Subsections of Homework
Week 1 HW: Principles and Practices

- First, describe a biological engineering application or tool you want to develop and why.
I want to develop a highly efficient bacterial chassis for rapid intracellular biosynthesis of novel PHA (polyhydroxyalkanoate) copolymers. More than 150 different hydroxyalkanoate monomers have been identified, and they can be combined into co-polymers (and potentially ter-/quad-polymers) with variable composition and sequence/microstructure, leading to an astronomical design space.
My goal is to build a chassis that makes the design–build–test loop fast and reliable: (1) use computational/AI approaches and literature review to prioritize promising copolymer compositions for target properties, then (2) rapidly prototype biosynthesis in a standardized host with predictable performance, and (3) generate experimental data to validate predicted properties and improve models.
This tool idea is directly inspired by my work: I’m the CEO of Bioplastix, a biotech startup with the mission to accelerate the transition to biodegradable bioplastics. We already have a promissing co-polymer (PLA-PHB with different proportions of PLA) and a highly efficient E.coli chassis to produce it. We want to accelerate the discovery of new co-polymers. A chassis that can quickly produce and screen new PHA copolymers would let us create radically better biopolymers and accelerate transition to bioplastics.
- Next, describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future, like ensuring non-malfeasance (preventing harm). Break big goals down into two or more specific sub-goals.
The goal of this biological tool is to contribute to an ethical future by reducing harm to human and planetary health through the replacement of oil-based plastics. Conventional plastics accumulate in ecosystems and human bodies (Nihart, A.J., Garcia, M.A., El Hayek, E. et al. Bioaccumulation of microplastics in decedent human brains. Nat Med 31, 1114–1119 (2025)), have high associated carbon footprints, and often rely on toxic ingredients across their life cycle.
For this tool to be successful, I suggest two core policy and governance goals:
Goal 1: Encourage the adoption of bio-based and biodegradable plastics.
At present, biodegradable bioplastics such as PHAs remain significantly more expensive than oil-based plastics (e.g., PHAs at approximately 4–5 USD/kg versus PET at ~1.3 USD/kg). Policies that reduce this gap during early scale-up phases are therefore essential to enable meaningful market penetration.
Goal 2: Prevent harm from new biological strains and novel polymer compositions.
Because this platform enables the rapid creation of many new copolymers and engineered microbial strains, governance is critical to avoid unintended biological or environmental consequences. Like any powerful biological technology, it must operate within clearly defined biosafety, material safety, and ethical boundaries.
Note: I could state Goal 1 as: Ban traditional plastics or sidcourage the use of traditinal plastics. A global ban or restrictions on conventional plastics could, in principle, accelerate this transition, but such an outcome appears unlikely in the near term, as illustrated by the limited progress of the UN Global Plastics Treaty negotiations.
- Next, describe at least three different potential governance “actions” by considering four aspects (Purpose, Design, Assumptions, Risks of Failure & “Success”).
Action 1: Tax incentives for biodegradable and bio-based plastics
Purpose: Today, in most countries and subnational jurisdictions, there are no meaningful tax incentives that favor biodegradable bioplastics over conventional fossil-based plastics. This action proposes targeted tax reductions for products made predominantly from certified biodegradable plastics, with the goal of narrowing the price gap between products made with conventional plastics and those made with biodegradable alternatives.
Design: This action would require: Legislation establishing tax reduction and scope. Clear regulation and enforcement, since many environmental laws fail at the implementation stage, including: i. eligibility criteria for tax incentives (e.g., minimum biodegradable content, verified biodegradability standards). ii. auditing mechanisms to prevent misuse, such as products labeled “bioplastic” that are bio-based but not biodegradable, or products containing only a small fraction of biodegradable material. iii. phased implementation, potentially starting with high-impact applications where replacing conventional plastics yields the greatest environmental benefit.
Key actors include national and/or subnational governments, tax authorities, certification bodies, manufacturers opting into the program, and end users opting into buying products.
Assumptions: This action assumes that: Governments are willing and able to implement and enforce differentiated tax schemes. Traditional plastics resin producers will not lobby enough so as to stop the law. Price reductions at the product level are sufficient to meaningfully influence purchasing and adoption decisions. Certification systems can accurately distinguish between genuinely biodegradable materials and greenwashed alternatives.
Risks of Failure & “Success”: The policy could fail if enforcement is weak, allowing non-compliant products to benefit from incentives, or if administrative complexity discourages participation. Jurisdictional differences in taxation could also lead to production shifting across borders rather than reducing overall plastic harm. Even “successful” implementation may have unintended consequences, such as encouraging overconsumption of disposable products simply because they are labeled biodegradable, rather than reducing total plastic use.
Action 2: Large-scale public awareness campaigns on the human health impacts of traditional plastics
Purpose: Currently, public discourse around plastics focuses primarily on environmental damage, while the human health impacts of conventional plastics—such as chemical exposure, bioaccumulation, and endocrine disruption—remain relatively under-communicated. This limits public pressure for change and weakens demand for safer alternatives. This action proposes coordinated awareness campaigns that frame conventional plastics as a public health issue, similar to past campaigns against tobacco or excessive sugar consumption.
Design: This action would require: Global actors (e.g., WHO, UN agencies, international NGOs) to support and legitimize messaging at a global scale. National and local governments to adapt campaigns to local contexts and regulatory priorities. Startups and research institutions working on plastic alternatives to contribute evidence-based narratives and real-world solutions. Influencers, educators, and media organizations to amplify messages beyond traditional policy channels.
Assumptions: This action assumes that Increased awareness of health risks will meaningfully influence consumer behavior and political support.
Also, that influencers and media actors can communicate complex health information responsibly. That global and local actins can be integrated.
Risks of Failure & “Success”: The campaign could fail if messages are oversimplified, sensationalized, or perceived as fear-based, leading to public distrust. There is also a risk of backlash from industry actors framing the campaign as anti-innovation or anti-consumer or leading to jobs reduction.
Action 3: Establish dedicated biopolymer research and governance centers at national and global levels
Purpose: At present, governance and reaseadrh and development of new biopolymers and engineered production strains is fragmented across regulatory agencies, academic labs, and industry actors. This fragmentation slows safe innovation and creates uncertainty around implementation, standards, and scale-up. This action proposes the creation of dedicated research and governance centers at national or subnational levels, complemented by a global network coordinating best practices and knowledge sharing.
Design: These centers would: i. Support regulatory implementation, including biosafety and material safety evaluation for new biopolymers.
ii Conduct applied research on performance, degradation, and real-world applications of biodegradable plastics. iii. Serve as interfaces between academia, startups, industry, and regulators. iv. Participate in global networks to harmonize standards, share data, and reduce duplication of effort. Key actors include governments (as funders), universities, public research institutes, and international coordination bodies, as well as industry and startups.
Assumptions: This action assumes that: Governments are willing to fund long-term, interdisciplinary centers rather than short-term projects.
Centralized expertise improves both safety and innovation outcomes. International collaboration is feasible despite differences in regulation and economic priorities.
Risks of Failure & “Success”: These centers could fail by becoming overly bureaucratic or disconnected from real industrial needs. They may also privilege dominant technological pathways, limiting diversity in approaches. A “successful” network might unintentionally centralize decision-making power, creating gatekeepers that slow innovation or disadvantage smaller players without access to these institutions.
- Next, score (from 1-3 with, 1 as the best, or n/a) each of your governance actions against your rubric of policy goals.
| Does the option: | Action 1 | Action 2 | Action 3 |
|---|
| G1: Encourage the adoption of bioplastics | | | |
| • By discouraging oil-based plastics | 2 | 1 | 3 |
| • By reducing the price gap with oil.based | 1 | n/a | 3 |
| • By encouraging innovation | 1 | 3 | 2 |
| • By improving the implementation of laws | 1 | 3 | 2 |
| G2: Prevent NEW harm | | | |
| • By preventing incidents at lab scale | 3 | n/a | 1 |
| • By preventing environmental consequences | 2 | 3 | 1 |
| • By preventing human health consequences | 2 | 3 | 1 |
| Overarching GOAL: Protect the environment | | | |
| • By preventing plastics in ecosistems | 1 | 2 | 3 |
| • By preventing GHG emmissions | 1 | 2 | 3 |
| Overarching GOAL: Protect human health | | | |
| • By preventing plastics in bodies | 1 | 2 | 3 |
| • By reducing the use of toxic ingredients | 1 | 2 | 3 |
| • By preventing GHG emmissions | 1 | 2 | 3 |
- Last, drawing upon this scoring, describe which governance option, or combination of options, you would prioritize, and why. Outline any trade-offs you considered as well as assumptions and uncertainties.
Based on the expected impact and feasibility of the proposed actions, I would prioritize Action 1: tax incentives for biodegradable and bio-based plastics as the primary governance intervention.
The main reason is that cost remains the dominant barrier to large-scale adoption of biodegradable plastics, despite their availability and well-documented environmental benefits. Awareness campaigns (Action 2) and research centers (Action 3) already exist to some degree, and bioplastics are commercially available today. What has not been broadly or consistently implemented is a policy mechanism that directly and effectively reduces the price gap between fossil-based plastics and biodegradable alternatives through governmental incentives.
For a national or subnational policymaker audience (e.g., ministries of industry, environment, or finance), Action 1 offers the highest near-term leverage. In contrast, awareness campaigns rely on slower cultural change, and research centers operate on longer innovation timelines.
Action 3—the creation or reorientation of dedicated biopolymer research and governance centers—would be a strong second priority because will be important to enhance laws. Importantly, this does not necessarily require creating entirely new institutions. Existing research centers could be re-scoped through regulation or funding requirements.
Action 2 is expected to emerge organically once Action 1 is implemented. As economic incentives shift markets, industry groups, startups, researchers, and civil society actors are likely to amplify awareness efforts and public communication, especially around human health impacts. In this sense, Action 1 can act as a catalyst for the other two actions.
Trade-offs, assumptions, and uncertainties: This prioritization assumes that governments are willing to intervene in markets through fiscal policy and that tax incentives will translate into lower end-user prices. There is also uncertainty around political feasibility, especially in jurisdictions resistant to environmental taxation or subsidies.
Week 2 Prep:
Homework Questions from Professor Jacobson:
Error Rate: 1:106 Throughput: 10 mS per Base Addition
basic error rate of DNA polymerase without proofreading is roughly 1 error per 10² to 10⁶ bases
Biology resolves this discrepancy through multiple layers of fidelity control, including polymerase proofreading, mismatch repair systems, and cell-cycle checkpoints that prevent propagation of damaged DNA.
Average Human Protein: 1036 bp • Longest Human Proteins (PKS): >100kbp
Since most amino acids are encoded by multiple synonymous codons, the number of possible DNA sequences that could encode the same protein is astronomically large.
Not all work due to biological constrains: Codon usage bias: different organisms preferentially use certain synonymous codons, which affects translation efficiency and protein yield. mRNA structure: different nucleotide sequences form different local structures that affect ribosome binding and stability. Embedded regulatory elements: coding regions can contain splice sites, RNA-binding motifs, or secondary signals that influence expression.
Homework Questions from Dr. LeProust:
phosphoramidite DNA synthesis.
it is difficult to synthesize oligos longer than ~200 nt
Each nucleotide addition step has a small but non-zero error rate (incomplete coupling, deletions, or side reactions). As oligo length increases, these errors accumulate multiplicatively, leading to a rapidly decreasing fraction of full-length, error-free molecules. Beyond ~150–200 nt, yield and fidelity drop sharply, making purification inefficient and expensive.
A 2000 bp sequence would require ~2000 sequential synthesis steps, which would result in near-zero yield of full-length correct molecules due to accumulated errors. Instead, long genes are built by assembling shorter oligos (e.g., 50–200 nt) using enzymatic methods such as PCR-based assembly or Gibson assembly
Homework Question from George Church:
1.
Histidine
Isoleucine
Leucine
Lysine
Methionine
Phenylalanine
Threonine
Tryptophan
Valine
lysine is universally essential across animal biology (not unique to any engineered or extinct species), to prevent organisms from surviving without an external lysine supply is not biologically plausible or unique—it simply reflects an existing nutritional requirement. This makes the fictional Lysine Contingency concept not a realistic.
Note on AI use:
A large language model (ChatGPT) was used solely to assist with English grammar, spelling, and clarity of writing. All ideas, goals, policy actions, and arguments presented in this assignment were developed independently by the author and not generated by the language model.
Week 11 HW: Bioproduction & Cloud Labs
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
- Each component’s role is in the cell-free reaction.
- E. coli Lysate (BL21 (DE3) Star): Provides the essential biological “hardware,” including ribosomes, tRNAs, and various translation factors; it specifically includes T7 RNA Polymerase to drive high-level transcription from T7 promoters.
Salts and Buffers:
- Potassium Glutamate: Acts as the primary potassium source and a major intracellular salt, which is critical for maintaining proper osmotic pressure and supporting protein-DNA interactions during transcription and translation.
- HEPES-KOH pH 7.5: Serves as a chemical buffering agent to maintain a stable physiological pH throughout the reaction, preventing acidification from metabolic byproducts.
- Magnesium Glutamate: Provides essential $Mg^{2+}$ ions that act as necessary cofactors for ribosome assembly and the enzymatic activity of polymerases.
- Potassium Phosphate (Monobasic & Dibasic): Maintains phosphate homeostasis and contributes to pH stability, while also providing inorganic phosphate for nucleotide recycling.
Energy and Nucleotide System:
- Ribose and Glucose: Serve as secondary energy substrates that can be metabolized by residual glycolytic enzymes in the lysate to regenerate ATP sustainably.
- AMP, CMP, GMP, UMP, and Guanine: These act as the fundamental precursors (nucleotides and bases) for mRNA synthesis during transcription and are necessary components for maintaining the energy charge of the system.
Translation Mix (Amino Acids):
- 17 Amino Acid Mix, Tyrosine, and Cysteine: Provide the raw building blocks required for the ribosome to assemble the polypeptide chain; Tyrosine and Cysteine are often added separately due to their lower solubility at neutral pH.
Additives and Backfill:
- Nicotinamide: Often acts as a cofactor or stabilizer to inhibit the degradation of essential metabolic intermediates like NAD+, thereby extending the reaction’s metabolic activity.
- Nuclease Free Water: Used to adjust the final volume (backfill) of the reaction while ensuring no contaminating enzymes degrade the DNA template or mRNA products.
- Describe the main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the Google Slide above. (2-3 sentences)
The 1-hour PEP-NTP mix is designed for high-speed, short-term bursts of protein synthesis by providing direct, high-energy building blocks like Nucleoside Triphosphates (NTPs) and a potent phosphagen (PEP) for immediate ATP regeneration. In contrast, the 20-hour NMP-Ribose-Glucose mix is optimized for long-term production and cost-efficiency; it utilizes cheaper precursors (Nucleoside Monophosphates) and a dual-sugar metabolic pathway (Ribose/Glucose) to regenerate energy slowly and steadily over several hours, preventing the rapid accumulation of inhibitory byproducts.
- Bonus question: How can transcription occur if GMP is not included but Guanine is?
Transcription can occur even if GMP is not explicitly included because the E. coli lysate contains residual metabolic enzymes (such as Phosphoribosyltransferases) that can utilize Guanine as a substrate. Through a “salvage pathway,” the system attaches a ribose-5-phosphate to the Guanine base to synthesize GMP. Once GMP is formed, kinases in the lysate further phosphorylate it into GDP and finally GTP, which is the actual nucleotide required by the RNA Polymerase to build the mRNA chain.
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
- Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each)
Functional Properties of Fluorescent Proteins
sfGFP (Superfolder GFP): This protein is engineered for extremely rapid folding and high stability, making it highly resistant to aggregation even when expressed at high rates in cell-free extracts.
mRFP1 (monomeric Red Fluorescent Protein): A key limitation is its relatively slow maturation time and lower photostability, which often results in a delayed fluorescence signal compared to green variants in short-term reactions.
mKO2 (monomeric Kusabira Orange 2): This protein features high pH sensitivity (acid sensitivity); its fluorescence can be significantly quenched if the cell-free reaction undergoes acidification due to metabolic byproduct accumulation.
mTurquoise2: Known for its exceptional quantum yield and brightness, it requires very precise folding conditions to achieve its maximum fluorescence intensity in a cyan readout.
mScarlet-I: This is a high-performance red protein that is notably oxygen-dependent for the final step of chromophore maturation, meaning insufficient aeration in the reaction vessel can limit its readout.
Electra2: Designed for enhanced photostability and rapid folding, this protein provides a very fast readout, though it can be sensitive to the ionic strength of the buffer (specifically magnesium levels) during the initial translation phase.
- Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
Protein: mKO2 (monomeric Kusabira Orange 2)
Reagents to Adjust: HEPES-KOH (Buffer) and Potassium Phosphate.
Hypothesis: Increasing the concentration of HEPES-KOH and Potassium Phosphate in the master mix will improve the fluorescence of mKO2 by enhancing the system’s buffering capacity.
Expected Effect: In a long 36-hour incubation, the cell-free reaction typically produces organic acids as byproducts of glucose and ribose metabolism. Since mKO2 is acid-sensitive, a drop in pH would normally quench its signal mid-incubation. By strengthening the buffer, we can maintain the pH near 7.5 for the entire duration, preventing the quenching of the chromophore and maximizing the cumulative fluorescence readout.
Week 12 HW: Building Genomes
Week 2 HW: DNA Read, Write, and Edit

