Week 6 HW: Genetic Circuits Part I: Assembly Technologies
DNA ASSEMBLY
Answer these questions about the protocol in this week’s lab:
What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The Phusion High-Fidelity PCR Master Mix is a pre-mix solution used to amplify DNA during PCR. There are a few components that make the master mix efficient and accurate.
Phusion High-Fidelity DNA Polymerase synthesises new DNA strands by adding new nucleotides to the primer and has very low error rates with proof-reading abilities from (3’—>5’)exonuclease. The mix also includes a reaction buffer that maintains the optimal pH, salt concentration, and ionic environment for the polymerase and stabilises it during PCR. Mg²⁺ ions, magnesium ions, also help stabilise the reaction and enable enzyme and nucleotide catalysis during DNA synthesis (Thermo Fisher Scientific).
What are some factors that determine primer annealing temperature during PCR?
The primer annealing temperature is primarily determined by the primers’ melting temperature (Tm). At least 50% of the primers need to melt at this temperature. Primers containing a high content of GC require higher melting temperatures because G-C base pairs have 3 hydrogen bonds, while A-T only have two. Longer primers also have higher melting temperatures because they have more nucleotides to denature. Salt and ion concentration also affect the annealing temperature, with higher salt concentration increasing binder stability and thus requiring a higher Tm. Lastly, primers with more complementarity have a higher Tm, whereas mismatches have weaker binding and thus don’t require a high Tm.
There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR and restriction enzyme digestion are both methods for generating linear DNA fragments, but they differ significantly in their mechanisms, protocols, and applications.
PCR (Polymerase Chain Reaction) is an amplification-based method using primers, a DNA polymerase, and thermal cycling to selectively copy specific regions of a DNA. The steps involve repeated cycles of denaturation, primer annealing, and extension, resulting in exponential amplification of the target sequence. PCR is highly specific, and primers can be designed to amplify almost any sequence. Additionally, PCR enables the introduction of mutations or additional sequences via primer design (Addgene).
In contrast, restriction enzyme digestion relies on enzymes to recognise specific short DNA sequences and cleave the DNA at or near these sites. The protocol is relatively simple, typically involving incubation of DNA with one or more restriction enzymes under appropriate buffer and temperature conditions. This method does not amplify DNA; instead, it cuts existing DNA into predictable fragments based on the locations of restriction sites (PubMed Central).
In terms of preferred methods, PCR is ideal when a specific DNA fragment needs to be amplified, when the starting DNA is limited, or when sequence modification is required. On the other hand, restriction digestion is preferable when working with larger quantities of DNA, and when precise, reproducible cutting at known sequences is needed, for example, in cloning where compatible ends are required for ligation (Addgene). Overall, PCR offers high specificity and flexibility, while restriction digestion provides precision and simplicity when suitable recognition sites are already present.
How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
To ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning, the fragments must have homologous ends. Gibson assembly relies on 20 - 40 base pair overlaps between adjacent DNA fragments, which enables them to anneal and be joined seamlessly.
How does the plasmid DNA enter the E. coli cells during transformation?
There are two main methods in which plasmid DNA enter E. coli cells during transformation. The first and most common method amongst labs is Heat shock, where the membrane of the cell becomes permeable by first being treated with calcium chloride (CaCl₂) and then “heat shocked” for 30-60 seconds at a temperature of 42 degrees Celsius. The DNA is then pulled into the cell and after enters a repaired and expressed plasmid gene.
The second and most efficient method is electroporation, in which cells and DNA are placed in a cuvette and a short, high-voltage pulse is applied, creating tiny pores in the membrane. The DNA then enters through these pores and then reseals.
