Week 2 HW: DNA Read Write & Edit
Part 1: Benchling & In-silico Gel Art
https://benchling.com/s/seq-B1mFk0Oh2ZF9coVeqBMH?m=slm-ySVijsqZThHxK3Qq6kwq
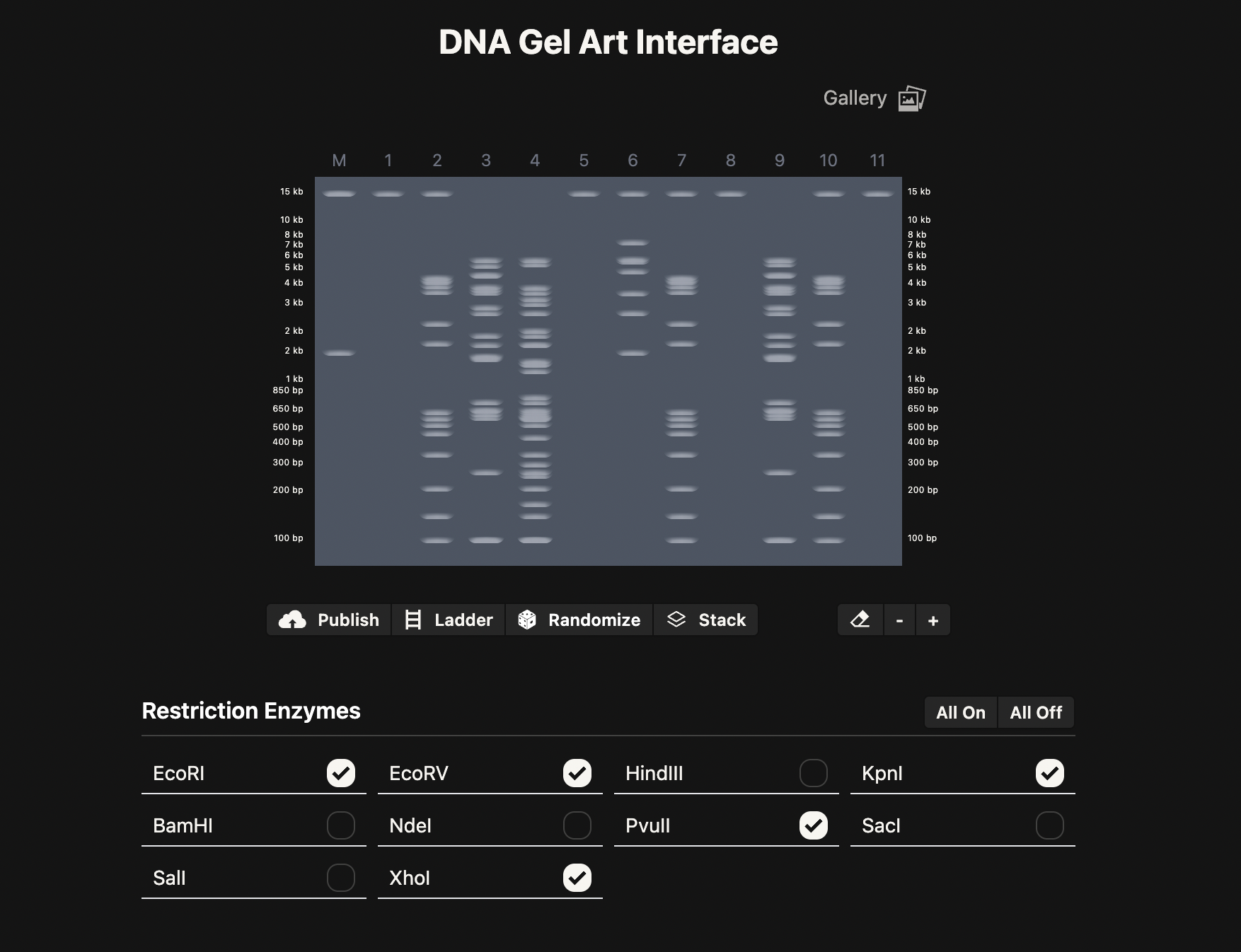

Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
I am interested in the protein Reflectin. This protein is common in Cephalopods and is responsible for their structural colour as their skin changes with the environment. I am interested in the potential applications of Reflectin in smart materials/textiles. Reflectins’ optical properties can be reversibly engineered to change colour under different conditions. There is one study in which Reflecin was used in this way in a thin film substrate, and responsive to hydration or dehydration of the material.