Part 1: Benchling & In-silico Gel Art 🦠
Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks:
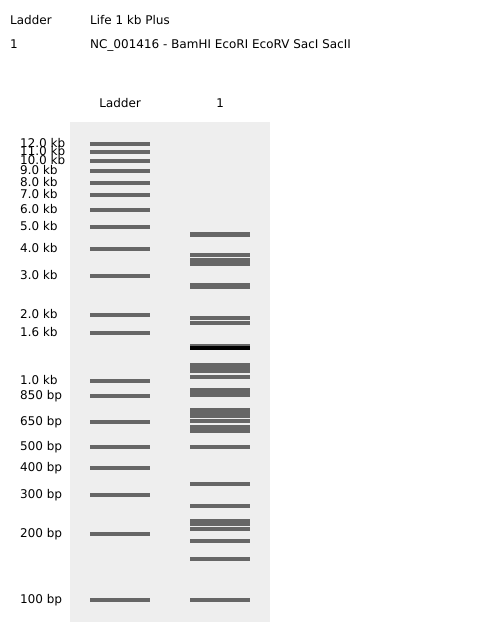

Part 3: DNA Design Challenge
I chose poly(3-hydroxyalkanoate) polymerase / PHB synthase (PhaC) from Cupriavidus necator (UniProt accession P23608) because it is a key enzyme in microbial bioplastic production. PhaC catalyzes the polymerization of (R)-3-hydroxybutyryl-CoA monomers to form poly(3-hydroxybutyrate) (PHB), and engineered variants of PhaC are widely used to broaden substrate specificity and produce other polyhydroxyalkanoates (PHAs). I obtained the amino-acid sequence from UniProt (entry P23608) in FASTA format.
MA T G K G A A A S T Q E G K S Q P F K V T P G P F D P A T W L E W S R Q W Q G T E G N G H A A A S G I P G L D A L A G V K I A P A Q L G D I Q Q R Y M K D F S A L W Q A M A E G K A E A T G P L H D R R F A G D A W R T N L P Y R F A A A F Y L L N A R A L T E L A D A V E A D A K T R Q R I R F A I S Q W V D A M S P A N F L A T N P E A Q R L L I E S G G E S L R A G V R N M M E D L T R G K I S Q T D E S A F E V G R N V A V T E G A V V F E N E Y F Q L L Q Y K P L T D K V H A R P L L M V P P C I N K Y Y I L D L Q P E S S L V R H V V E Q G H T V F L V S W R N P D A S M A G S T W D D Y I E H A A I R A I E V A R D I S G Q D K I N V L G F C V G G T I V S T A L A V L A A R G E H P A A S V T L L T T L L D F A D T G I L D V F V D E G H V Q L R E A T L G G G A G A P C A A A A G L E L A N T F S F L R P N D L V W N Y V V D N Y L K G N T P V P F D L L F W N G D A T N L P G P W Y C W Y L R H T L P A E R A Q G T G Q A D R V R R A G G P G Q H R R P Y I Y G S R E D H I V P W T A A Y A S T A L L A N K L R F V L G A S G H I A G V I N P P A K N K R S H W T N D A L P E S P Q Q W L A G A I E H H G S W W P D W T A W L A G Q A G A K R A R P A N Y G N A R Y R A I E P A P G D T S K P R H
Although the genetic code is universal, most amino acids are encoded by multiple codons. However, different organisms do not use these synonymous codons at the same frequency. This phenomenon is known as codon bias. If a gene from one organism is expressed in a different host, the original codons may be rare in the new host, which can reduce translation efficiency, among other problems.
Codon optimization is necessary to modify the DNA sequence so that it uses codons preferred by the host organism. Optimizing codon usage can significantly increase protein expression levels, improve translation efficiency, enhance mRNA stability, and reduce the likelihood of misfolding or premature termination.
For this project, I chose to optimize the phaC gene for Escherichia coli. I selected E. coli because it is the chassis organism we use at Bioplastix.
I used https://www.genscript.com/ as a code optimizattion tool (Job ID: 20260216015729685186).
At Bioplastix, the engineered bacteria produce the enzymes intracellularly. Under specific metabolic conditions, these enzymes catalyze the polymerization of monomers inside the cell, leading to intracellular accumulation of the biopolymer.
To achieve that, once the codon-optimized DNA sequence is obtained, it is inserted into a plasmid (an expression vector containing the necessary regulatory elements, such as a promoter and terminator). This plasmid is then be inserted into the host organism, Escherichia coli, for protein production.
In such a cell-dependent system, the DNA sequence is transcribed into messenger RNA (mRNA) by RNA polymerase. The mRNA is then translated by ribosomes into the PhaC protein. Depending on the design of the genetic construct, the gene can be placed under a constitutive promoter, where the protein is continuously produced, or under an inducible promoter, where expression is triggered by specific conditions (such as the presence of IPTG or oxigen).
Alternatively, the protein could also be produced using a cell-free expression system, where the DNA is added to a reaction mixture containing ribosomes, tRNAs, nucleotides, and enzymes necessary for transcription and translation. This allows protein production without living cells, although large-scale industrial production until now typically relies on cell-based systems.
Part 4: Prepare a Twist DNA Synthesis Order
Link Sharing From Benchling: https://benchling.com/s/seq-l6sbaCHItMO0QD5lHD4M?m=slm-38MASpmHAGVSi3hySqfU
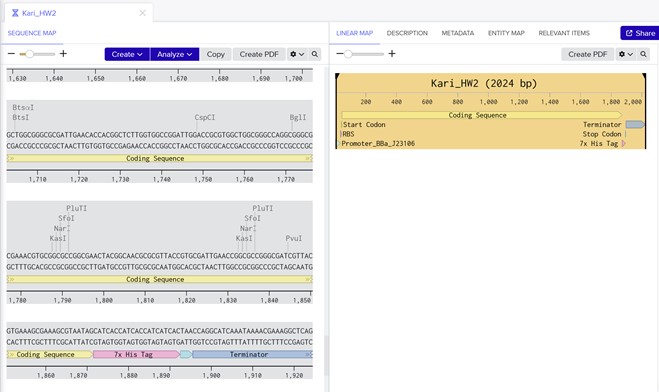

Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
I would like to sequence DNA from environmental microbial communities, particularly from diverse and extreme environments such as soil, marine ecosystems, open dumps, and high-stress habitats. These environments often contain microorganisms with unique metabolic capabilities.
By sequencing DNA from environmental samples (metagenomic sequencing), we could identify novel enzymes involved in polymer biosynthesis, including new variants of PHA synthases or related polymerizing enzymes. These enzymes may have improved catalytic efficiency, altered substrate specificity, or greater stability under industrial conditions. Discovering new enzymes through DNA sequencing could enable the development of more efficient bioplastic production systems and potentially allow the synthesis of novel polymers with enhanced material properties.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
I would use Illumina sequencing technology, a second-generation (next-generation sequencing, NGS) platform. Illumina sequencing is well-suited for metagenomic analysis because it provides high-throughput, highly accurate short reads at relatively low cost, making it ideal for sequencing complex microbial communities. It is second generation because it performs massively parallel sequencing, requires DNA amplification (cluster generation), and sequences millions of fragments simultaneously.
The input is extracted environmental DNA (metagenomic DNA) from microbial communities.
Sample preparation steps:
- DNA extraction from environmental samples (soil, water, etc.)
- Fragmentation of DNA into smaller pieces (typically ~200–500 bp)
- End repair and A-tailing
- Adapter ligation (short known sequences attached to fragment ends)
- PCR amplification to enrich properly ligated fragments
- Library quantification and quality control
Illumina sequencing works as follows:
- The prepared DNA library is loaded onto a flow cell.
- Fragments bind to complementary oligos attached to the flow cell.
- Bridge amplification creates clusters of identical DNA fragments.
- Fluorescently labeled reversible terminator nucleotides are added.
- During each cycle, one nucleotide is incorporated per cluster.
- A camera detects the fluorescent signal emitted by the incorporated base.
- The terminator is cleaved, and the next cycle begins.
Each nucleotide (A, T, C, G) carries a different fluorescent signal. The imaging system detects the color emitted at each cycle, and software converts fluorescence intensity into base calls (A, T, C, or G).
5.2 DNA Write
The DNA I would want to “write” (synthesize) would be a set of candidate polymerizing enzymes identified from the environmental metagenomic library, especially novel PHA synthase (PhaC-like) genes or other enzymes predicted to catalyze polymer formation. The main reason to synthesize these genes is to rapidly test them in a standardized host. I would order codon-optimized versions of each candidate enzyme gene for Escherichia coli, because E. coli is the chassis organism we use at Bioplastix.
I would use solid-phase phosphoramidite DNA synthesis, the standard chemical DNA synthesis technology used by companies such as Twist Bioscience.
In the past we already used Twist for synthesis of new enzymes. It is very useful because enables codon optimization, Introduction of specific mutations, Removal of unwanted restriction sites, parallel synthesis of multiple variants.
With this methodd DNA is synthesized one nucleotide at a time on a solid support using phosphoramidite chemistry. This produces short DNA fragments (typically 150–200 bp). Since genes are longer than individual oligos, overlapping oligonucleotides are assembled into full-length genes using: PCR-based assembly, Gibson Assembly or Enzymatic ligation methods. The assembled gene is amplified and sometimes enzymatically corrected to reduce synthesis errors. The final construct is cloned into a plasmid and verified by DNA sequencing before delivery.
From a practical perspective at Bioplastix, the main limitation of outsourcing DNA synthesis to companies such as Twist Bioscience is cost. Synthesizing long gene sequences can be expensive, particularly when testing multiple enzyme variants. Each synthesis order costs approximately $1,500 USD. After synthesis, there are further costs associated with Transforming it into our production strains and Screening and validating expression. Overall, with a typical budget, we are able to test approximately 5–10 new enzyme candidates per round.
5.3 DNA Edit
The DNA I would want to edit is the genome of our production strain (e.g., Escherichia coli) in order to improve its efficiency as a bioplastic-producing chassis organism. Specifically, I would focus on :
1️⃣ Expanding substrate utilization: One of our current priorities is enabling the strain to efficiently consume alternative and low-cost carbon sources, particularly sucrose. Editing the genome to introduce or optimize sucrose transporters and metabolic pathways would allow the bacteria to convert a wider range of feedstocks into biopolymer, improving economic feasibility and sustainability.
2️⃣ Increasing carbon flux toward polymer production: I would edit genes involved in central carbon metabolism to redirect more carbon toward the PHA biosynthesis pathway.
3️⃣ Engineering polymerizing enzymes: I would also edit the genes encoding polymerizing enzymes (such as PhaC) to Increase substrate affinity, Improve catalytic efficiency and Enhance thermostability. Thermotolerant enzymes would be particularly valuable at industrial scale, where higher fermentation temperatures reduce cooling costs and contamination risk.
To perform these genome edits, I would use CRISPR-Cas9 genome editing, combined with homologous recombination. CRISPR-based systems are precise, programmable, and highly efficient, making them ideal for metabolic engineering in bacterial systems. CRISPR-Cas9 uses: i. A guide RNA (gRNA) designed to match a specific DNA target sequence. ii. The Cas9 nuclease, which creates a double-strand break at the target site
After the break is introduced, the cell repairs the DNA. If a repair template is provided (homology-directed repair, HDR), specific edits such as insertions, deletions, or point mutations can be introduced.
Limmitations include Metabolic burden (Multiple edits can stress the cell and reduce growth rate) and Regulatory complexity (Metabolic networks are highly interconnected; editing one pathway may produce unexpected downstream effects).
Week 3 HW: Lab Automation

