Week 2 HW: DNA, Read, and Write
Part 1
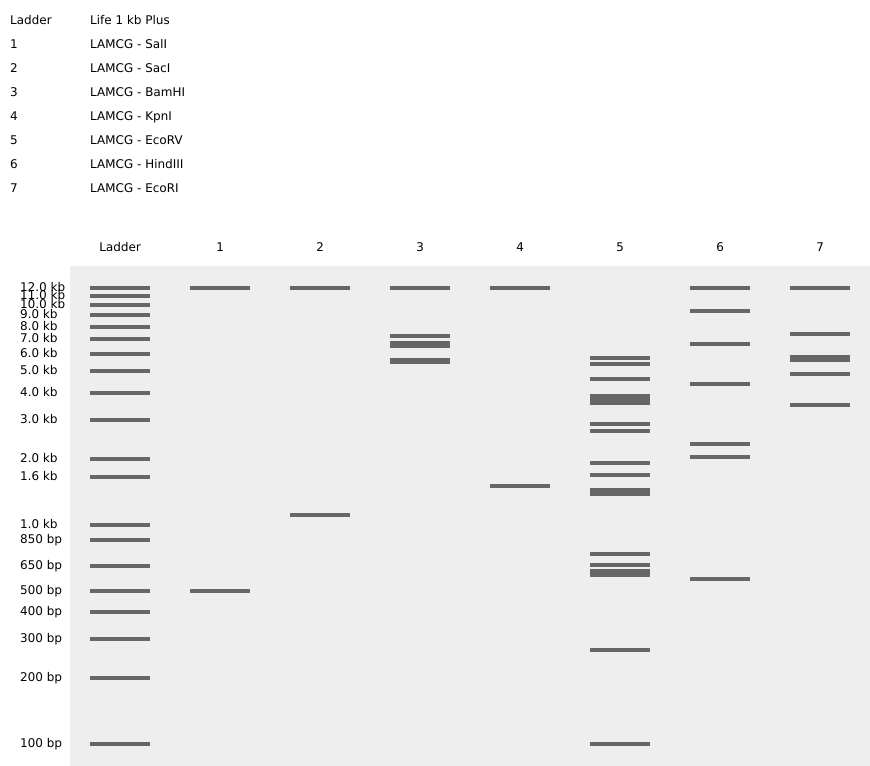



Part 3
3.1. My chosen protein: Green Fluorescent Protein (GFP)
I chose Green Fluorescent Protein (GFP) originally discovered in the jellyfish Aequorea victoria. I chose this protein because:
- It is one of the most important tools in modern biotechnology.
- It glows bright green under blue/UV light.
- Scientists use it as a reporter protein to see when genes are turned on inside living cells.
- It lets researchers literally watch biology happen.
I retrieved the sequence using UniProt (a protein database).
Example FASTA-style entry:
3.2 Reverse Translate:
ATGAGTAAAGGAGAAGAACTTTTCACTGGAGTTGTCCCAATTCTGGTTGAACTGGACGGCGATGTTAACGGCCACAAATTCAGTGTCTCAGGAGAA This DNA sequence would produce the beginning of the GFP protein.
3.3 codon optimisation
codon optimisation is important in synthetic biology because different organisms prefer different codons even if they code for the same amino acid. This is called codon bias.
For example:
Both code for glycine:
- GGT → commonly used in bacteria
- GGG → rarely used in bacteria
Codon optimization rewrites a gene using codons preferred by the host organism so its ribosomes can efficiently translate the mRNA, leading to faster translation and higher protein yield.
Organism chosen:
I optimised the gene for Escherichia coli (E. coli) because:
- It is the most common laboratory host
- Cheap and fast growing
- Standard organism for protein production
- Used in insulin production
After optimization, the DNA sequence changes but the amino acid sequence stays identical.
I optimized the gene for Escherichia coli (E. coli). I chose E. coli because it is the most commonly used organism for recombinant protein expression in laboratories and biotechnology. It grows quickly, is inexpensive to culture, and its genetics are well understood. Because it is widely used for producing proteins such as insulin and research enzymes, optimizing the codons for E. coli increases the likelihood that the protein will be successfully expressed at high levels.
