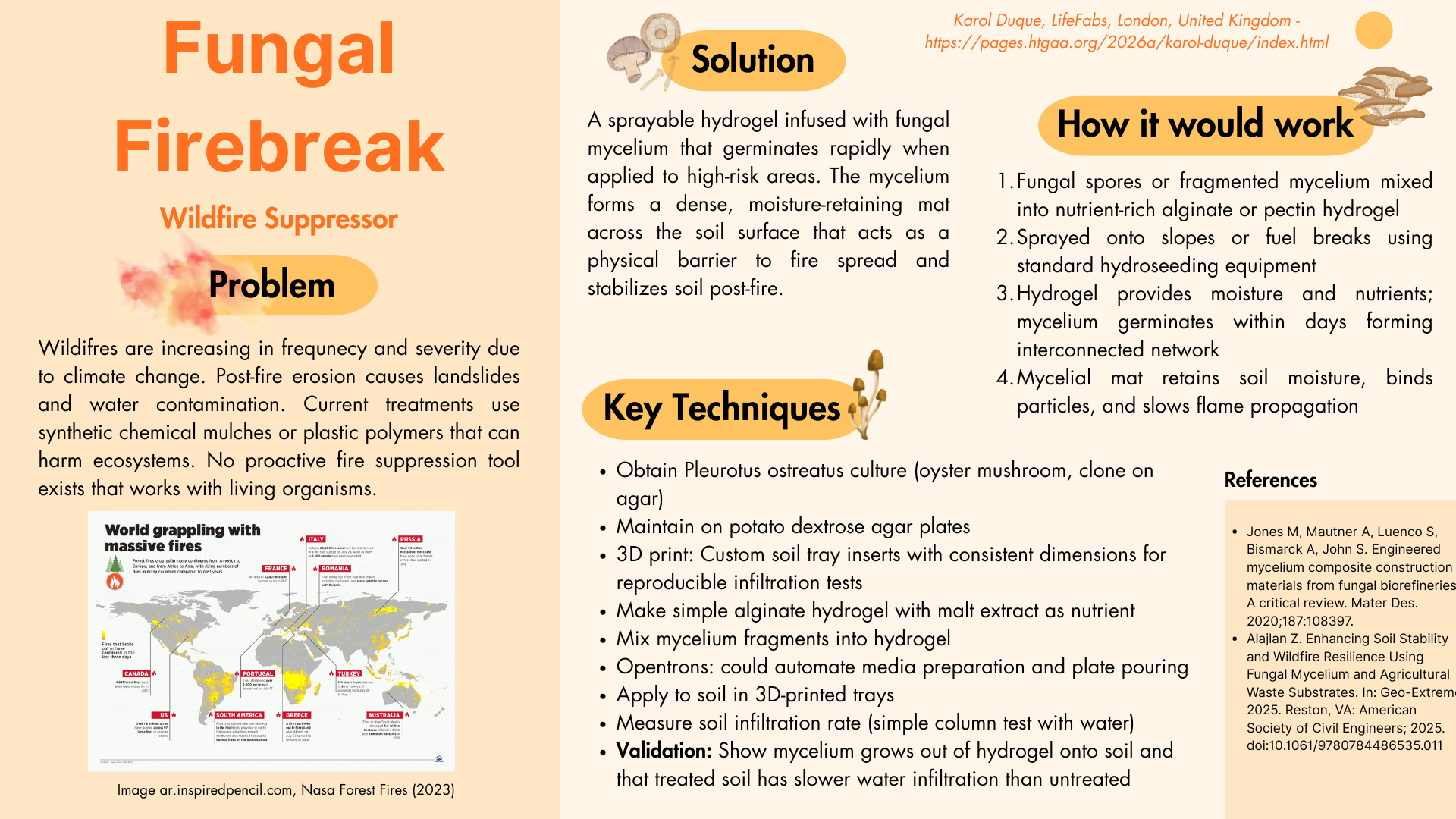
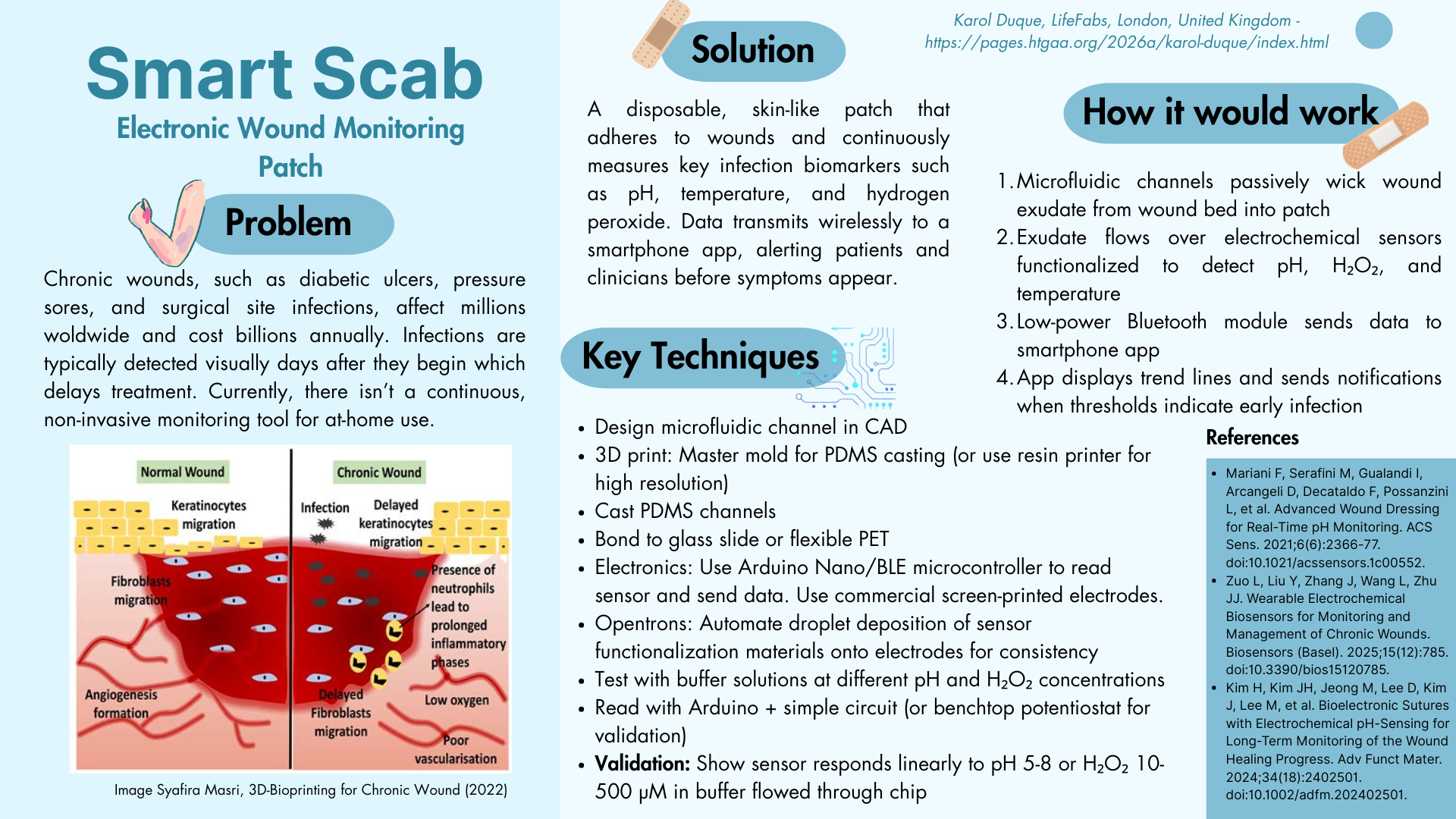
The colors were achieved using eqFP578 (blue), TagRFP (pink), and mCherry (purple.)
Then, using the coordinates from the GUI, I followed the instructions in the HTGAA26 Opentrons Colab to write my own Python script with the assistance of Claude (Anthropic) which helped me structure the pipetting logic. This is the code:
from opentrons import types
metadata = { # see https://docs.opentrons.com/v2/tutorial.html#tutorial-metadata
'author': 'Karol Duque',
'protocolName': 'Fluorescent Protein Scene',
'description': 'Three-color artistic scene using fluorescent bacteria: '
'mCherry (purple under UV) in A1, '
'TagRFP (pink under UV) in B1, '
'EqFP578 (blue under UV) in C1. '
'Coordinates generated at opentrons-art.rcdonovan.com. '
'AI assistance: Claude (Anthropic) helped structure the pipetting logic.',
'source': 'HTGAA 2026 Opentrons Lab',
'apiLevel': '2.20'
}
##############################################################################
### Robot deck setup constants - don't change these
##############################################################################
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
# Using CSS color names that closely match actual fluorescence appearance:
# mCherry → 'darkorchid' (fluoresces purple)
# TagRFP → 'hotpink' (fluoresces pink)
# EqFP578 → 'dodgerblue' (fluoresces blue)
#
# Confirm physical well positions with your TA before running!
well_colors = {
'A1' : 'darkorchid', # mCherry
'B1' : 'hotpink', # TagRFP
'C1' : 'navy', # EqFP578
}
def run(protocol):
##############################################################################
### Load labware, modules and pipettes
##############################################################################
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
temperature_module = protocol.load_module('temperature module gen2', COLORS_DECK_SLOT)
temperature_plate = temperature_module.load_labware('opentrons_96_aluminumblock_generic_pcr_strip_200ul', 'Cold Plate')
color_plate = temperature_plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate') ## TA MUST CALIBRATE EACH PLATE!