Python Script for Opentrons Artwork: https://colab.research.google.com/drive/14m54uLCM5UtsggVjU2Ucxh5hhtNELWD2#scrollTo=pczDLwsq64mk&line=4&uniqifier=1
- Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
I found the paper ¨Opentrons for automated and high-throughput viscometry¨ very interesting.
¨The operating protocol involves measuring the amount of liquid dispensed over a set time for given dispense conditions. Data collected at different set dispense flow rates was used to train an ensemble machine learning regressor to predict Newtonian liquid viscosity¨. They demonstrated the ability of the proxy viscometer to characterize the rheological behavior of two types of power-law fluids.
Write a description about what you intend to do with automation tools for your final project.
For my final project, I intend to focus on a challenge that is directly relevant to my startup, Bioplastix, which develops technologies for the production of bio-based copolymers with plastic properties. I am particularly interested in exploring how low-cost automation tools, such as the Opentrons OT-2, can accelerate enzyme discovery and biopolymer development workflows. I have two potential project directions:
Idea 1: High-Throughput Screening of Polymerizing Enzymes: Bioplastix has access to a diverse environmental enzyme library. Our goal is to identify enzymes capable of polymerizing specific monomers into useful copolymers. An initial in silico screening step can be performed by identifying genetic sequences that are homologous to known polymerizing enzymes. However, the key bottleneck occurs during experimental validation in the wet lab. Using automation tools, we could perform parallel screening of up to 96 enzyme candidates per plate, standardizing reaction setup to reduce variability.
This will be possible Implementing colorimetric assays to detect polymer formation. The automation platform would allow us to screen dozens (or hundreds) of enzymes under consistent conditions, significantly accelerating discovery while reducing human error.
Idea 2: Automated Cell-Free Production of Copolymers: A second direction would be to explore automated workflows for cell-free production systems. Instead of expressing enzymes in living cells, we could use cell-free systems to produce polymers. This would involve screen different reaction conditions (pH, cofactors, substrate ratios, temperature), optimize copolymer production in a high-throughput format.
Week 4 HW: Protein Design - Part 1
Part A. Conceptual Questions
- How many molecules of amino acids do you take with a piece of 500 grams of meat? (on average an amino acid is ~100 Daltons)
Assumptions: Meat ≈ 20% protein. 500 g meat → ~100 g protein. Average amino acid residue mass ≈ 100 Da ≈ 100 g/mol.
So: 100 g ÷ 100 g/mol ≈ 1 mole of amino acid residues. 1 mole = 6.022 × 10²³ molecules.
👉 You ingest on the order of 6 × 10²³ amino acid residues in 500 g of meat.
- Why do humans eat beef but do not become a cow, eat fish but do not become fish?
Because digestion destroys biological structure. Proteins are hydrolyzed into amino acids. The original sequences (information) are lost. Your body reassembles amino acids according to human DNA instructions. Biological identity is encoded in sequence and genomic regulation — not in the raw amino acid building blocks. You absorb matter, not identity.
Note: While reading this response, I was prompted to ask ChatGPT the following question:
“Do the 20 amino acids explain all the DNA of all species? Are there more evolved species that use more amino acids?”
ChatGPT’s response was: Yes, the same 20 amino acids (with two rare exceptions) account for virtually all proteins in all known living organisms.
There are no “more evolved” species that use a greater number of standard amino acids.
This follow-up helped clarify the universality of the genetic code and reinforced the idea that biological complexity arises from sequence variation and regulation, rather than from an expanded set of amino acid building blocks.
- Why are there only 20 natural amino acids?
This is evolutionary, not chemical necessity. The 20 canonical amino acids: Provide sufficient chemical diversity: Hydrophobic, Polar, Charged, Aromatic, Reactive, Enable folding into complex structures.
Likely reflect optimization of the genetic code to minimize error impact. More than 500 amino acids exist in nature, but only 20 are universally encoded in the genetic code (with rare additions like selenocysteine). Evolution found a minimal but sufficient toolkit.
- Can you make other non-natural amino acids? Design some new amino acids.
Yes. Synthetic biology routinely incorporates non-natural amino acids. Examples: A) Fluorinated phenylalanine. Replace para-H with F. Increased hydrophobicity. Enhanced stability. Altered electronic properties.
B) Long-chain hydrophobic amino acid. Extend leucine side chain by two carbons. Stronger hydrophobic core packing. Enhanced β-sheet stabilization.
- Where did amino acids come from before enzymes that make them, and before life started?
Prebiotic chemistry. Evidence includes: Miller–Urey experiments (electric discharge in reducing atmosphere). Meteorites (e.g., Murchison contains amino acids). Hydrothermal vent synthesis. Interstellar ice chemistry. Amino acids likely formed abiotically before enzymatic systems existed. Life inherited pre-existing chemistry.
- If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
Natural α-helices (L-amino acids) are: 👉 Right-handed. If composed of D-amino acids: 👉 The helix becomes left-handed.
This inversion arises directly from chirality.
- Can you discover additional helices in proteins?
Yes. Beyond α-helices: 3₁₀ helix, π-helix, Polyproline type II helix, Coiled-coils, Collagen triple helix. Protein design has also generated synthetic helices not found in nature. Helical geometry depends on backbone torsion angles and side-chain packing.
- Why are most molecular helices right-handed?
Because life uses L-amino acids. Chirality at the monomer level restricts energetically favorable torsion angles, biasing right-handed helices.
Handedness is an emergent property of stereochemistry.
- Why do β-sheets tend to aggregate? What is the driving force for β-sheet aggregation?
Because β-strands expose: Extended backbone hydrogen bond donors/acceptors. Flat, planar surfaces. They readily align and form intermolecular hydrogen bonds, leading to stacking and fibril formation.
Their geometry promotes self-association.
Primary forces: Backbone hydrogen bonding. Hydrophobic interactions. Aromatic stacking. Water exclusion (hydrophobic effect).
Aggregation often lowers free energy significantly.
- Why do many amyloid diseases form β-sheets? Can you use amyloid β-sheets as materials?
Because partially unfolded proteins expose aggregation-prone segments. These segments reorganize into cross-β structures: Highly ordered. Extremely stable. Resistant to degradation
Examples: Alzheimer’s (Aβ), Parkinson’s (α-synuclein), Prion diseases.
Amyloid represents a misfolded but energetically favorable state.
amyloid β-sheets can be used as materials. Amyloid fibrils have: High tensile strength, Nanofibrillar architecture, Structural robustness. They are being explored for: Hydrogels, Nanowires, Bio-scaffolds, Sustainable biomaterials. The main challenge is controlled assembly.
- Design a β-sheet motif that forms a well-ordered structure.
Design principle: Alternate hydrophobic and polar residues to create amphipathic β-strands.
Example motif: Val–Thr–Val–Thr–Val–Thr–Val–Thr
This yields: One hydrophobic face. One polar face. Stable sheet stacking.
For improved order: Design a β-hairpin: Val–Thr–Val–Thr–Asn–Gly–Val–Thr–Val–Thr
Asn–Gly promotes a tight turn. Alternating residues favor sheet formation. Terminal charged residues can prevent uncontrolled aggregation.
This design promotes controlled, ordered β-sheet assembly.
NOTE: The answers presented in this section were generated using ChatGPT as a starting point and were subsequently reviewed and edited by me. The AI output served as a draft to help structure and articulate the responses. The prompt I used was: “Provide detailed answers to the following questions about amino acids, protein structure, helices, β-sheets, aggregation, amyloids, and biomaterial applications. Answer with scientific depth and clarity.” All final revisions, interpretations, and refinements reflect my own understanding.
Part B: Protein Analysis and Visualization
- Briefly describe the protein you selected and why you selected it.
I selected a PhaC synthase because we use it at Bioplastix to polimerize our co-polymer.
- Identify the amino acid sequence of your protein.
The length of the protein is: 562 aminoacids. The most common amino acid is: L, which appears 67 times.
Using RCSB Sequence Similarity tool. The query matched 24 Polymer Entities, for example:
orgnism: Pseudomonas aeruginosa PAO1
Macromolecule: Poly(3-hydroxyalkanoic acid) synthase 1
Sequence Match: Sequence Identity: 99%, E-Value: 0, Region: 1-559

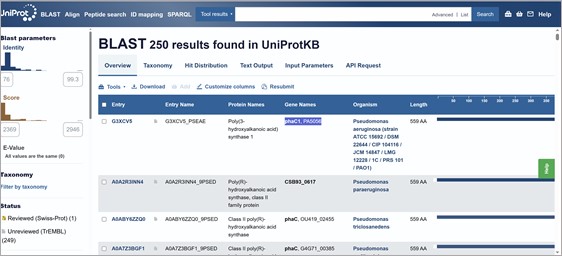
Also, using UniProtKB BLAST 250 results were found (most of them pseudomonas). For example: Poly(3-hydroxyalkanoic acid) synthase 1, phaC1, PA5056. Protein family/group databases: ESTHER pseae-PHAC1PHA_synth_II

