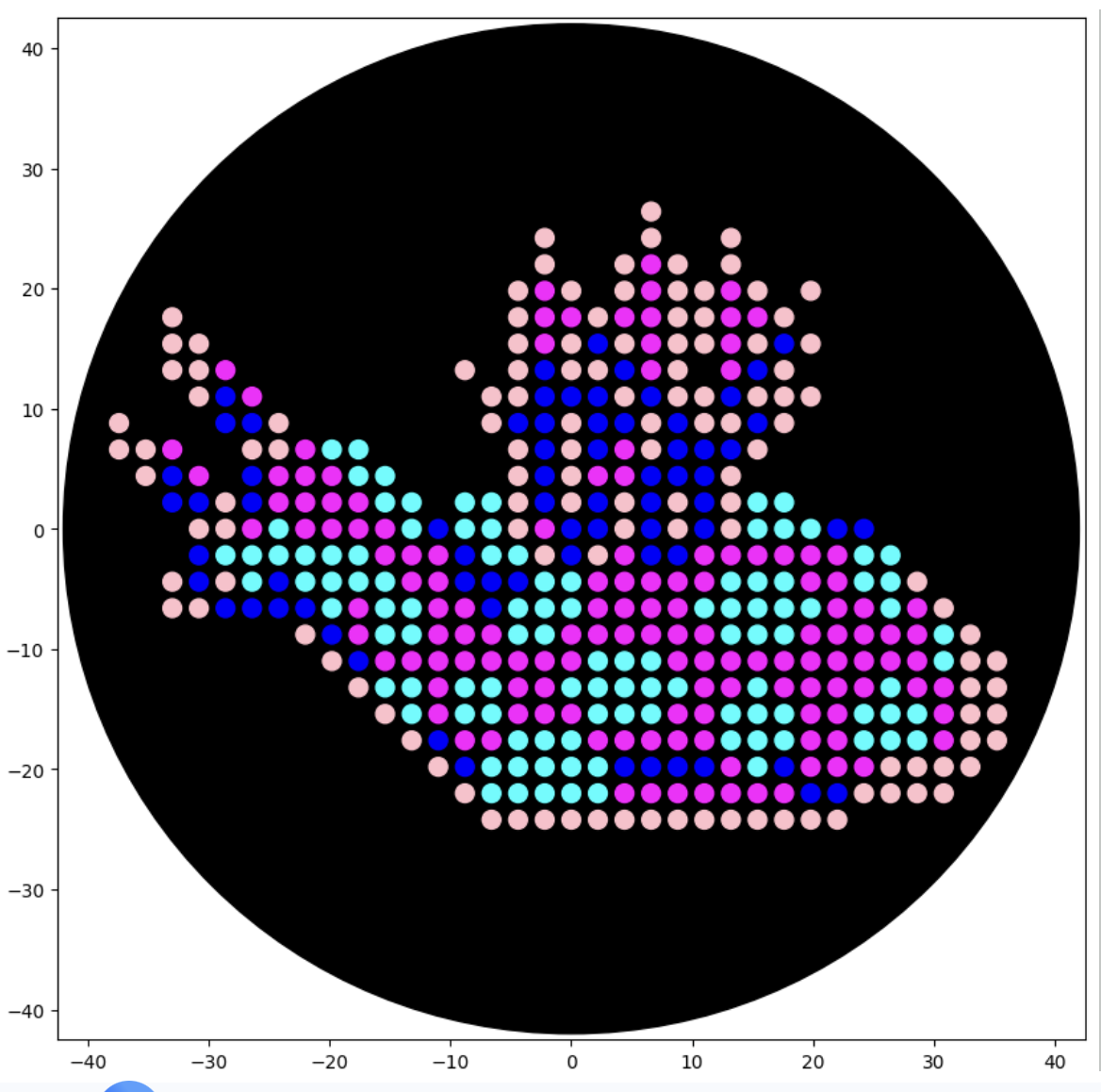
After following the instructions, I downloaded the Python script from the GUI website to insert it into my Google Drive code. The code seemed to be all correct, but I encountered an error when it came to the proteins available on the GUI website and the actual colours in the code. I therefore used the Gemini function available in Google Colab to debug my code when it couldn’t recognise some of the colours. Gemini helped identify missing arguments in the load_labware function and suggested remapping custom protein names to standard colours for visualisation. This is why my final image colours don't match those on the OpenTrons art website.