3.4 Next Steps After Sequence:
After obtaining and codon-optimizing the DNA sequence, the protein can be produced using recombinant protein expression technologies. The DNA does not automatically become a protein — it must first be inserted into a system that contains the cellular machinery needed for transcription and translation.
Two major approaches can be used: cell-dependent expression** and **cell-free expression.
Cell-dependent protein production (inside living cells)
The most common method is to express the gene inside bacteria such as E. coli using recombinant DNA technology.
Step 1: Gene synthesis and cloning
The optimized DNA sequence is chemically synthesized and inserted into a circular DNA vector called a plasmid.
The plasmid contains:
- a promoter (turns the gene on)
- a ribosome binding site
- the protein coding sequence
- a terminator
- an antibiotic resistance marker (to select cells that received the plasmid)
This process is called molecular cloning.
Step 2: Transformation
The plasmid is introduced into bacterial cells in a process called transformation (heat shock or electroporation). Some bacteria take up the plasmid and now carry the new gene.
Step 3: Transcription
Inside the bacteria, RNA polymerase binds to the promoter and copies the DNA sequence into messenger RNA (mRNA).
DNA:
mRNA:
(Thymine (T) is replaced with Uracil (U))
Step 4: Translation
A ribosome attaches to the mRNA and reads it in 3-nucleotide codons. Transfer RNAs (tRNAs) bring amino acids that match each codon. The ribosome links the amino acids together into a growing polypeptide chain.
Example:
- AUG → Methionine (start)
- GAA → Glutamate
- UUU → Phenylalanine
The chain continues to elongate until a stop codon is reached.
Step 5: Folding and protein formation
After translation, the amino acid chain folds into its 3-dimensional structure. Once folded correctly, it becomes a functional protein. The bacteria now produce large quantities of the protein, which can be purified.
Cell-free protein production (in a test tube)
Instead of living cells, the protein can also be produced using a cell-free expression system. In this method, cellular machinery extracted from cells (ribosomes, enzymes, tRNAs, amino acids, and nucleotides) is mixed in a reaction tube.
When the synthesized DNA is added:
- RNA polymerase transcribes DNA → mRNA
- Ribosomes translate mRNA → protein
This system essentially recreates the central dogma outside a living organism.
Advantages:
- faster (hours instead of overnight growth)
- no need to maintain living genetically modified organisms
- useful for rapid prototyping proteins
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
I would sequence DNA from the slime mould Physarum polycephalum, specifically genes involved in its oscillatory signalling, environmental sensing, and network formation behaviour. Rather than sequencing the entire genome, I would focus on candidate gene families related to calcium signalling, cytoskeletal contraction, and chemo-sensing receptors, which are believed to underlie the organism’s ability to dynamically reorganise its network in response to environmental conditions.
The motivation is connected to my project: I am developing a biohybrid interface in which the organism participates in spatial decision-making for ecological design. Currently the system treats the slime mould as a black box: it grows, moves, and produces electrical oscillations that are interpreted externally. Sequencing specific regions of its DNA would help identify the biological mechanisms that allow the organism to detect gradients such as moisture, nutrients, and chemical repellents.
Understanding these genes would allow me to distinguish between:
- behaviours that reflect environmental sensing
- behaviours that are internal metabolic rhythms
This matters because the device relies on interpreting the organism’s behaviour as ecological information. If I can identify the genetic pathways associated with sensing versus internal physiological cycles, I can better calibrate which outputs meaningfully correspond to environmental variables (such as soil quality or resource distribution) and which are unrelated background activity.
More broadly, the sequencing would support a longer-term goal: developing a biological chassis for ecological sensing. Rather than engineering the organism immediately, the first step is reading and mapping the genetic basis of its distributed computation. This would establish whether the organism could, in the future, be tuned to respond more specifically to environmental variables (for example pollutants, salinity, or nutrient availability) and therefore act as a living environmental sensor and participatory planning interface.
In this way, the DNA sequencing is not only basic biological curiosity. It is a foundational step toward understanding how a non-neural organism performs spatial computation and whether its sensing behaviour can be responsibly interpreted within civic ecological design systems.
ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why? Also answer the following questions: Is your method first-, second- or third-generation or other? How so? What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps. What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)? What is the output of your chosen sequencing technology?
hybrid sequencing strategy: Nanopore (long reads) + Illumina (accurate short reads).