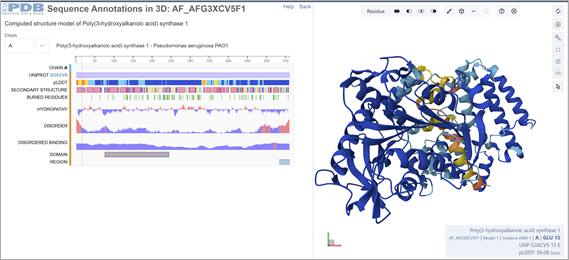
- Identify the structure page of your protein in RCSB
This was solved with a model. ¨COMPUTED STRUCTURE MODEL OF POLY(3-HYDROXYALKANOIC ACID) SYNTHASE 1. There are no experimental data to verify the accuracy of this computed structure model. See Model Confidence metrics below for all regions of the polypeptide chain.
AlphaFold DB: AF-G3XCV5-F1
Released in AlphaFold DB: 2021-12-09
Last Modified in AlphaFold DB: 2022-09-30
Organism(s): Pseudomonas aeruginosa PAO1
UniProtKB: G3XCV5
Model Confidence pLDDT (global): 88.77¨
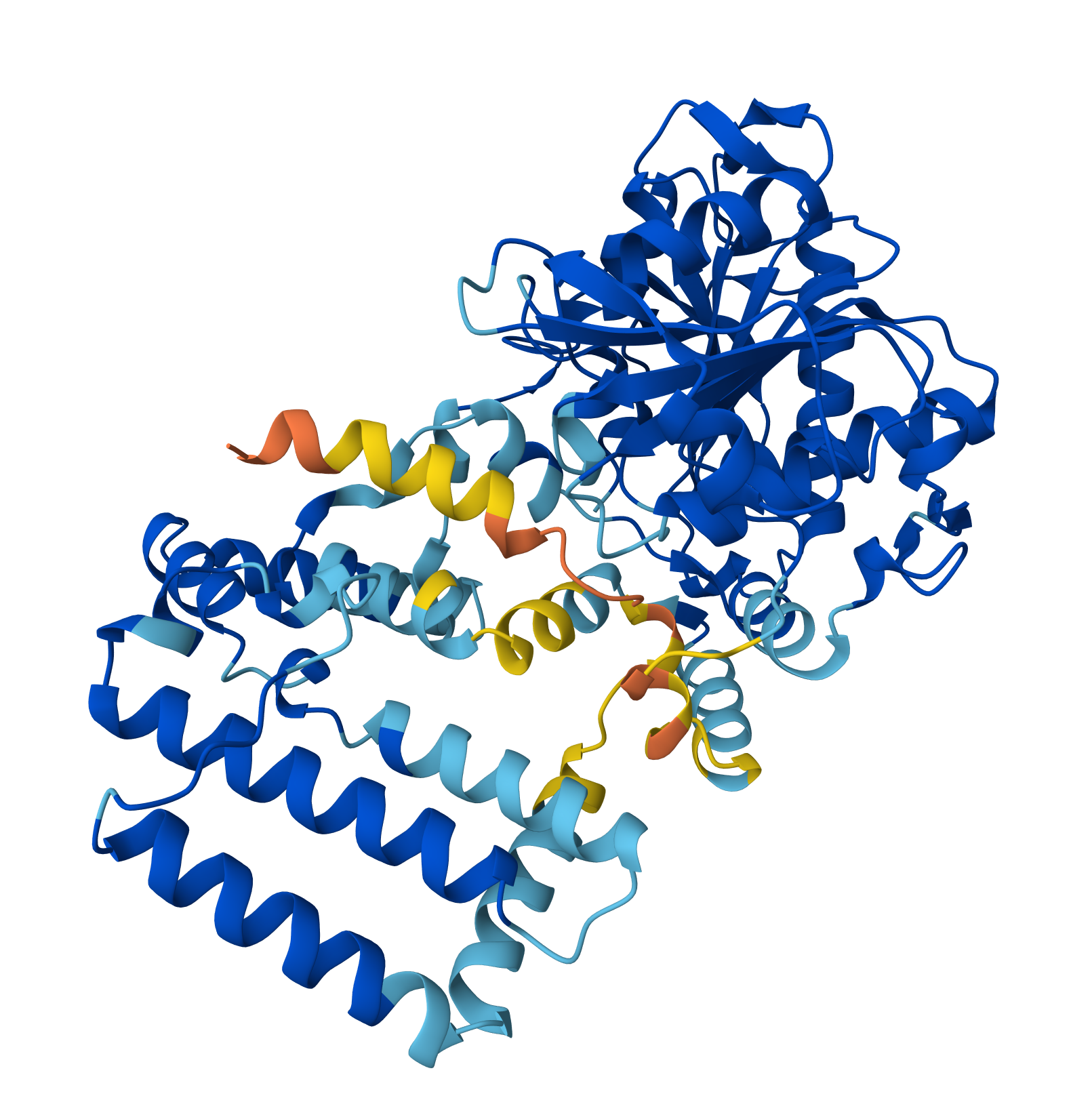

I found a crystal structure for a PhaC with Resolution: 2.70 Å:
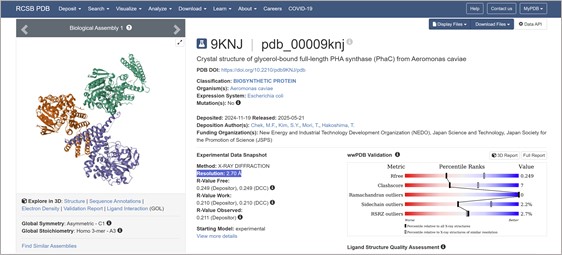

- Open the structure of your protein in any 3D molecule visualization software
Part C: Using ML-Based Protein Design Tools
I selected PHA synthase (PhaC) from Aeromonas caviae.
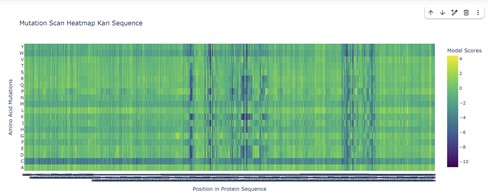

The mutational scanning heatmap highlights residue positions that are highly sensitive to mutation (dark blue/purple columns), likely corresponding to structurally or functionally critical residues. In contrast, regions with predominantly neutral or positive scores represent mutationally tolerant positions that could serve as potential targets for protein engineering.
The mutational heatmap suggests that different regions of the protein show varying tolerance to amino acid substitutions. Several positions display strong intolerance to mutation, visible as vertical dark-blue bands across many amino acid substitutions. These regions likely correspond to structurally or functionally important residues where mutations would strongly destabilize the protein or disrupt its activity.
In contrast, other regions show mostly green or light-colored scores, indicating positions that are more tolerant to mutation. These sites may correspond to surface-exposed residues or flexible loop regions, and could represent promising targets for protein engineering.
Interestingly, substitutions to tryptophan (W) and cysteine (C) tend to produce consistently negative scores across many positions, appearing as predominantly blue rows in the heatmap. This suggests that introducing these residues is generally unfavorable, likely because tryptophan is bulky and cysteine can introduce disulfide bonds or structural instability in inappropriate contexts.
Overall, the heatmap highlights a pattern of heterogeneous mutational tolerance, with conserved regions that are highly sensitive to mutation and more permissive regions that may accommodate sequence variation. These insights can guide rational protein engineering by identifying positions where mutations are more likely to be tolerated without compromising protein folding.

Part D: Group Brainstorm on Bacteriophage Engineering
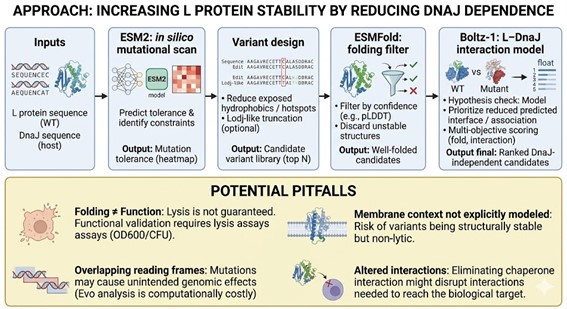
Bacteriophage Engineering Proposal: L Protein Stabilization
Primary Goal: Increased stability (easiest)
Specific Approach: Engineering DnaJ-independence by reducing chaperone-recognition signals while preserving the structural scaffold of the L protein.
Computational Tools and Pipeline Justification To achieve this goal, we propose a three-step computationally efficient pipeline:
Step 1: Sequence-level Mutational Scanning using ESM2
Approach: We will perform a zero-shot in silico mutational scan across the L protein sequence using the ESM2 Protein Language Model (PLM). We aim to identify exposed hydrophobic patches (typical DnaJ recognition motifs) and propose polar/hydrophilic substitutions.
Why this helps: ESM2 has learned deep evolutionary constraints across millions of protein sequences. It allows us to rapidly differentiate between highly constrained residues (which are structurally vital and “untouchable”) and mutation-tolerant positions. This ensures we only disrupt chaperone-binding motifs without breaking the core evolutionary scaffold of the protein, all at a fraction of the computational cost of molecular dynamics.
Step 2: Rapid Structural Filtering using ESMFold
Approach: The top candidate sequences from the ESM2 scan will be predicted using ESMFold. We will filter out any variants that collapse, show low pLDDT (confidence) scores, or have a high RMSD compared to the Wild-Type (WT) backbone.
Why this helps: While ESM2 evaluates sequence-level fitness, we need explicit 3D structural validation. ESMFold is significantly faster than AlphaFold2, making it ideal for high-throughput filtering. This step ensures that our hydrophilic mutations do not inadvertently destroy the L protein’s ability to fold independently.
Step 3: Complex Modeling using Boltz-1
Approach: We will model the L protein + DnaJ complex for both the WT and our top folded mutant candidates. We will analyze the predicted interface contacts and Predicted Aligned Error (PAE) to assess binding affinity.
Why this helps: Folding correctly in isolation is not enough; we must explicitly prove reduced chaperone dependency. By comparing the mutant-DnaJ interface against the WT-DnaJ interface, we can prioritize variants that maintain a stable fold but show a significantly weakened or abolished interaction with the DnaJ chaperone.
Potential Pitfalls
- Pitfall 1: Overlapping Reading Frames and Genomic Constraints. Phage genomes are highly compact, meaning the DNA sequence encoding the L protein might also encode parts of other proteins or regulatory elements in alternative reading frames. Our targeted mutations could have unintended, fatal consequences for the phage’s overall viability. While genomic foundation models like Evo could assess these genome-wide constraints, their computational cost is prohibitive for our current scope.
- Pitfall 2: The Stability vs. Function Trade-off. ESMFold guarantees that the protein adopts a stable 3D conformation in solution, but it does not guarantee biological function (membrane lysis). Lytic activity heavily depends on complex factors like membrane insertion dynamics, oligomerization, and reaction kinetics. Furthermore, completely abolishing chaperone interaction might inadvertently prevent the L protein from being properly delivered to its target membrane.

Week 5 HW: Protein Design - Part 2
Part A. SOD1 Binder Peptide Design
The perplexity scores for the candidate peptides were:
‘SDGAVLLGSDGE’ (Candidate 1): 16.25
‘LLGSDGALQVGS’ (Candidate 2): 14.65
‘SGVAVLCSDGQG’ (Candidate 3): 25.34
‘AVGVCGVAVLGN’ (Candidate 4): 17.20
Lower perplexity scores suggest that the model finds a sequence more “familiar” or “expected,” potentially correlating with higher biological plausibility, conformational stability, or likelihood of interaction. Candidate 2, with a perplexity of 14.65, appears to be the most promising candidate from the mutated SOD1 sequence for further investigation, being closest to the known binding peptide’s score.
Candidate 1: ipTM = 0.52 pTM = 0.88
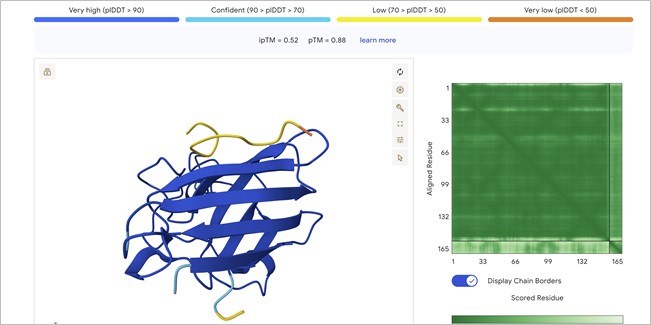

Localization: The peptide (the small chain with yellow and orange segments at the top) is localized near the N-terminus. It is observed to sit atop the beta-barrel, specifically in the region where the initial helix and the N-terminal end of the SOD1 (shown in dark blue) connect with the rest of the protein structure.Interaction with the beta-barrel: The peptide directly engages the $\beta$-barrel region. It appears to be “resting” on the upper beta-sheets, effectively acting as a molecular cap.Surface-bound vs. Buried: The peptide appears primarily surface-bound. While it is not deeply buried within the protein core, it maintains extensive contact with the exposed surface, a characteristic typical of stabilizing peptides.Relationship to A4V: Being situated at the top, near the start of the sequence, the peptide is in the immediate vicinity of residue 4. This suggests it may help “anchor” the N-terminus, preventing it from detaching or unfolding due to the destabilizing effect of the A4V mutation.
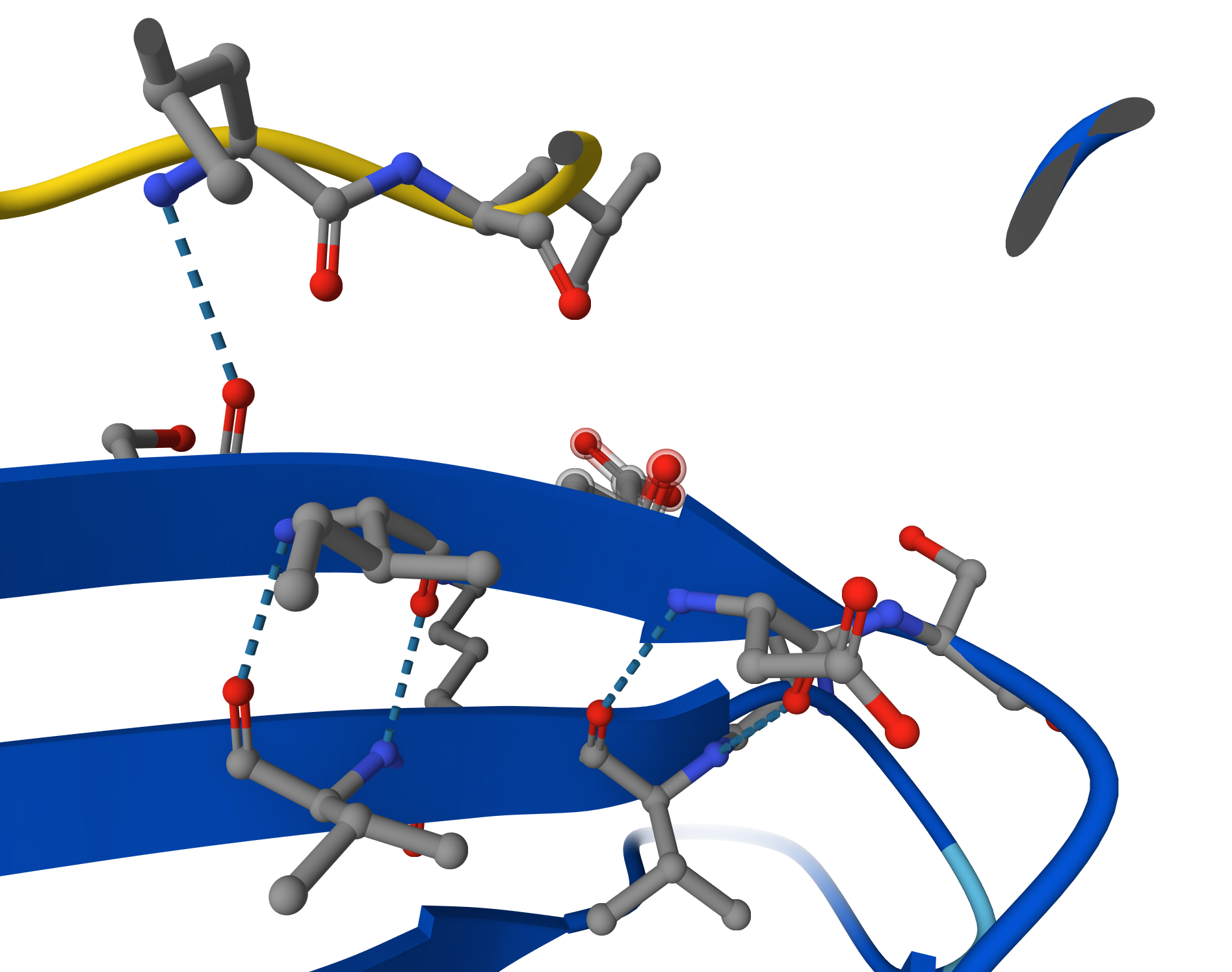