Protein Sequence:
tr|Q6WDN6|Q6WDN6_EUPSC Reflectin 3a OS=Euprymna scolopes OX=6613 PE=4 SV=1 MNRYMNRFRNFYGNMCRNRNRGMMEPMSRMTMDFQGRYMDSQGRMVDPRYYDYYGRYNDY DRYYGRSMFNYGWMMDGDRYNRYNRWMDYPERYMDMSGYQMDMYGRWMDMQGRHCNPYSQ WMMYNYNRHGYYPNYSYGRHMFYPERWMDMSNYSMDMYGRYMDRWGRYCNPFYHYYNHWN RSGNNPGYYSYYYMYYPERYFDMSNWQMDMQGRWMDMQGRYCSPYWYNWYGRQMYYPYQN YYWYGRWDYPGMDYSNWQMDMQGRWMDMQGRYMDPWWMNDSYYNNYYN
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
reverse translation of tr|Q6WDN6|Q6WDN6_EUPSC Reflectin 3a OS=Euprymna scolopes OX=6613 PE=4 SV=1 to a 864 base sequence of most likely codons. atgaaccgctatatgaaccgctttcgcaacttttatggcaacatgtgccgcaaccgcaac cgcggcatgatggaaccgatgagccgcatgaccatggattttcagggccgctatatggat agccagggccgcatggtggatccgcgctattatgattattatggccgctataacgattat gatcgctattatggccgcagcatgtttaactatggctggatgatggatggcgatcgctat aaccgctataaccgctggatggattatccggaacgctatatggatatgagcggctatcag atggatatgtatggccgctggatggatatgcagggccgccattgcaacccgtatagccag tggatgatgtataactataaccgccatggctattatccgaactatagctatggccgccat atgttttatccggaacgctggatggatatgagcaactatagcatggatatgtatggccgc tatatggatcgctggggccgctattgcaacccgttttatcattattataaccattggaac cgcagcggcaacaacccgggctattatagctattattatatgtattatccggaacgctat tttgatatgagcaactggcagatggatatgcagggccgctggatggatatgcagggccgc tattgcagcccgtattggtataactggtatggccgccagatgtattatccgtatcagaac tattattggtatggccgctgggattatccgggcatggattatagcaactggcagatggat atgcagggccgctggatggatatgcagggccgctatatggatccgtggtggatgaacgat agctattataacaactattataac
3.3. Codon optimization.
Optimized Codon Sequence
Using Yeast (Pichia Pastoris)
While E-Coli is a great organism to use for simple codon optimization, in this case with the codon optimization for Reflectin 3a Protein, Yeast ‘Pichia Pastoris’, is a better choice for a more complex fold and protein like Reflectin a more sufisticated and complex eukaryotic protein.
1 ATGAATAGAT ACATGAATAG GTTTAGAAAC TTCTACGGAA ACATGTGTAG GAACAGAAAT AGGGGAATGA 71 TGGAACCTAT GTCCAGGATG ACAATGGACT TCCAAGGAAG GTATATGGAC TCTCAAGGTA GGATGGTTGA 141 TCCTAGGTAC TACGACTACT ACGGTCGTTA TAACGACTAC GACAGATACT ACGGTAGATC TATGTTCAAT 211 TACGGTTGGA TGATGGATGG TGACAGGTAT AATAGATACA ACCGTTGGAT GGATTACCCC GAAAGGTACA 281 TGGATATGAG TGGATATCAA ATGGATATGT ATGGCAGATG GATGGATATG CAAGGAAGAC ATTGCAACCC 351 ATACTCACAA TGGATGATGT ACAATTATAA CAGGCACGGT TATTATCCTA ACTATTCCTA CGGCCGACAT 421 ATGTTCTACC CTGAACGTTG GATGGACATG TCTAACTATT CAATGGACAT GTATGGAAGA TACATGGATA 491 GGTGGGGAAG GTACTGCAAC CCATTTTACC ACTACTATAA CCATTGGAAC AGAAGTGGAA ATAATCCTGG 561 CTACTACTCC TACTACTATA TGTACTATCC CGAGAGATAC TTCGACATGT CCAACTGGCA GATGGACATG 631 CAAGGAAGAT GGATGGATAT GCAAGGAAGA TATTGTTCTC CTTACTGGTA TAACTGGTAT GGTAGACAGA 701 TGTACTATCC ATATCAGAAT TATTATTGGT ACGGAAGGTG GGATTATCCT GGAATGGATT ACTCCAATTG 771 GCAGATGGAC ATGCAAGGTA GATGGATGGA CATGCAAGGC AGATACATGG ATCCATGGTG GATGAACGAC 841 TCCTACTACA ATAACTACTA TAAT