Final image:

Post Lab-Questions
1) Find and describe a published paper that utilises the Opentrons or an automation tool to achieve novel biological applications.
I found a 2025 paper named 'Real‑time AI‑driven quality control for laboratory automation: a novel computer vision solution for the opentrons OT‑2 liquid handling robot'that was
I found a 2025 paper named: Real‑time AI‑driven quality control for laboratory automation: a novel computer vision solution for the Opentrons OT‑2 liquid handling robot (Khan et al., 2025), which I found particularly interesting.
This paper explores a novel AI-driven computer vision model called YOLOv8 (object detection model), created for the enhancement of quality that limits the accuracy and reliability of liquid handling robots like Opentrons OT-2. Combining these two systems would allow for the precise detection of pippette tips as well as liquid volumes, consequently providing real-time feedback on potential errors like incorrect placement or the absence of pipette tips as well as liquid levels. The paper highlights that the results obtained with their model present an effective, accessible and affordable solution for the improvement of laboratory automation for both academic and research laboratories.
In the introduction, the authors discuss the advantages ( enhancing reproducibility, efficiency and safety) but also disadvantages (high costs, protocol variability and limited expertise) of automation as well as how using it in life sciences remains limited compared to its high success rates in other industries like manufacturing and food production. The paper, therefore, addresses this gap through their YOLOv8 model in combination with the Opentrons OT-2 liquid handling robot.
The paper continues into the Methodology section, which was divided into 4 sections:
The following are the experimental results and analysis, where they began by evaluating the YOLOv8 performance and concluding that the trend indicates the model had progressively enhanced its ability to recognise and, in turn, categorise objects more accurately throughout the training. They continued by looking at the readings from the detected tips and liquids by testing the YOLOv8 system through the conduction of 50 experiments. The experiments consisted of running the OT-2 robot in real time, and intentionall exclude certain pipette tips. The tests resulted in a 98% accuracy of identifying missing tips and their exact location. In addition, the authors tested the measurement of liquids within the pipette tips by conducting 100 experiments using a variety of liquid quantities. Their results showed a 95% accuracy in the assessment of liquid within the tips, which suggested that it was an effective approach but that there is still room for improvement. The authors suggest the implementation of liquid segmentation techniques to do so.
In the discussion section, the authors mainly highlight the potential of their technology as an affordable alternative to costly proprietary solutions and how it aids in the improvement of the quality of produced results, which consequently increases the throughput of the laboratory.
Lastly, in the conclusion and future direction section, the paper describes the potential of the YOLOv8 coupled with OT-2 in the advamncement in life science laboratory automation. The authors also state the limitations of their study, such as their experiment focusing on specific pipette types and liquid volumes. They suggest that future works should address these by expanding the dataset to include a wider range of experimental conditions as well as additional liquid handling tasks. Furthermore, they aim to further develop the YOLOv8 AI computer to detect challenges such as air bubbles, which also affect liquid handling accuracy.
Paper Reference:
Khan, S.U., Møller, V.K., Frandsen, R.J.N. and Mansourvar, M. (2025). Real-time AI-driven quality control for laboratory automation: a novel computer vision solution for the Opentrons OT-2 liquid handling robot. Applied Intelligence, 55(6). doi:https://doi.org/10.1007/s10489-025-06334-3.
2) Write a description about what you intend to do with automation tools for your final project.
For my final project (if I continue the direction of my initial project proposition), I would use a Python script to automate E. coli transformation for screening Psilocybe-derived psiH pathway variants.(Bryant, 2022)
The automation tools I would need/ want to use apart from Python:
As a biodesign student, I want to combine industrial automation (OT-2) with hands-on fabrication challenges. The 3D modelling is a new territory for me, but pushing these design boundaries alongside molecular automation is core to biodesign practice. Additionally, since lab access through my node is uncertain and equipment availability unknown, I'll prioritise developing the Python script and 3D printed components, as these I can prototype independently while planning for potential laboratory access.
Sources