I would use a hybrid sequencing approach combining Oxford Nanopore long-read sequencing and Illumina sequencing-by-synthesis.
The reason is that Physarum polycephalum has a large and repetitive genome and complex gene regulation. Long-read sequencing allows reconstruction of full genes and regulatory regions, while short-read sequencing provides higher base accuracy. Using both allows discovery of candidate sensing and oscillatory signaling genes that may underlie the organism’s network-forming computation.
Nanopore sequencing helps assemble the genome structure, while Illumina sequencing helps confirm base accuracy and detect smaller mutations or gene variants.
Is your method first-, second-, or third-generation?
- Illumina sequencing-by-synthesis → Second-generation sequencing
- massively parallel, short accurate reads
- Oxford Nanopore sequencing → Third-generation sequencing
- single-molecule, long reads without amplification
Second-generation sequencing reads many short fragments simultaneously, while third-generation sequencing reads individual DNA molecules directly as they pass through a pore.
What is your input and how do you prepare it?
Input
Genomic DNA extracted from cultured slime mould plasmodium.
Because Physarum is multinucleate, the sample contains many nuclei within one cell mass, so I would isolate and purify high-molecular-weight DNA to preserve long fragments.
Preparation steps (library preparation)
1. Cell lysis
- Break open plasmodium cells
- Release nuclei and DNA
2. DNA purification
- Remove proteins, lipids, and RNA
- Keep long DNA strands intact
3. Fragmentation
- For Illumina: shear DNA into short fragments (~200–500 bp)
- For Nanopore: keep DNA long (no fragmentation or minimal shearing)
4. Adapter ligation
- Attach synthetic DNA adapters to fragment ends
- These allow the sequencing machine to recognize and bind the DNA
5. (Illumina only) PCR amplification
- Copy fragments to create enough signal for imaging
6. Load library onto sequencer
Illumina requires amplified libraries attached to a surface, while nanopore sequencing reads native single molecules directly.
Essential sequencing steps & base-calling
A. Illumina (Sequencing-by-Synthesis)
How it works:
- DNA fragments bind to a flow cell
- Fragments form clusters by bridge amplification
- Fluorescently labeled nucleotides are added one at a time
- A camera records the color signal after each cycle
- The color identifies A, T, C, or G
Each nucleotide has a different fluorescent dye, and the sequence is determined by tracking colour changes during repeated cycles.
Base calling:
Colour detected → nucleotide identity.
B. Nanopore Sequencing
How it works:
- A motor protein feeds single-stranded DNA through a nanopore protein embedded in a membrane
- An electric current flows through the pore
- Each base changes the current in a characteristic way
- Software interprets the signal
As DNA passes through the pore, each nucleotide causes a distinct disruption in ionic current, which is converted into sequence information.
Base calling:
Electrical signal pattern → nucleotide identity.
What is the output?
Illumina output
- Millions of short reads (100–300 base pairs)
- Very accurate
- FASTQ files containing:
- sequence
- quality score per base
Nanopore output
- Long reads (10,000–1,000,000+ base pairs)
- Lower accuracy but reveals gene structure
- FASTQ signal-derived sequences
Why this matters for my slime-mould project
The goal is to understand how biological computation happens.
Nanopore reads would reveal:
- gene clusters
- regulatory regions
- large signaling genes
Illumina reads would confirm:
- mutations
- receptor proteins
- ion channel genes
Together, the sequencing flow is: genetic pathways → cellular oscillations → spatial decision-making behaviour.
This directly supports my biohybrid living computation interface.
5.2 DNA Write
(i) What DNA would you want to synthesize and why?
I would design and synthesize a reporter gene circuit that makes cellular signaling activity in Physarum polycephalum visible and measurable. Specifically, I would synthesize a construct that couples an activity-responsive promoter to a fluorescent or luminescent reporter gene.
The goal is not to immediately reprogram the slime mould, but to make its internal computational state observable. In my project the organism functions as a spatial decision-making system, but currently it behaves as a black box: we only see network growth or electrical oscillations after they occur. A genetic reporter would allow the organism to directly communicate its internal signaling state.
The specific biological process I want to observe is the calcium-dependent oscillatory signaling that drives cytoplasmic streaming and network reorganisation. These rhythmic contractions are believed to be the mechanism underlying the organism’s path-finding and resource distribution behavior.