Upon closer inspection of the AlphaFold3 model, a clear hydrogen bond is visible between the peptide backbone and the SOD1 $\beta$-sheet. This specific inter-chain interaction confirms that the peptide is not just near the enzyme, but actively docking through electrostatic stabilization, which contributes to the observed ipTM of 0.52
Candidate 2: ipTM = 0.54 pTM = 0.89
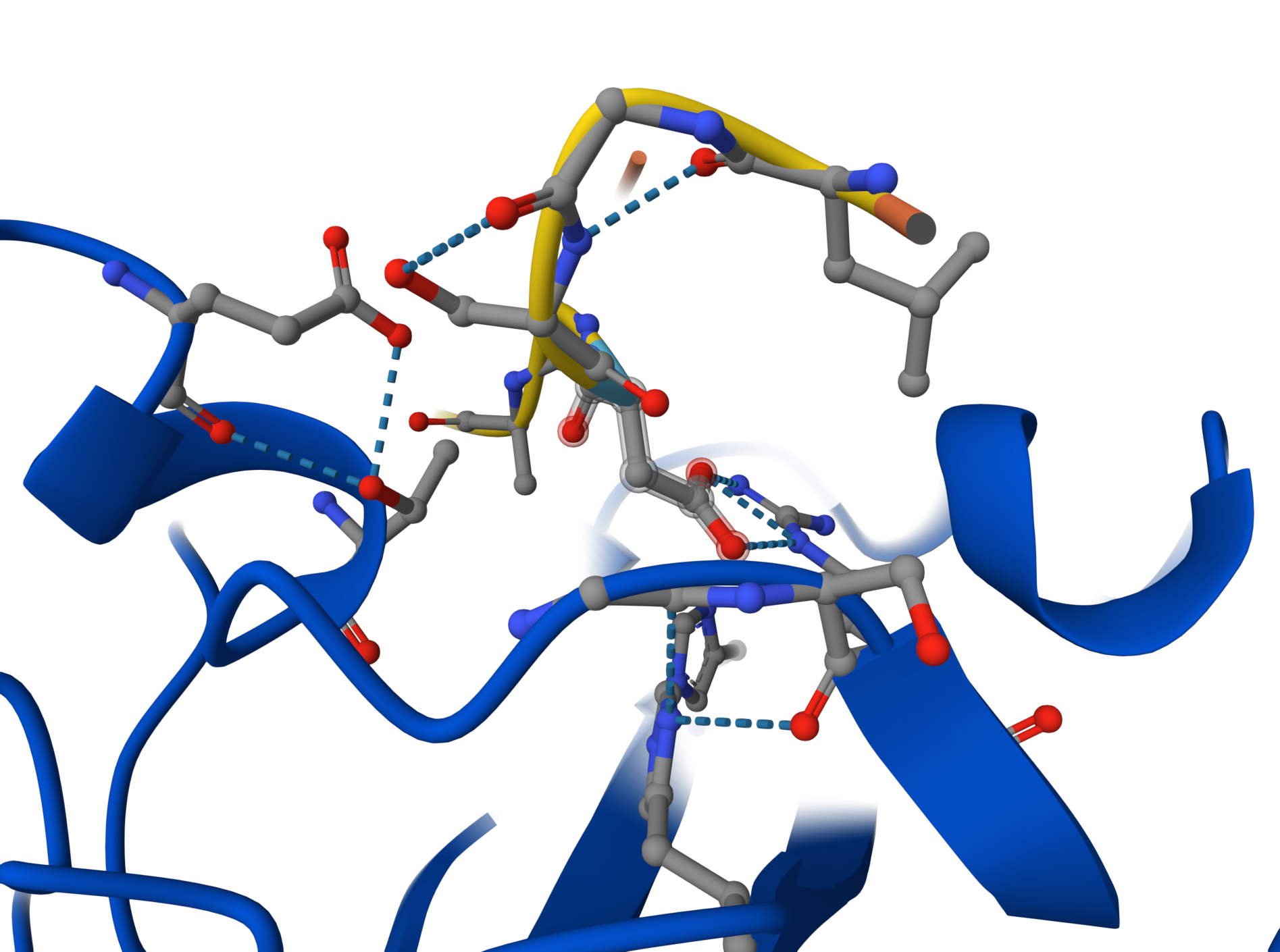

Although Candidate #2 shows a similar confidence score (ipTM = 0.54), the structural model reveals a significantly more robust interaction network. Detailed inspection shows multiple hydrogen bonds between the peptide backbone and the SOD1 surface loops, compared to the single-point attachment of previous candidate. This increased ‘molecular velcro’ effect likely provides better stabilization of the N-terminus, making this peptide a stronger therapeutic lead against the A4V mutation.
Candidate 3: ipTM = 0.35 pTM = 0.88
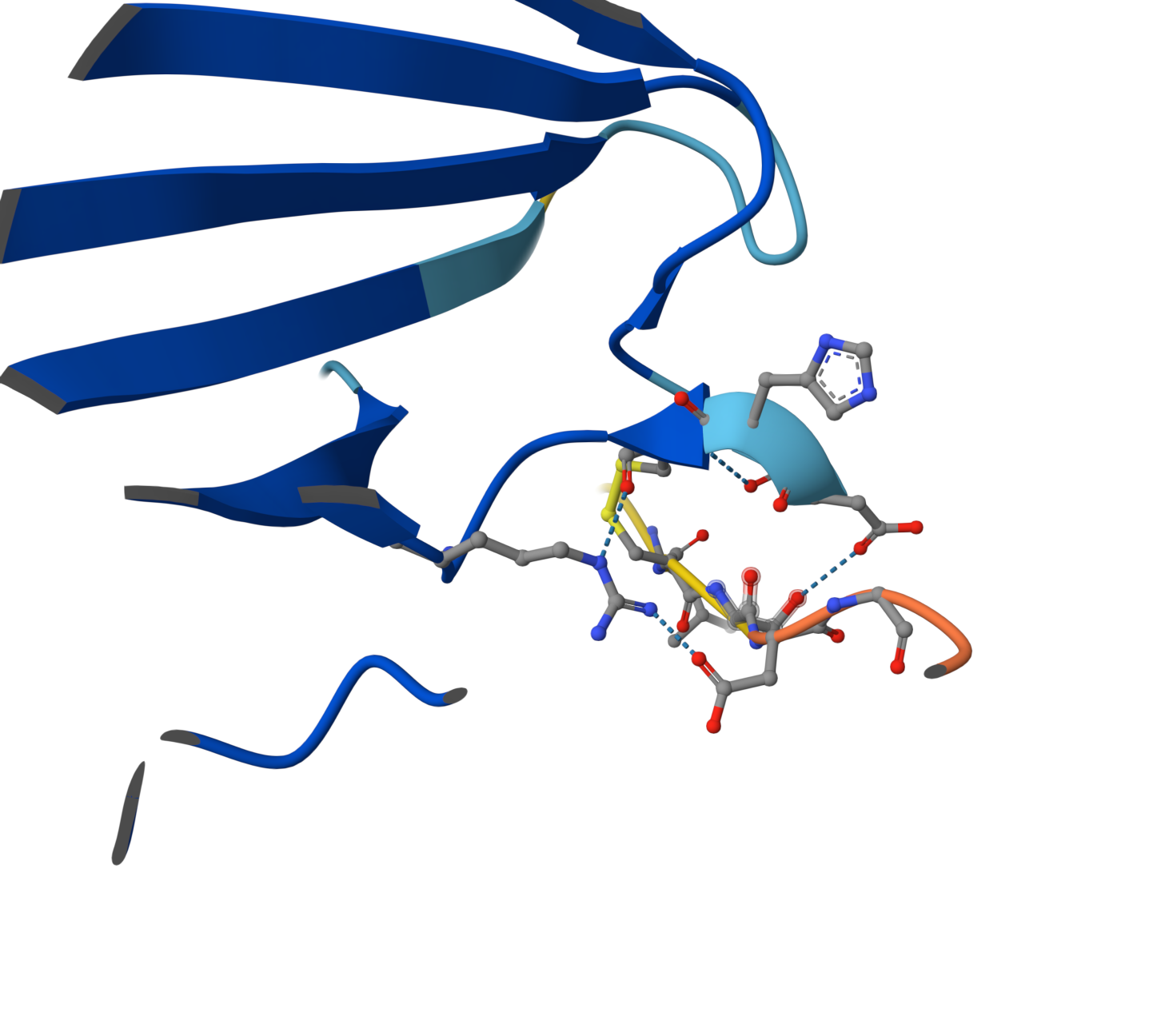

Although the global view suggests a weak interaction (ipTM ~0.35), detailed close-up inspection reveals a highly specific docking event. The peptide binds to a flexible disordered loop (cyan region) rather than the rigid $\beta$-barrel core. This interaction is mediated by a sophisticated hydrogen-bond network and includes crucial electrostatic contacts between a peptide Arginine ($Arg$) and acidic residues on the enzyme’s surface loop. This suggests the peptide acts as an allosteric stabilizer, reducing the flexibility of critical loops near the N-terminus. A key residue (appearing as a nitrogen-rich heterocycle) is perfectly positioned to stabilize the enzyme’s loop through electrostatic contacts.
Candidate 4: ipTM = 0.49 pTM = 0.89
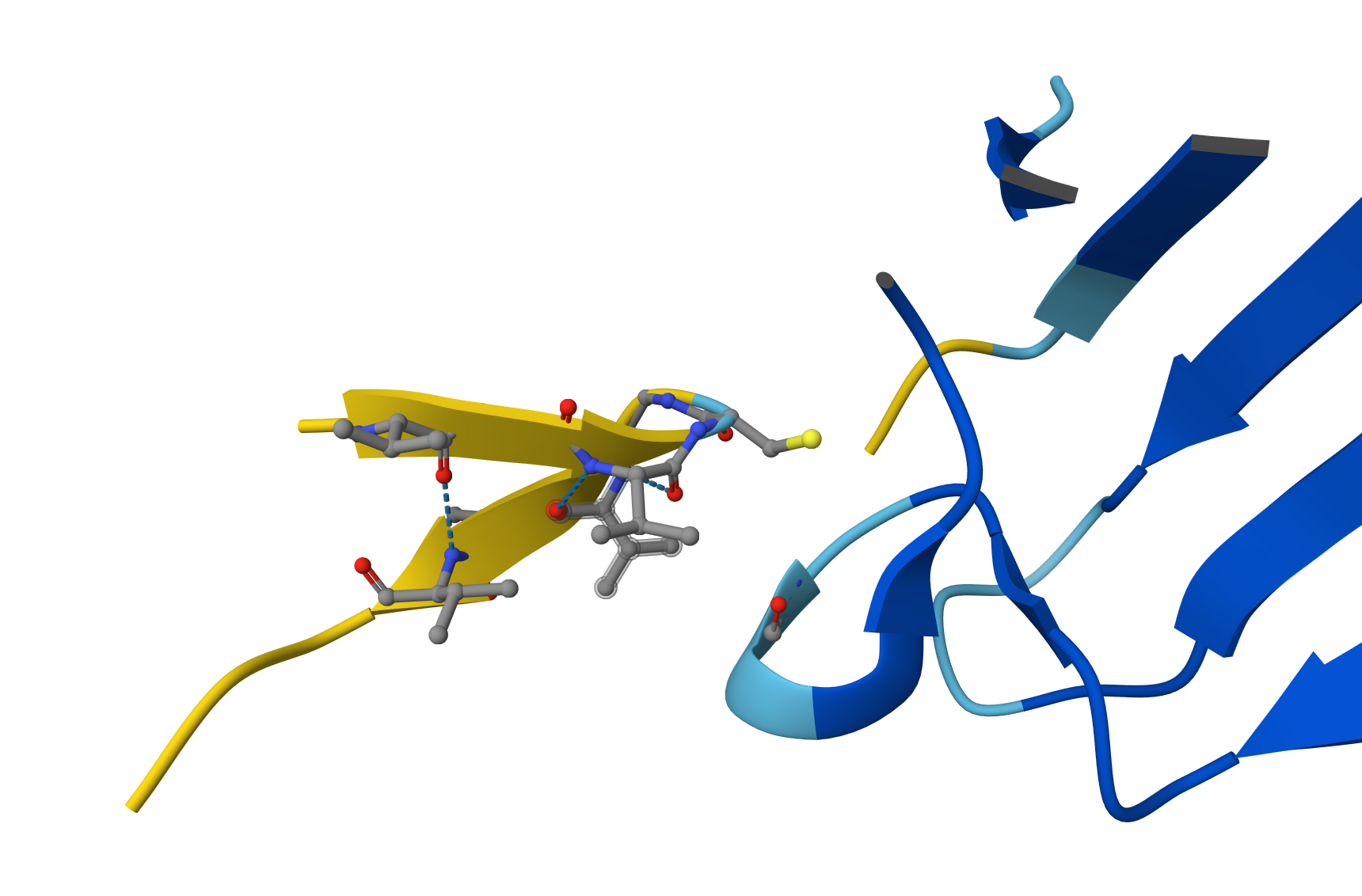