3.4. You have a sequence! Now what?
Reflectin, as a protein, has many potential applications, especially in textiles. Once the yeast has harvested the reflectin protein with optimised codons for yeast Pichia Pastoris, the reflectin protein can be purified and used as a structural colour material in textiles, with multiple different gradients. Relfectin could also be engineered to be expressed in cells with biological signals. In the same way that reflectin is used in cephalopods to express a change in environment in response to stimuli, Reflectin can be engineered into a material that responds to its environment. Reflectin also has electrical and dialectic properties which could be used for bioelectronic systems and technologies. There is increasing research on the future of bioelectronics, including deriving electrical signals from biological organisms and engineered systems in which they are embedded and used to power systems. In the case of Reflectin, it can be incorporated into a bioconductive system, for example, EMG, and respond to one’s own muscular electrical pulses, changing depending on one’s skin conductivity,(Cai.T). Skin conductivity can also reflect one’s neurological system, thereby revealing one’s internal worlds and expressing emotions such as excitement and adrenaline.
Part 4: Prepare a Twist DNA Synthesis Order
4.2. Build Your DNA Insert Sequence
Expression cassette
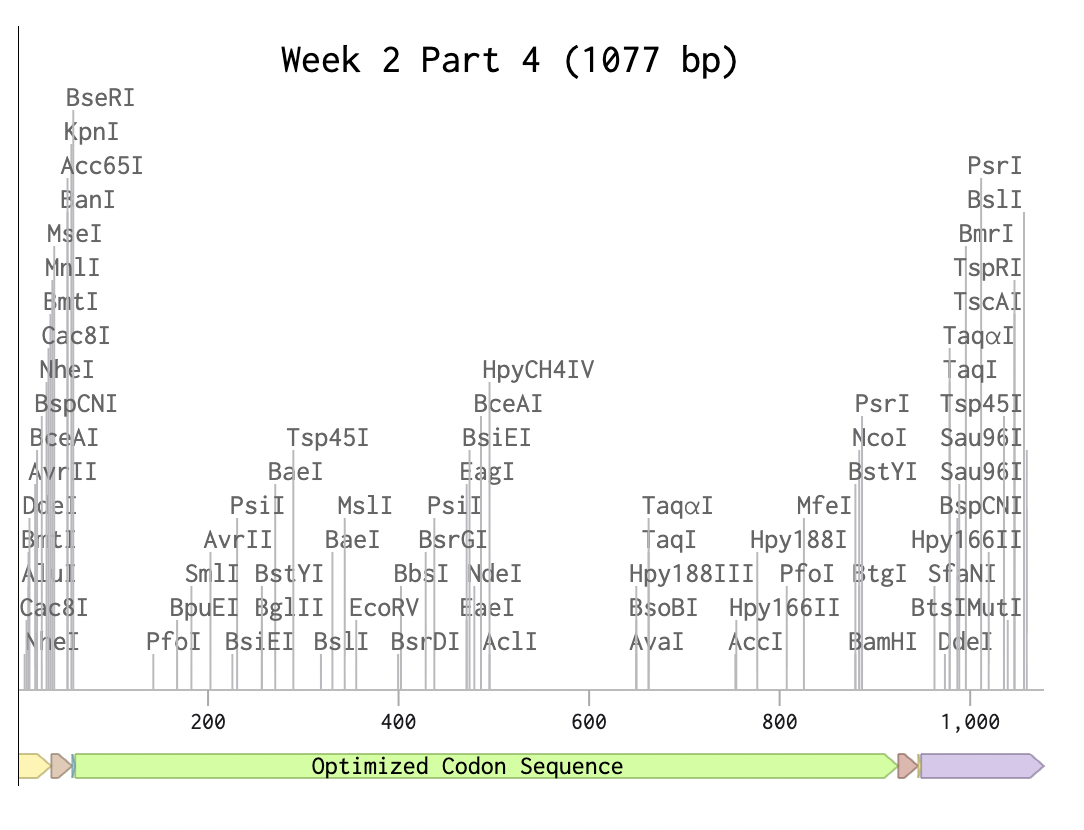
 https://benchling.com/s/seq-gDhXeHq7OCFYtyBIH9bE?m=slm-7Ec7akbFEDQuYeDYt6eK