Therefore I would synthesize a calcium-responsive reporter construct.
Proposed genetic construct
The DNA I would synthesise is a simple eukaryotic expression cassette composed of:
[Calcium-responsive promoter] → [reporter protein gene] → [terminator]
Function:
When intracellular signaling activity increases, the organism produces a detectable signal (light or fluorescence). This turns slime-mould computation into a readable biological output rather than requiring electrodes.
This makes the organism not just a growth-based computer, but a living sensing display.
Reporter gene choice
I would synthesize a codon-optimized reporter such as:
GFP (Green Fluorescent Protein)
or
Luciferase
Reason:
These reporters are widely used because they convert gene expression into visible output. The lecture slides emphasised how synthetic DNA enables applications across research, medicine, materials, and sensing. My project falls under biological sensing: instead of detecting disease, the construct detects the organism’s internal signaling activity.
Example DNA sequence (simplified coding region)
Below is a shortened GFP coding sequence example (not the full-length gene, but representative of what would be synthesised):
(For an actual synthesis, the sequence would be codon-optimised for Physarum expression.)
Why this construct matters
Currently, slime-mould computing requires:
- cameras
- electrodes
- external interpretation
A genetic reporter changes the relationship:
Instead of humans interpreting the organism, the organism can directly express its internal state.
This enables:
- visualizing decision points
- mapping signaling waves
- correlating gene activity with spatial behavior
The result: the slime mould becomes a biological interface, not just a biological substrate.
Future extension
Once validated, additional constructs could be synthesized, such as:
- pollutant-responsive promoters
- moisture-responsive expression
- nutrient sensing circuits
This would allow the organism to function as a living environmental sensor rather than a passive computational material.
Why synthesis is necessary
As discussed in the DNA synthesis lecture, synthetic DNA allows researchers to design specific sequences and functions rather than relying on naturally occurring genes slides-lecture-2-leproust. In this case, synthesis enables building a custom interface between cellular signaling and human observation — a foundational step toward a programmable biohybrid ecological computing system.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
To synthesize the reporter construct, I would use solid-phase chemical DNA synthesis (phosphoramidite synthesis) combined with enzymatic gene assembly.
Rather than copying DNA from an organism, this method builds DNA from individual nucleotides in a programmable way. Synthetic DNA platforms can manufacture custom genes designed in software, which is necessary because my calcium-responsive reporter construct does not naturally exist in Physarum.
Modern synthetic DNA platforms fabricate many oligonucleotides simultaneously using microarray-based synthesis and then assemble them into a full gene. Synthetic DNA tools allow researchers to design specific sequences and produce genes for applications in research, sensing, therapeutics, and materials.
I chose this technology because:
- I am designing a new genetic circuit
- there is no natural template to clone
- the gene must be codon-optimized and modular
- the DNA must be written not copied
What are the essential steps of the chosen synthesis method?
1. Oligonucleotide chemical synthesis (base-by-base writing)
The gene is first broken into short fragments (~150–300 bp oligos). Each oligo is chemically synthesized on a solid support surface.
Solid-phase phosphoramidite synthesis proceeds in repeating chemical cycles:
- Attach first nucleotide to solid surface
- Add a protected nucleotide phosphoramidite
- Coupling reaction attaches base
- Cap unreacted strands
- Oxidize phosphate backbone
- Remove protecting group (deblock)
- Repeat cycle
These steps are repeated many times to build a sequence base-by-base, to essentially make a chemical writing of DNA.
2. Cleavage and purification
After synthesis, the oligos are:
- released from the surface
- chemically deprotected
- purified
3. Gene assembly
The short oligos are then assembled into a full gene:
- overlapping sequences are designed
- fragments anneal
- DNA polymerase fills gaps
- PCR amplifies the full gene
classical gene synthesis is assembling many short oligos into a full gene through PCR-based assembly
4. Cloning and sequence verification
The final gene is:
- inserted into a plasmid vector
- propagated in bacteria
- sequence-verified (often using next-generation sequencing)
What are the limitations of this synthesis method?
Accuracy limitations
Chemical DNA synthesis is not perfect. Errors occur because each chemical addition step is slightly inefficient.
Typical issues:
- deletion mutations
- substitution errors
- incomplete strands
As sequences get longer, cumulative error rates increase. Therefore longer genes must be assembled from shorter verified fragments.