ipTM of 0.49: By falling below the 0.5 threshold, AlphaFold is indicating a lack of confidence in the existence of a stable binding interface. Visual Evidence: The image shows the peptide (yellow chain) physically separated from the SOD1 enzyme (blue chain). Although the peptide attempts to adopt a self-folded conformation, there are no hydrogen bonds connecting it to the protein. pTM of 0.89: As in previous cases, this high value confirms that the SOD1 structure is correctly modeled and stable; the issue is strictly a lack of peptide affinity.
Why does it fail to bind? Despite having numerous Valines (V) and Glycines (G), this peptide appears to be excessively hydrophobic and prone to self-folding. Instead of targeting the SOD1 surface, the peptide prefers to interact with itself, remaining “afloat” in the solvent without engaging the target.
After a comprehensive analysis using PepMLM for generation and AlphaFold3 for structural validation, two distinct strategies for stabilizing the mutant SOD1 emerge:
Candidate 2 (LLGSDGALQVGS) - The Structural Lead: > With the lowest perplexity score (14.65) and a superior ipTM of 0.54, this peptide stands out as the most structurally stable binder. Its aliphatic composition allows it to dock firmly against the protein’s core, acting as a reliable “patch” for the hydrophobic vulnerability created by the A4V mutation. 💎
Candidate 3 (SGVAVLCSDGQG) - The “Dark Horse” Candidate: > While its global confidence metrics are lower, high-resolution inspection reveals a fascinating “allosteric” mechanism. Candidate 3 demonstrates a sophisticated hydrogen-bond network that specifically “clamps” onto disordered surface loops. By inmovilizing these flexible regions near the N-terminus, it could provide a unique form of protection against the unfolding process that leads to toxic aggregation. 💎
Final Recommendation: > While Candidate 2 is the primary choice for advancement toward therapy due to its overall stability, Candidate 3 warrants further investigation. Its ability to “freeze” specific protein loops offers a complementary approach to traditional binding, potentially providing a more nuanced way to rescue the native fold of the SOD1-A4V enzyme.
The known binder presentes ipTM = 0.33 pTM = 0.79 clearly not a better choice 👎 👎 👎
NOTE: During the interaction with Gemini, the following suggestion was received: “SOD1 is a metalloenzyme, meaning it requires Copper (Cu) and Zinc (Zn) to be stable. If you want an ultra-precise model, you could add this.” That is to say, adding a third element (ligand) in AlphaFold using their specific SMILES strings or chemical identifiers. While the current model focuses on the protein-peptide interface, including these metallic cofactors would better simulate the native, stabilized state of the SOD1 enzyme.
PeptiVerse
Candidate 4 (AVGVCGVAVLGN) presents a contradictory profile. While PeptiVerse predicts the highest binding affinity of the set (6.651 $pKd$), this contradicts the AlphaFold3 structural model, which showed no physical contact with the enzyme (ipTM 0.49).This discrepancy is likely explained by the peptide’s high hydrophobicity (GRAVY: 1.83). Such extreme hydrophobicity often leads to non-specific interactions or self-aggregation rather than targeted docking at the A4V site. Furthermore, its hemolysis probability (0.132) is significantly higher than other candidates, making it a less safe therapeutic option.
A comparison between structural modeling and pharmacological prediction reveals a compelling trade-off. Candidate 2 (LLGSDGALQVGS) maintains the highest structural confidence (ipTM 0.54), but Candidate 3 (SGVAVLCSDGQG) shows a much stronger predicted binding affinity in PeptiVerse (6.242 $pKd$ vs 4.502 $pKd$). Visually, this is supported by the dense hydrogen-bond network observed in the AlphaFold3 close-up, where Candidate 3 effectively “clamps” onto the surface loops. Both peptides show ideal therapeutic profiles with maximum solubility and negligible hemolysis probability, confirming that PepMLM-generated sequences successfully avoid the toxic traits of highly hydrophobic non-binders like Candidate 4.
PeptiVerse Evaluation: Candidate 3 (SGVAVLCSDGQG)Binding Affinity: 6.242 ($pKd$). This is significantly higher than Candidate 2 (4.502). In logarithmic terms, this represents a much stronger predicted affinity for the target.Solubility: 1.000. Like Candidate 2, it is predicted to be perfectly soluble.Hemolysis: 0.022. Even lower than Candidate 2, making it exceptionally safe for systemic use.Net Charge: -1.55. Its slightly more negative charge might contribute to its better solubility and specific interaction with the mutant site. Interestingly, Candidate 3 (SGVAVLCSDGQG) emerges as a superior pharmacological lead.
Optimized Peptides with moPPIt
The peptides generated by moPPIt represent a significant shift from “plausible sampling” to “precision engineering.” Compared to the PepMLM candidates, several key differences emerge:
- Chemical Diversity and Functional Groups: The moPPIt sequences incorporate a wider variety of amino acids, such as Cysteine (C), Tyrosine (Y), and Phenylalanine (F). While the PepMLM leads were primarily aliphatic or polar (rich in L, V, S, D, G), the presence of Cysteine in the moPPIt leads allows for potential disulfide bond formation, which can stabilize the peptide’s conformation and enhance its “clamping” effect on the SOD1 surface.
- Targeted Structural Anchoring: Unlike the stochastic nature of PepMLM, which sampled sequences that could theoretically bind anywhere, moPPIt was guided to specific residues near the A4V mutation site. This targeted approach results in sequences that are chemically optimized to interact with the specific structural pocket destabilized by the mutation.
- Pre-optimized Therapeutic Metrics: By incorporating solubility and hemolysis guidance during the generation process, moPPIt avoids the pitfalls of extreme hydrophobicity seen in some PepMLM candidates (like Candidate 4, which had a GRAVY score of 1.83). This ensures that the generated sequences are not just good binders but also safe pharmacological leads.
Pre-Clinical Evaluation Strategy: Before advancing these moPPIt candidates to clinical trials, they must be validated through a multi-step process:Structural Validation (In Silico): Molecular Dynamics (MD) Simulations: Static models like AlphaFold3 are insufficient. MD simulations are required to evaluate the binding residence time and ensure the peptide remains docked at the A4V site under physiological fluctuations. Biochemical Assays (In Vitro): Thioflavin T (ThT) Fibrillization Assay: This is the most critical functional test. It determines if the peptide can successfully inhibit the aggregation of mutant SOD1 into toxic fibrils. Surface Plasmon Resonance (SPR): This provides an accurate measurement of the Dissociation Constant ($K_d$) and binding kinetics (on/off rates) to verify the affinity predicted by PeptiVerse. Biological & Safety Testing:Proteolytic Stability: Since these are peptides, they must be tested for resistance to serum proteases to ensure a sufficient half-life in the human body. Cellular Toxicity: The leads must be tested on motor neuron cultures expressing the A4V mutation to confirm they reduce cellular stress and improve neuron survival without inducing off-target toxicity.
Note on AI Collaboration:
The technical responses and structural analyses presented in this work were developed with the assistance of Gemini, an artificial intelligence model by Google. Gemini provided the initial drafts and technical frameworks based on the raw data from AlphaFold3, PepMLM, PeptiVerse, and moPPIt. The final review, polishing, and scientific validation were performed by the student to ensure accuracy and alignment with the course objectives.
Part C: Final Project: L-Protein Mutants
Analysis of Clustal Omega Alignment
- Soluble Region (Residues 1–40)
This region is critical for DnaJ chaperone interaction.
Highly Variable (Ideal for Mutation):
- Positions 1–6: The start of the protein shows significant variation (METRFP vs METQSP vs MEIRFP). Position 4 is particularly flexible.
- Positions 15–19: This loop varies between STNRR, STNRF, and STNRY. Mutating these could alter the binding surface for DnaJ.
Conserved (Avoid Mutating):
- Positions 21–25 (PFKHE): These residues are almost identical across all sequences, suggesting they are structurally vital.
- Positions 30–38 (RRQQRSST): This motif is highly conserved.
- Transmembrane Region (Residues 41–75)
This region integrates into the membrane to form pores.
Variable (Ideal for Mutation):
- Position 45: Changes between F (Phenylalanine) and C (Cysteine).
- Position 73: Varies between Q (Glutamine) and R (Arginine). Adding a charge here could affect how the protein sits in the membrane.
Conserved (Avoid Mutating):
- Positions 48–60 (LAIFLSKFTNQLL): This hydrophobic core is very consistent, as it must maintain a specific shape to span the lipid bilayer.
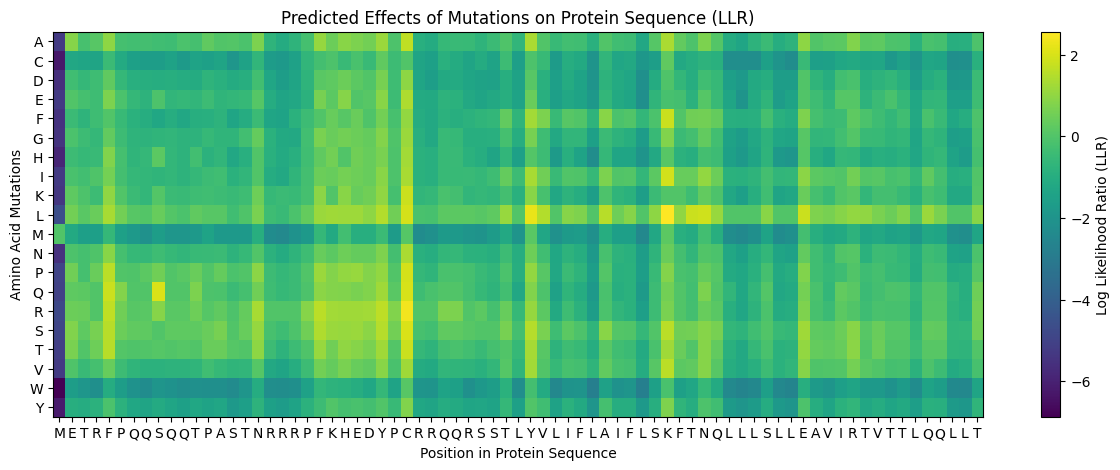