https://benchling.com/s/seq-gDhXeHq7OCFYtyBIH9bE?m=slm-7Ec7akbFEDQuYeDYt6eK
4.3. On Twist, Select The “Genes” Option
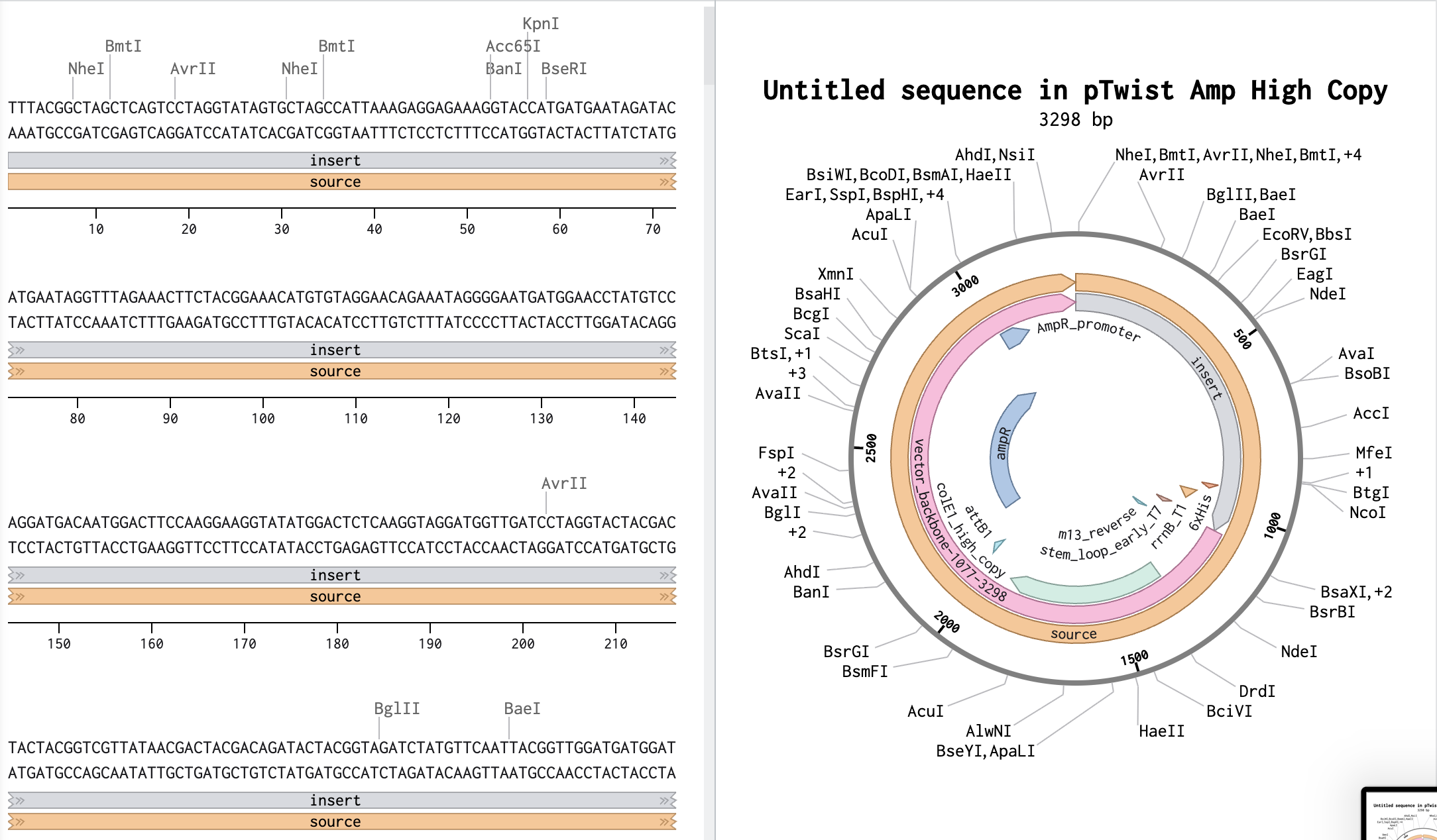

Part 5: DNA Read/Write/Edit
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
I would like to sequence the ‘16S rRNA gene’. This DNA is found in the skin microbiome and serves as a diversity metric for the bacteria in our skin’s gut and more! As we are all made of a diverse ecosystem of bacteria, I would love to further extract one’s metrics from this DNA, which serves as a ‘bacterial barcode’; it is also a cheap and efficient DNA to work with. Additionally, instead of sequencing a single DNA sample, I can sequence all microbial DNA in a single sample. This makes 16S a very efficient DNA. With its diverse micropalette, I would love to explore its potential for applications in materials that reflect one’s unique microflora.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
To perform the DNA edit of the 16S rRNA gene, I would choose the Illumina next-generation sequencing technology as 16S regions are short (250-500bp), and Illumina gives high accuracy, as well as allowing a multiplexity of samples. For more comprehensive functional analysis through whole metagenomic sequencing sampling, I would pick the Oxford Nanopore for long sequencing for long reads to improve genome assembly and detect structural variations. Together, these technologies can provide a functional analysis of the DNA and its microbial species.