Length limitations
Single oligos cannot be extremely long.
Typical constraints:
- individual oligos: ~150–300 nucleotides
- genes: assembled from many oligos
Highly repetitive DNA, extreme GC content, or hairpin structures are especially difficult to synthesize
Speed limitations
Gene synthesis is not instantaneous because it involves:
- chemical cycles
- purification
- assembly
- verification sequencing
However, modern platforms dramatically improve speed and scale; thousands of oligos can be produced simultaneously using array-based synthesis
Scalability trade-offs
Strength:
- extremely scalable (many genes at once)
Weakness:
- each individual gene still requires assembly and verification
- long custom constructs take longer than short fragments
Why this method fits the project
The purpose of my project is to create a new biological interface. Because the reporter circuit is a designed genetic construct rather than a naturally occurring gene, the appropriate technology is synthetic DNA manufacturing. Chemical oligo synthesis combined with gene assembly allows precise control over promoter, coding region, and regulatory elements, enabling the slime mould to express a visible signal corresponding to its internal computational state.
This moves the organism from being observed externally to being able to communicate biologically.
5.3 DNA Edit
(i) What DNA would you want to edit and why?
I would edit genes in the slime mould Physarum polycephalum that control its oscillatory signaling and environmental sensing behavior. My goal is not to fundamentally redesign the organism, but to make its computational behavior more interpretable and experimentally controllable.
The organism’s path-finding and network formation depend on rhythmic cytoplasmic streaming driven by intracellular calcium signaling and cytoskeletal contraction. These oscillations determine how the organism distributes resources and selects paths in response to gradients such as nutrients, moisture, and repellents. In my project, the slime mould functions as a biohybrid spatial decision-making system for ecological design, but currently its behavior can only be influenced indirectly using light, salt, or food placement.
I would therefore edit genes involved in three biological functions:
1. Editing sensory receptors (environmental responsiveness)
I would modify or insert chemosensory receptor genes so that the organism responds to specific environmental variables rather than only simple attractants (e.g., oats) or repellents (e.g., salt).
Goal:
Enable the slime mould to detect meaningful environmental conditions such as:
- soil salinity
- pollutants
- nutrient concentration
- moisture stress
Why:
Currently, the organism reacts to arbitrary laboratory proxies. By editing sensing pathways, its decisions could reflect real ecological signals instead of human-selected stimuli. This would allow the biohybrid system to function as a living environmental sensor.
Type of edit:
Targeted insertion of a sensing pathway promoter or receptor gene under native regulatory control.
2. Editing oscillatory signaling genes (control of computation speed)
The slime mould’s decision-making depends on rhythmic contraction waves. I would edit genes regulating calcium ion channels or actin-myosin cytoskeletal contraction to slightly alter oscillation frequency.
Goal:
Adjust how quickly the organism explores and stabilizes networks.
Why:
Currently, experiments can take many hours or days because the organism’s internal clock sets its computational speed. Modifying the oscillation period could make the system experimentally usable without changing its overall behavior.
Type of edit:
Regulatory modification (promoter tuning) rather than gene knockout adjusting expression levels rather than removing function.
3. Editing reporter expression (internal state visibility)
In addition to writing a reporter gene, I would edit its genomic insertion site so the reporter is expressed only during active signaling events rather than continuously.
Goal:
Allow the organism to visibly signal when it is “making a decision” (i.e., undergoing active network reorganization).
Why:
Right now the organism’s internal state is invisible unless measured electrically. A genomic integration tied to a signaling pathway would allow the organism to communicate its activity biologically.
Type of edit:
Knock-in insertion at a signaling pathway locus (for example, a calcium-responsive gene).
Why editing rather than only writing DNA
The write step adds new functions, but editing is necessary to integrate them into the organism’s native regulatory system. Without editing, the reporter gene would operate independently of the biological processes responsible for computation.
Editing allows:
- linking sensing → signaling → behavior
- reducing reliance on external interpretation
- making the organism a participant rather than an experimental object
Broader implications
The intention is not to optimise the organism for efficiency or industrial use, but to explore a new relationship between biological systems and human decision-making. The edit would allow the slime mould to mediate between environmental conditions and human planning processes, acting as a biological interface.