According to the graph, in general, we should avoid the aminoacids Cysteine, Methionine, and Tryptophan. Residue 4 and residues 21-28 seem good options to mutate.
Looking at the excel with experimental data, there is a clear “functional window” for engineering between residues 13 and 31 of the soluble domain. In this region, multiple mutations—such as R18G, R20W, and K23E—maintain a Lysis score of 1, demonstrating that this domain is structurally flexible and can tolerate amino acid substitutions without losing functional integrity. In contrast, the N-terminal start (residues 1–11) and the transmembrane core (residues 48–60) are highly sensitive, where most mutations result in Lysis 0 due to the disruption of protein production or pore-forming capability. Therefore, our engineering strategy focuses on introducing mutations within the residue 13–31 range to optimize DnaJ-independent folding and protein expression while preserving the essential lytic activity of the phage.
Functional Robustness in the TM Domain: * Between residues 38 and 75, there are numerous “safe” substitutions, such as T49S, A63V, and T69S, all of which maintain full lysis activity. This suggests that while the hélice must span the membrane, it can tolerate many conservative amino acid changes without losing its pore-forming ability.
The “Lethal” Exceptions: Even within this functional window, there are critical “black holes” where any change causes failure. For example, position 49 is highly sensitive; while S49L works, several other mutations at this exact spot lead to 0 lysis.
Position 60 is a “dead zone”: L60P, L60V, and L60Q all result in a total loss of function.
The Soluble Domain (Residues 1–40)
This region interacts with the host chaperone DnaJ. Highly Sensitive Sites (Lysis = 0): * Position 1 (M1I, M1T): Any change to the start codon abolishes protein production and lysis. Position 3 (T3I, T3S): Mutations here result in a total loss of function. Position 33 (Q33H): Changing Glutamine to Histidine at this position stops lysis. Tolerant Sites (Lysis = 1): Position 18 (R18G, R18I): The protein remains functional, suggesting this part of the soluble domain is flexible. Position 20 (R20W, R20L): These substitutions are well-tolerated. Position 23 (K23E): This site is resilient to change.
Proposed Mutations for MS2 L-Protein Engineering
R18G + R20W + K23E: Combines three sites proven to be functional (Lysis 1) in the lab data. By changing these three residues simultaneously, we drastically alter the electrostatic surface of the N-terminal domain to ensure DnaJ independence while maintaining high protein levels.
S15A + R19S + Q32E: Targets the highly variable “loop” residues identified in ClustalOmega. Replacing these with smaller or differently charged residues (S to A, R to S, Q to E) aims to create a “stealth” soluble domain that fails to bind the host’s mutated DnaJ chaperone.
F45A + A63V: Combines two experimentally validated “safe” sites in the lysis-active region. This combination aims to stabilize the hydrophobic hélice (A63V) while testing if the removal of the bulky Phenylalanine (F45A) facilitates faster pore assembly.
T69S + L73R: Uses a proven functional mutation (T69S) paired with an evolutionary change seen in Emesvirus (L73R). The goal is to optimize the C-terminal “anchor” to improve membrane penetration and accelerate bacterial killing.
R18I + T75S: Combines a high-expression soluble mutation (R18I) with a conservative C-terminal tail modification. This variant is designed to test if increasing the initial stability of the protein translates into more efficient processing at the membrane interface.
Week 6 HW: Genetic Circuits Part I: Assembly Technologies
Part A. DNA Assembly
- What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
Phusion High-Fidelity DNA Polymerase:
A Pfu-like enzyme fused to a dsDNA-binding domain (Sso7d). This increases processivity and ensures an error rate 50 times lower than Taq polymerase.
5X Phusion HF Buffer (including $MgCl_2$): Maintains optimal pH and provides Magnesium ions, which act as essential cofactors for the polymerase to catalyze the addition of dNTPs.
dNTPs (Deoxynucleotide Triphosphates): The molecular “bricks” (dATP, dTTP, dCTP, dGTP) used to synthesize the new DNA strand.
Stabilizers: Often including glycerol or mild detergents to maintain enzyme stability through repeated thermal cycling.
- What are some factors that determine primer annealing temperature during PCR?
The annealing temperature is usually calculated as $T_m - 5^\circ\text{C}$. Key factors include:GC Content: G-C pairs have three hydrogen bonds (compared to two in A-T), requiring more thermal energy to denature. Higher GC content increases $T_m$.Primer Length: Longer primers have more total hydrogen bonds, leading to a higher melting temperature.Salt Concentration (Cations): $Na+$ and $Mg{2+}$ ions in the buffer stabilize the DNA double helix by neutralizing the negative charges of the phosphate backbone.Primer Concentration: Higher concentrations can slightly shift the kinetics of hybridization.
There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
To ensure success in a Gibson Cloning reaction, you must verify the following:
- Overlapping Ends: Adjacent fragments must share 15–40 bp of identical sequences at their ends. This is achieved by designing PCR primers with “overhangs” that match the neighboring fragment.
- DNA Purity: You must remove the original template (via DpnI digestion) and residual primers to prevent non-specific products.
- Correct Concentration: Fragments should be added in specific molar ratios (e.g., 1:2 or 1:3 vector-to-insert) to optimize assembly efficiency.
- How does the plasmid DNA enter the E. coli cells during transformation?
During Chemical Transformation (using $CaCl_2$):Neutralization: Calcium ions ($Ca^{2+}$) neutralize the negative charges of both the DNA phosphate backbone and the cell membrane’s phospholipids, allowing them to come into close contact.Heat Shock: A sudden increase to 42°C creates a thermal gradient that generates temporary pores in the plasma membrane.Entry: The pressure difference and thermal motion “push” the DNA into the cytoplasm before the cells are moved to a recovery medium to heal the membrane.
- Describe another assembly method in detail (such as Golden Gate Assembly)
Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online).
Golden Gate Assembly is a molecular cloning method that utilizes Type IIS Restriction Enzymes (such as BsaI or BsmBI) and T4 DNA Ligase. Unlike standard enzymes, Type IIS enzymes cut outside of their recognition sites, allowing for the creation of custom, non-palindromic “sticky ends” (overhangs). The reaction is performed in a “one-pot” format, where digestion and ligation occur simultaneously; because the ligation product lacks the original restriction site, it cannot be re-digested, driving the reaction toward the final assembly. This method is scarless and highly efficient, enabling the directional assembly of 10+ fragments in a single step. It is the gold standard for creating complex genetic circuits and modular libraries in Synthetic Biology.
Model this assembly method with Benchling or Asimov Kernel!
Part B. Asimov Kernel
Week 7 HW: Genetic Circuits Part II: Neuromorphic Circuits
Part 1: Intracellular Artificial Neural Networks (IANNs)
- What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
Traditional genetic circuits operate like a light switch (0 or 1). IANNs, however, behave like a signal processor, offering several critical advantages:
Analog vs. Digital Processing. Boolean circuits only detect if a signal is “present” or “absent.” IANNs process signals analytically, so they can distinguish between low, medium, and high concentrations. This allows the cell to respond to gradients, which is much closer to how natural biological systems actually function.
Multivariable Pattern Classification. A Boolean circuit (using AND/OR gates) becomes incredibly complex and “brittle” as you add more inputs. By using a neural network architecture, IANNs can integrate multiple signals simultaneously (e.g., 5 different microRNAs). Instead of a simple “yes/no” gate, the IANN creates a complex decision boundary. This allows a cell to identify a specific state (like a cancer cell) with much higher precision, filtering out false positives that a simple Boolean circuit would miss.
Programmable “Weights” (Tunability). In a traditional circuit, if you want to change the behavior, you often have to re-engineer the entire genetic architecture. IANNs allow you to tune behavior simply by adjusting the weights of the connections (e.g., changing an RBS strength or a protein’s binding affinity). This makes the system modular and reprogrammable without changing the basic “wiring” of the circuit, allowing the same biological “brain” to be adapted for different tasks.
Robustness to Biological Noise. In Biology is “noisy”—molecule levels fluctuate randomly. Boolean circuits are sensitive to this noise and can “misfire” easily. IANN Advantage: By using ReLU activation functions and Sequestron mechanisms (molecular sequestration), IANNs act as filters. They can ignore small fluctuations (noise) and only “fire” a response when the weighted sum of signals is clear and consistent.
Multi-layer Composition (Deep Logic). IANN Advantage: IANNs are inherently multi-layer. This allows for sophisticated behaviors like Bandpass filters (activation only within a specific middle range), which are extremely difficult to achieve with pure Boolean logic. This “layered” capability allows biological computation to be much deeper and more “intelligent.”
- Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
In my HTGAA Final Project, I am exploring the use of an Intracellular Artificial Neural Network (IANN) as a sophisticated control strategy for a system designed for autonomous cell lysis. The core objective is to ensure that E. coli cells only undergo self-destruction and release their contents when they have reached a specific “Peak Harvest” state. To achieve this, the IANN acts as a biological classifier that integrates three distinct analog inputs: temperature-sensitive riboswitches to align with the fermentation phase, phosphate sensors to detect nutrient depletion, and membrane-tension riboswitches that signal high internal polymer accumulation.
- Input 1: Thermal Stress (Temperature). Using riboswitches that respond to temperature shifts. This ensures the lysis “arm” is only primed during the specific thermal phase of the industrial fermentation.
- Input 2: Phosphate Levels. A sensor for low phosphate (a common signal for the end of the growth phase), ensuring the bacteria don’t explode while they are still actively replicating.
- Input 3: Membrane Tension / Stress. Riboswitches or promoters that sense the physical stretching of the cell membrane or metabolic stress caused by high PHA/PHB (polymer) accumulation.
- The output: activation of the lysis cassette
The primary advantage of using an IANN over a simple Boolean “AND” gate is its ability to perform a weighted sum of these signals. By tuning the “weights” of the network—specifically the Translation Initiation Rate (TIR) of the RBS for each sensor—I can program the cell to ignore minor “leakiness” or noise from a single sensor. This ensures that the lysis cassette (SRRz) only triggers when the combined mathematical score of all three inputs crosses a precise threshold, preventing the premature loss of the batch.
However, implementing an IANN for this goal presents significant engineering challenges. The most critical limitation is the metabolic burden; producing multiple repressors and “decoy” binding sites to maintain the network’s logic consumes ATP and ribosomes that would otherwise be used for bioplastic synthesis. Furthermore, maintaining orthogonality among multiple sensors to avoid cross-talk is complex. While a simpler circuit might be more efficient, the IANN offers a level of programmable robustness that could be vital for scaling Bioplastix (the startup that would incorporate the auto-lysis strategy in its Bioprocess) to industrial-level bioreactors where environmental conditions are constantly fluctuating and are not homogeneous.
- Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
Part 2: Fungal Materials
- What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Fungal materials are primarily made from mycelium, the underground root-like network of a fungus. This mycelium acts as a natural “glue” that can bind agricultural waste into solid structures.
Mushroom Packaging: Companies like Ecovative grow mycelium around husks or stalks in molds to replace Styrofoam. It is used for shipping everything from electronics to wine.
Mycoleather: Textiles like Mylo or Reishi mimic the look and feel of animal leather. High-end fashion brands are using it for bags and garments as a sustainable alternative.
Fungal Bricks and Insulation: Experimental architecture uses mycelium blocks for their natural fire resistance and acoustic insulation properties.
Acoustic Panels: Mycelium-based tiles are used in interior design to absorb sound in offices or studios.
- Pros: They are biodegradable, carbon-negative (they sequester carbon as they grow), fire-resistant, and non-toxic. Production requires very little energy compared to plastic or leather tanning.
- Cons: They are often hydrophilic (absorb water), which can lead to rot if not properly coated. They generally have lower tensile strength than synthetic plastics or traditional leather and can vary in consistency.
It is crucial to clarify that mycelium is not a species or a family, but an anatomical part of a fungus. It consists of a dense, branching network of thread-like filaments called hyphae. In the field of biomaterials, this network acts as a natural binder to create structural composites.
Key Filamentous Fungi (Mycelium-based):
- Ganoderma lucidum (Reishi): Widely used in the production of mycoleather. Its hyphae grow extremely dense, creating a flexible and durable material that serves as a sustainable alternative to animal hides.
- Pleurotus ostreatus (Oyster Mushroom): The standard for bio-packaging. It is a fast-growing, aggressive colonizer that can quickly turn agricultural waste into molded shapes, replacing expanded polystyrene (Styrofoam).
- Trametes versicolor (Turkey Tail): Frequently studied for bioremediation. It produces powerful extracellular enzymes (laccases) capable of breaking down complex chemical toxins and dyes in environmental applications.
Unicellular Fungi (Yeasts): Beyond complex mycelial networks, yeasts represent a vital category of unicellular fungi.
Saccharomyces cerevisiae: This is perhaps the most industrially significant fungus. Beyond its traditional roles in baking and brewing, it is a primary “chassis” in synthetic biology for the large-scale production of bioethanol and high-value recombinant proteins (like insulin). Its well-understood genetics make it an ideal eukaryotic model for metabolic engineering.
- What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
If we apply synthetic biology to fungi, we can transform these materials from “passive” objects into “living” materials:
- Self-Healing Materials: Engineering fungi to remain dormant within a structure and “wake up” to grow and seal cracks when moisture is detected.
- Sensing and Reporting: Modifying fungi to change color or glow in the presence of environmental toxins (like heavy metals in soil).
- Enhanced Secretion: secrete specific enzymes.
Advantages of Fungi vs. Bacteria:
While E. coli is the workhorse of synthetic biology, fungi offer unique engineering advantages:
- Macro-scale Structure: Unlike bacteria, which are unicellular, fungi form massive, interconnected multicellular networks. This allows for the creation of large-scale physical materials that hold their shape.
- Eukaryotic Processing: Fungi are eukaryotes. They can perform complex post-translational modifications on proteins that bacteria cannot, making them better for producing specialized enzymes or human-like proteins.
- Superior Secretion: Fungi are natural “secretory machines.” They can pump out vast quantities of proteins and metabolites into their environment, which is much more efficient for industrial harvesting than lysing bacteria.