Describe another assembly method in detail (such as Golden Gate Assembly)
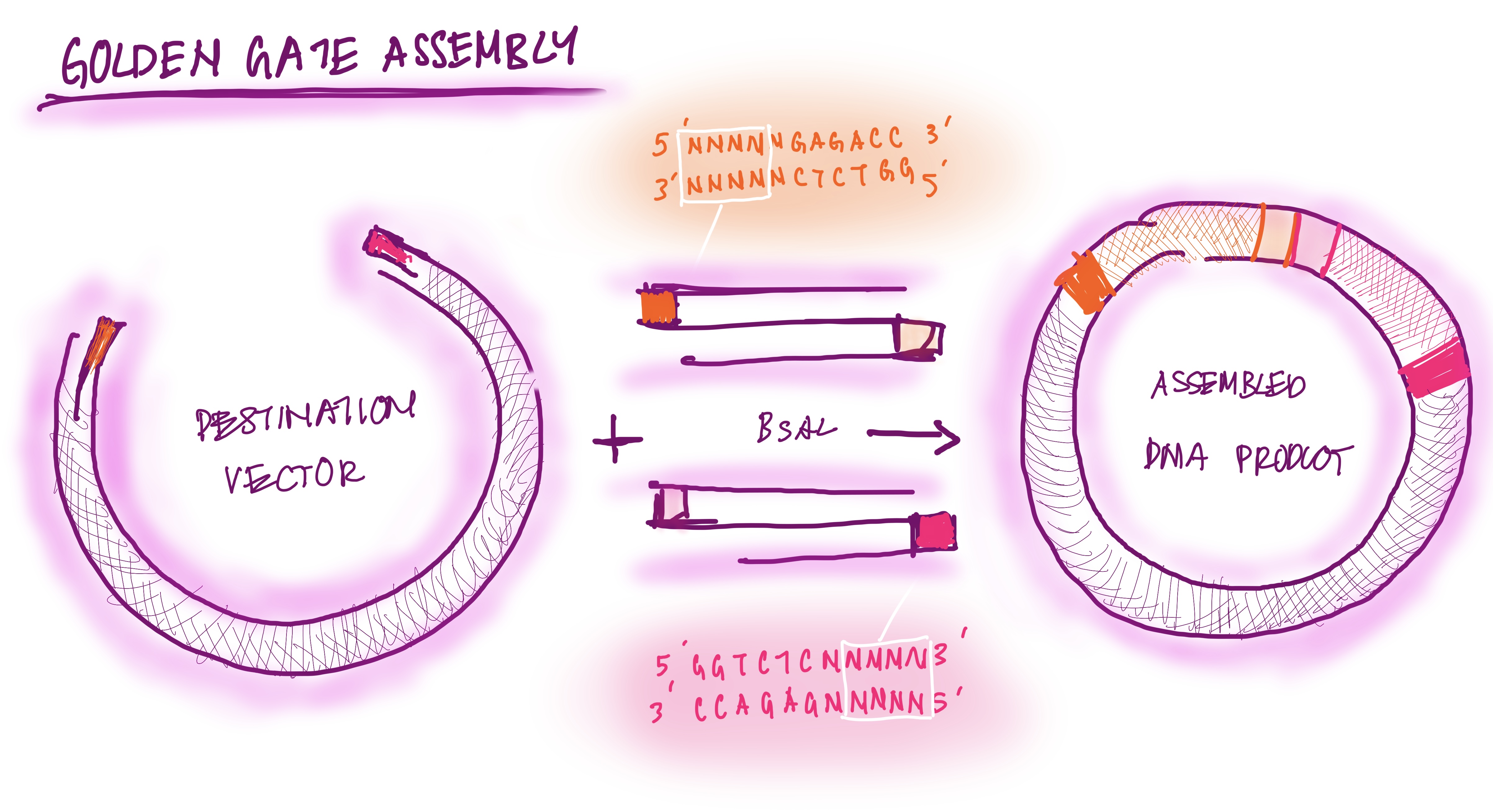
The Golden Gate assembly is a process for molecular cloning where DNA fragments are bound with ISS restriction enzymes and DNA ligase in a single seamless reaction. The first type of ISS restriction enzyme cuts the DNA outside the recognition site, creating sticky sites (overhangs). The DNA fragments are designed to have reciprocal overhangs. DNA ligase connects matching overhang ends, forming continuous DNA strands. The reaction alternates cutting and ligating. This assembly process is efficient because it can assemble multiple fragments at once and avoids waste of DNA base pairs; DNA is “scarless” (New England Biolabs).
GOLDEN GATE ASSEMBLY DIAGRAM DRAWING

RESOURCES
Addgene, n.d. PCR Protocol. Available at: https://www.addgene.org/protocols/pcr/ (Accessed: 10 March 2026).
New England Biolabs. “Golden Gate Assembly Domestication Tutorial.” YouTube, 26 Mar. 2020, https://www.youtube.com/watch?v=JuiJASyG9gc
PubMed Central, n.d. Polymerase Chain Reaction (PCR). Available at: https://pmc.ncbi.nlm.nih.gov/articles/PMC4846334/ (Accessed: 29 March 2026).
Thermo Fisher Scientific, n.d. Phusion High-Fidelity PCR Master Mix. Available at: https://www.thermofisher.com/order/catalog/product/F531S (Accessed: 13 March 2026).
USE OF AI
Golden Gate Assembly Drawing: prompt “feedback on drawing of the Golden Gate assembly”
“Edit my answer to this question”