However, this also raises ethical considerations: editing a living organism to participate in civic decision systems changes how agency and responsibility are distributed. For that reason, I would prioritise reversible and minimal edits (regulatory changes rather than permanent loss-of-function mutations) and maintain contained laboratory use.
I would use DNA editing to make the slime mould not a programmable machine, but a legible ecological collaborator, an organism whose internal biological processes can be meaningfully interpreted rather than inferred indirectly.
(ii) What technology would you use and why?
I would use CRISPR-Cas9 genome editing, combined with homology-directed repair (HDR) and, in some cases, CRISPR activation (CRISPRa).
I chose CRISPR because it allows targeted, programmable editing of specific genes rather than random mutation. My project requires modifying signaling and sensing pathways in Physarum polycephalum while preserving the organism’s viability and behavior. CRISPR enables precise insertion of reporter genes and fine control of gene regulation, which is necessary for connecting biological signaling to observable outputs.
CRISPR is appropriate because I am not trying to create a new organism, but to:
- insert a reporter gene
- tune expression of signaling pathways
- minimally perturb existing functions
How does CRISPR edit DNA?
CRISPR works by using a programmable RNA molecule to guide a nuclease enzyme (Cas9) to a specific DNA sequence.
Essential mechanism
- A guide RNA (gRNA) is designed to match a target DNA sequence.
- The Cas9 protein binds the guide RNA.
- The complex scans DNA in the cell.
- When the matching sequence is found, Cas9 cuts the DNA (double-strand break).
- The cell repairs the break.
The repair process is what produces the edit.
There are two repair pathways:
1. Non-homologous end joining (NHEJ)
- error-prone repair
- produces insertions or deletions
- used for gene knockouts
2. Homology-directed repair (HDR)
- precise repair using a template
- allows insertion of new DNA
For my project I would mainly use HDR, because I want to insert a reporter gene and modify regulatory regions rather than destroy genes.
Essential experimental steps
- Identify target gene (e.g., calcium signaling gene)
- Design guide RNA sequence
- Construct editing plasmid
- Deliver CRISPR system into slime mould cells
- Cas9 cuts DNA at target site
- Cell repairs DNA using provided template
- Screen for successful edits
What preparation is required?
Design steps
- Choose a target gene locus
- Design guide RNA (20 bp complementary sequence)
- Design repair template with homology arms (~500–1000 bp)
- Insert reporter gene into template DNA
Inputs needed
Biological components
- Cas9 nuclease protein (or gene encoding it)
- guide RNA (gRNA)
- donor DNA repair template (containing reporter gene)
- promoter and terminator sequences
Delivery materials
- plasmid vector OR ribonucleoprotein complex (Cas9 + gRNA)
- transformation method (e.g., electroporation or microinjection)
Cells
- cultured Physarum polycephalum plasmodium
What is CRISPRa and why use it?
In addition to cutting DNA, a modified Cas9 (dead Cas9 or dCas9) can regulate genes without breaking DNA.
dCas9:
- binds DNA
- does not cut
If fused to an activator protein, it increases expression of a gene.
I would use CRISPRa to:
- adjust oscillation frequency
- tune sensing sensitivity
This avoids permanent genome disruption.
Limitations of CRISPR editing
1. Efficiency
Not every cell receives the edit.
Possible issues:
- low transformation efficiency
- multinucleate cells (especially relevant in Physarum)
- mosaic editing
Result: many cells must be screened.
2. Off-target effects
The guide RNA may bind similar DNA sequences and cut unintended locations.
Consequences:
- unintended mutations
- altered behavior
Careful guide design reduces this risk.
3. Repair pathway bias
Cells often prefer NHEJ instead of HDR.
This means:
- insertions may fail
- knockouts occur instead of precise edits
HDR is typically less efficient than gene disruption.
4. Precision limitations
Even successful edits may:
- integrate partially
- rearrange DNA
- produce variable expression
Therefore sequencing verification is required after editing.
Why this editing method fits the project
My project depends on linking biological signaling to visible outputs. CRISPR allows insertion of a reporter gene into a native signaling pathway and fine-tuning of sensing behavior. This makes the organism experimentally interpretable without fundamentally redesigning its biology.
Sequencing lets me understand the organism, synthesis lets me add a component, and CRISPR lets me connect the component to the organism’s natural processes.