Also answer the following questions:
Is your method first-, second- or third-generation or other? How so? What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps. What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)? What is the output of your chosen sequencing technology?
Illumina sequencing is a second-generation ’next-generation’ sequencing technology. It is considered second-generation due to a few factors. Firstly, it performs massively parallel sequencing, where DNA fragments are sequenced simultaneously; it also relies on sequencing by synthesis of clonal amplification. This is unlike technologies like Sanger, which sequence one fragment at a time, and require PCR amplification.
The preparation steps begin with collecting microbial DNA from skin swabs. Next, total microbial DNA is extracted through cell lysis and purification. The 16S rRNA hypervariable regions are then amplified using PCR with region-specific primers. In a subsequent indexing PCR step, Illumina adapters and sample-specific barcodes are added to both ends of the amplicons to enable multiplexing and flow cell binding. The amplified libraries are purified to remove primer dimers and unwanted fragments, quantified, normalised, and pooled. The final product is an adapter-ligated DNA library ready for cluster generation and sequencing on an Illumina platform.
Illumina decodes DNA through ‘sequencing by synthesis’. Flourecently labeled nucelotideswith reversible terminators are incorporated one base at a time. Fluorescent imaging then detects each nucleotide. The software then converts the visible fluorescent signals into base calls (A, C, T, G). The output is millions of sequence reads stored as FASTQ files, including all nucleotide sequences and quality scores for each base.
5.3 DNA Edit (i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
Based on the DNA sequencing of 16S, and identifying the microuniverse, microflora and diverse rich ecosystems of one’s skin, I would edit specific pigment biosynthesis in microbial genes to produce select pigments, creating chromatic responses in a textile. I would insert promoters responsive to environmental factors such as pH, humidity, or metabolite concentration. The engineered microbes could then produce pigment in dynamic gradiations according to the wearer and their environment. This design interface approach leverages gene editing as a translational medium for expressing microworlds as a visual living and smart material.
(ii) What technology or technologies would you use to perform these DNA edits and why? Also answer the following questions:
To perform this gene edit, the use of a CRISPR-Cas9 system would be needed for pigment biosynthesis. Crispr Cas9 can offer precise base pair editing of promoter regions, tuning expression without breaking the double strand. Editing machinery into bacterial cells could be achieved via electroporation or conjugative plasmids. Inducing pigment production responsive to the environment, synthetic gene circuits would be engineered through inducible promoters or quorum-sensing systems. PCR sequences would also need to be validated edited strains before integrating into a bioresponsive textile system.
How does your technology of choice edit DNA? What are the essential steps? What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
Cas9 is an RNA-guided endonuclease which induces a double-strand break at a target DNA sequence chosen by the complementary guide RNA and recircularized PM site. The Cas9 enzyme then cuts the DNA in the target site. The cell then repairs the cut, and DNA sequences can be changed during the repair by disrupting the gene or inserting a new sequence with a provided repair template. For the editting process I would first need to design an RNA reciprocale with the target gene and ensure a PAM site is close to the target. The input would contain a plasmid including the CAS9, guide RNA, bacterial cells and the repair template. The process would first include the plasmid construction, introducing the plasmid into cells, CRISPR cutting the DNA, and lastly selecting the successful edits of the cell.
What are the limitations of your editing methods (if any) in terms of efficiency or precision?
One limitation of CRISPR-Cas9 editing tools is the potential for off-target effects, in which the guide RNA partially binds to unintended genomic regions, leading to unintended DNA cuts. Also, as this technology requires a PAM site near the target site, it limits the locations that can be edited. The cell’s DNA repair mechanisms can also reduce efficiency, as homology-directed repair may often be unpredictable and can require screening of multiple clones. Efficiency can also be reduced by the delivery of CRISPR components into bacterial cells, especially in non-model organisms. Lastly, viability may be reduced by continuous double-strand breakage, which can be toxic to cells and thus reduce viability.
RESOURCES:
Cai, T., Han, K., Yang, P., Zhu, Z., Jiang, M., Huang, Y., Xie, C. (2019) Reconstruction of dynamic and reversible color change using reflectin protein. Scientific Reports, 9, 5201. Available at: https://www.nature.com/articles/s41598-019-41638-8
USE OF AI
“Polish/Edit this paragraph to my conceptual idea”