- Environmental Resilience: Fungi thrive in harsh, acidic, or low-moisture environments where most lab bacteria would die. This makes them ideal for “out-of-lab” applications like bioremediation or outdoor construction.
Week 9 HW: Cell-Free Systems
Part 1: General and Lecturer-Specific Questions
General homework questions
- Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
The fundamental advantage of cell-free protein synthesis (CFPS) lies in the removal of the cellular membrane, which effectively transforms a “black box” biological process into an open, accessible engineering platform. By eliminating the cell wall, researchers gain unprecedented flexibility and direct control over experimental variables; the reaction environment can be precisely manipulated by adding non-natural amino acids, specific chaperones, or tailored energy sources without the constraints of cellular transport or homeostasis. Furthermore, CFPS decouples protein production from host viability, allowing for the synthesis of highly cytotoxic proteins that would otherwise trigger cell death and halt production in traditional in vivo systems.Beyond throughput, the “open” nature of the system significantly enhances real-time monitoring and process optimization. Unlike the opaque interior of a living E. coli cell, a cell-free reactor allows for millisecond-scale sampling and mid-process adjustments of critical concentrations—such as magnesium levels or pH—to maximize yields. Perhaps most importantly for rapid prototyping, CFPS enables a drastically accelerated iteration cycle. By bypassing time-consuming steps like transformation, plating, and overnight culturing, researchers can transition from a linear DNA template (such as a PCR product) to a functional protein in just a few hours, representing a paradigm shift in the speed of biological design.
Case A: Production of Cytotoxic Proteins
Many useful proteins, such as antimicrobial peptides (AMPs) or certain lytic enzymes (like the ones you might use in Bioplastix), kill the host cell as soon as they are expressed. CFPS allows you to synthesize these “suicide” proteins because the system lacks the physiological targets that the toxins would otherwise destroy.
Case B: Incorporation of Non-Standard Amino Acids (nsAAs)
If you want to create a protein with expanded chemical properties (e.g., for site-specific labeling, “click” chemistry, or enhanced stability), CFPS is superior. In a cell, it is extremely difficult to “force” the machinery to use a synthetic amino acid without interfering with the cell’s own survival. In a cell-free extract, you can simply “starve” the reaction of a natural amino acid and flood it with the synthetic version.
- Describe the main components of a cell-free expression system and explain the role of each component.
- The Biological Extract (The Machinery): usually obtained by lysing cells (such as E. coli, wheat germ, or rabbit reticulocytes) and removing the cell wall and genomic DNA. It provides the Ribosomes for translation, RNA Polymerase for transcription, and various tRNAs, aminoacyl-tRNA synthetases, and initiation/elongation factors. Without the extract, there is no hardware to read the genetic code.
- The DNA Template (The Instructions): Unlike in vivo systems that require circular plasmids, cell-free systems can often use linear DNA (like PCR products). It provides the genetic sequence of the protein of interest. It must contain specific regulatory elements that the extract’s machinery can recognize, such as a T7 or endogenous promoter, a Ribosome Binding Site (RBS), and a terminator.
- Energy Regeneration System (The Fuel): Protein synthesis is energetically expensive. Since the cell’s natural mitochondria or metabolic pathways are no longer intact, we must provide an external energy source. It consists of NTPs (ATP, GTP, UTP, CTP) which act as the direct building blocks for mRNA and the energy source for the ribosome. It also includes an energy-rich secondary substrate (like phosphoenolpyruvate (PEP) or creatine phosphate) and a corresponding kinase to “recharge” the ATP as it is consumed.
- Small Molecules and Buffers (The Environment): A precise chemical environment is required to keep the enzymes stable and active. Amino Acids: The raw building blocks used to assemble the protein chain. Magnesium and Potassium salts: Critical cofactors for ribosome assembly and stability. Buffers (e.g., HEPES): To maintain a stable pH, as the metabolic byproducts of the reaction can quickly acidify the mixture.
- Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
Protein synthesis requires a massive amount of energy at every stage. For every single amino acid added to a polypeptide chain, four high-energy phosphate bonds are consumed: two during the “charging” of the tRNA with an amino acid and two during the translation elongation steps (GTP hydrolysis).In a static cell-free batch, the initial supply of ATP would be depleted almost instantly. Furthermore, the accumulation of Inorganic Phosphate —a byproduct of ATP hydrolysis—can inhibit the reaction by chelating magnesium ions, which are essential for ribosome stability. Therefore, a regeneration system is critical not just to keep the “fuel tank” full, but to maintain a chemical environment that isn’t poisoned by its own waste.
To ensure a steady supply of ATP in your HTGAA experiments, two strategies are:
The Creatine Phosphate / Creatine Kinase (CP/CK) System. This is the most common “plug-and-play” method for cell-free experiments. You add Creatine Phosphate (the high-energy substrate) and the enzyme Creatine Kinase to the mixture. Every time an ATP molecule is used and becomes ADP, the Creatine Kinase transfers a phosphate group from the Creatine Phosphate directly back to the ADP, “recharging” it into ATP instantly. It is highly efficient and maintains a very high [ATP]/[ADP] ratio, which is vital for high-yield protein production.
PURE System or Secondary Carbon Sources. If you are using an E. coli extract, you can utilize the cell’s own residual glycolytic enzymes. You provide a secondary energy source like Phosphoenolpyruvate (PEP) or Glucose-6-Phosphate. The enzymes already present in the extract (like pyruvate kinase) process these substrates to regenerate ATP. The “Continuous” Approach: For very long experiments, you can use a Dialysis System (Continuous Exchange Cell-Free - CECF). The reaction happens inside a dialysis membrane submerged in a large reservoir of buffer, energy substrates, and nucleotides. Fresh fuel diffuses in, and inhibitory byproducts like inorganic phosphate diffuse out, allowing the reaction to run for days.
- Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
The choice between prokaryotic and eukaryotic cell-free expression systems is primarily dictated by the complexity of the protein and the requirement for post-translational modifications (PTMs). Prokaryotic systems, typically based on E. coli S30 extracts, are the “workhorses” of the field due to their high protein yields (often reaching mg/mL levels), low cost, and rapid synthesis rates. They utilize 70S ribosomes and simpler energetic pathways but lack the machinery for complex folding or PTMs like glycosylation. In contrast, eukaryotic systems—such as Wheat Germ Extract (WGE) or Rabbit Reticulocyte Lysate (RRL)—employ 80S ribosomes and offer a more sophisticated folding environment. While they generally produce lower total yields and are more expensive to prepare, they are indispensable for synthesizing large, multi-domain eukaryotic proteins that require authentic folding or specific modifications like disulfide bond formation, phosphorylation, or lipidation.
70S and 80S Ribosomes: These are the molecular machines responsible for protein synthesis. The “S” (Svedberg unit) indicates their size and sedimentation rate during centrifugation.
70S Ribosomes: Found in prokaryotes (bacteria) and organelles like mitochondria. They are smaller and simpler in structure.
80S Ribosomes: Found in eukaryotes (animals, plants, fungi, and humans). They are larger, more complex, and capable of more sophisticated regulation of translation.
PTMs are chemical changes made to a protein after it has been synthesized by the ribosome. These modifications are essential for the protein to become biologically functional. Examples: Glycosylation (adding sugars), phosphorylation (adding phosphate groups), and disulfide bond formation.
Importance: While bacteria are efficient at making simple proteins, they lack the machinery for most complex eukaryotic PTMs, which is why human proteins often require eukaryotic expression systems to function correctly.
For a prokaryotic system (E. coli), an ideal protein to produce would be Green Fluorescent Protein (GFP) or a bacterial enzyme like Pyruvate Kinase. GFP is a robust, relatively small protein that folds efficiently in the bacterial cytoplasm without needing PTMs. Using an E. coli cell-free extract allows for massive production in just a few hours, making it perfect for high-throughput screening of genetic circuits or biosensors where speed and quantity are prioritized over structural complexity.
For a eukaryotic system (such as Rabbit Reticulocyte Lysate), a strategic choice would be a human therapeutic protein like Erythropoietin (EPO) or a complex Single-Chain Variable Fragment (scFv) antibody. EPO requires extensive and specific glycosylation to be biologically active and stable in the human body—a process that E. coli machinery cannot perform. By using a mammalian-derived cell-free system (often supplemented with microsomal membranes), researchers can ensure the protein is correctly glycosylated and folded with the necessary disulfide bridges, providing a functional product that closely mimics its natural human counterpart.
- How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
Use a cell-free system supplemented with liposomes or nanodiscs (synthetic lipid bilayers). This allows the membrane protein to integrate into a stable environment as it is synthesized.
Challenge 1: Hydrophobicity and Aggregation. Membrane proteins often precipitate in aqueous extracts. Solution: Add detergents (at sub-CMC levels) or chaperones to maintain solubility.
Challenge 2: Lack of Energy/Substrates. Solution: Use a Continuous-Exchange (CECF) system to provide a steady supply of energy and remove inhibitory byproducts, ensuring longer reaction times for difficult folding.
- Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
- DNA Template Quality: Verify DNA purity (A260/280 ratio) and ensure the promoter/RBS sequences are optimized for the specific extract used.
- Resource Exhaustion: Increase the concentration of the energy regeneration system (e.g., Creatine Phosphate) or optimize the Magnesiu concentration.
- Protein Degradation: Add Protease Inhibitors to the extract or use a specialized strain (like BL21 E. coli) that is deficient in endogenous nucleases and proteases.
Homework question from Kate Adamala
Design of a Synthetic Minimal Sentinel Cell for Autolysis Testing
- Function and Operation: A “Self-Destructing Sentinel” that replicates the dual-sensing logic. It is designed to sequester a payload and release it only when two specific environmental conditions are met: phosphate starvation and a temperature shift. Input: 1. Low concentration of inorganic phosphate. 2. Thermal shift (e.g., transition from 30°C to 37°C).Output: Expression of the Lambda phage lysis cassette (SRRz) leading to membrane rupture and release of internal contents.
- Realization and Feasibility: Cell-free Tx/Tl alone: Without encapsulation, the “lysis” has no physical meaning. You could produce the proteins, but you wouldn’t be able to measure the structural failure or the release kinetics that your industrial process needs. Genetically modified natural cell: While this is an end goal for E. coli, testing it first in a minimal cell is safer. In a natural cell, phosphate starvation triggers many “survival” pathways that might interfere with your promoter’s strength. The minimal cell provides a “clean” signal-to-noise ratio.
- Components and Encapsulation: Membrane: A lipid bilayer composed of POPC and DOPE (1,2-dioleoyl-sn-glycero-3-phosphoethanolamine). DOPE adds curvature tension, making the membrane more sensitive to the “holes” created by the holins (S protein) of the SRRz cassette. Internal Content: Extract: E. coli S30 cell-free extract (Prokaryotic system). Machinery: Endogenous E. coli RNA polymerase and ribosomes. Small Molecules: NTPs, Amino Acids, and a buffer with initially high phosphate that will be “consumed” or diluted to trigger the circuit.
- Communication and System Origin: Bacterial (E. coli). The circuit uses the pPhoA promoter, which is native to E. coli. Using a bacterial cell-free extract ensures that the transcription factors (like PhoB) required for the pPhoA promoter to work are present. To simulate phosphate starvation, we can use Alpha-hemolysin pores. These allow phosphate to diffuse out of the minimal cell into a phosphate-free external buffer, triggering the internal sensor.
- Experimental Details: The minimal cell will encapsulate:Lipids: POPC and DOPE. Genes (The Circuit): pPhoA Promoter: Driving the first stage of the cascade. It responds to the lack of phosphate.Temperature-Sensitive Riboswitch: Placed upstream of the SRRz sequence. Even if pPhoA is active, the mRNA won’t be translated unless the temperature reaches the threshold (e.g., 37°C), which unfolds the riboswitch. SRRz Cassette: The “actuator” consisting of S (holin), R (endolysin), and Rz (spanin).
- Measuring Function: We will validate the design using: Phase-Contrast Microscopy to visually observe the “bursting” of the vesicles when phosphate is depleted and temperature is raised. Fluorescence Release Assay: We will co-encapsulate a large fluorescent protein (like mCherry). We will measure the increase in external fluorescence over time. Success Metric: If fluorescence remains internal at 30°C regardless of phosphate levels, but releases at 37°C under low phosphate, the logic is proven.
Homework question from Peter Nguyen
Proposal Pitch: The “Sentinel Bio-Textile”
One-sentence summary pitch: A smart, biodegradable textile infused with freeze-dried cell-free systems that acts as a “living” safety suit, detecting environmental pathogens and neutralizing them through the inducible secretion of antimicrobial peptides.
How will the idea work? The textile fibers are embedded with a freeze-dried E. coli cell-free extract containing the genetic instructions for a pathogen-sensing riboswitch and an actuator gene. When a specific pathogen (or a chemical marker of contamination) is detected, the dehydrated machinery is activated by ambient moisture or sweat, triggering the transcription and translation of the circuit. The system then produces and secretes Antimicrobial Peptides (AMPs) or lytic enzymes directly onto the fabric surface. This creates a localized, on-demand decontamination zone without the need for living, genetically modified organisms to survive on the wearer’s skin.
What societal challenge or market need will this address? This addresses the growing need for advanced Personal Protective Equipment (PPE) in healthcare and industrial settings, where traditional fabrics only provide a passive physical barrier. Currently, PPE often becomes a vector for cross-contamination; a bio-active textile that actively “cleans” itself or alerts the user to invisible threats would significantly reduce the spread of hospital-acquired infections and enhance worker safety in bio-hazardous environments.
Addressing limitations (Activation, Stability, One-time use) Activation: The freeze-dried components remain dormant and stable at room temperature until they come into contact with a specific trigger, such as a localized application of water or a “developer” spray.
Stability: By using lyoprotectants (like sucrose or trehalose) during the freeze-drying process, the enzymatic machinery is shielded from thermal degradation, allowing for a long shelf-life in standard warehouse conditions.
One-time use: While the reaction is currently a one-time “pulse,” we envision the textile as a modular patch system. Once the “biological fuse” has been spent, the functionalized patch can be swapped out or discarded, taking advantage of the biodegradable nature of the underlying Bioplastix material.
Homework question from Ally Huang



