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
##############################################################################
### Patterning
##############################################################################
def location_of_color(color_string):
for well, color in well_colors.items():
if color.lower() == color_string.lower():
return color_plate[well]
raise ValueError(f"No well found with color {color_string}")
def dispense_and_detach(pipette, volume, location):
assert(isinstance(volume, (int, float)))
above_location = location.move(types.Point(z=location.point.z + 5))
pipette.move_to(above_location)
pipette.dispense(volume, location)
pipette.move_to(above_location)
###
### YOUR CODE HERE to create your design
###
# Coordinates from opentrons-art.rcdonovan.com
# mCherry → 'darkorchid' well A1 (fluoresces purple)
# TagRFP → 'hotpink' well B1 (fluoresces pink)
# EqFP578 → 'navy' well C1 (fluoresces blue)
mcherry_points = [(-5, 25),(5, 25),(-20, 20),(10, 20),(-12.5, 17.5),(-27.5, 15),(-25, 15),(-30, 12.5),(-27.5, 12.5),(-25, 12.5),(-22.5, 12.5),(27.5, 12.5),(-30, 10),(-27.5, 10),(-25, 10),(-22.5, 10),(-20, 10),(-17.5, 10),(-15, 10),(25, 10),(27.5, 10),(-32.5, 7.5),(-30, 7.5),(-27.5, 7.5),(-25, 7.5),(-22.5, 7.5),(-17.5, 7.5),(-15, 7.5),(-12.5, 7.5),(-10, 7.5),(0, 7.5),(2.5, 7.5),(5, 7.5),(7.5, 7.5),(10, 7.5),(12.5, 7.5),(22.5, 7.5),(25, 7.5),(27.5, 7.5),(30, 7.5),(-32.5, 5),(-27.5, 5),(-25, 5),(-22.5, 5),(-20, 5),(-17.5, 5),(-15, 5),(-12.5, 5),(-10, 5),(-5, 5),(-2.5, 5),(0, 5),(2.5, 5),(10, 5),(12.5, 5),(15, 5),(17.5, 5),(20, 5),(22.5, 5),(27.5, 5),(30, 5),(-32.5, 2.5),(-30, 2.5),(-27.5, 2.5),(-25, 2.5),(-22.5, 2.5),(-17.5, 2.5),(-15, 2.5),(-12.5, 2.5),(-10, 2.5),(-7.5, 2.5),(-5, 2.5),(-2.5, 2.5),(2.5, 2.5),(12.5, 2.5),(15, 2.5),(17.5, 2.5),(20, 2.5),(22.5, 2.5),(25, 2.5),(27.5, 2.5),(30, 2.5),(-32.5, 0),(-30, 0),(-27.5, 0),(-25, 0),(-22.5, 0),(-20, 0),(-17.5, 0),(-15, 0),(-12.5, 0),(-10, 0),(-5, 0),(-2.5, 0),(2.5, 0),(10, 0),(15, 0),(17.5, 0),(20, 0),(22.5, 0),(25, 0),(-20, -2.5),(-17.5, -2.5),(-12.5, -2.5),(-10, -2.5),(-7.5, -2.5),(-5, -2.5),(-2.5, -2.5),(2.5, -2.5),(17.5, -2.5),(-7.5, -5),(-5, -5)]
tagrfp_points = [(-10, 15),(10, 15),(-7.5, 12.5),(12.5, 12.5),(-20, 7.5),(-30, 5),(25, 5),(-20, 2.5),(10, 2.5),(-7.5, 0),(27.5, 0),(30, 0),(-32.5, -2.5),(-30, -2.5),(-27.5, -2.5),(-25, -2.5),(-22.5, -2.5),(-15, -2.5),(10, -2.5),(15, -2.5),(20, -2.5),(22.5, -2.5),(25, -2.5),(27.5, -2.5),(30, -2.5),(-32.5, -5),(-30, -5),(-27.5, -5),(-25, -5),(-22.5, -5),(-20, -5),(-17.5, -5),(-15, -5),(-12.5, -5),(-10, -5),(-2.5, -5),(10, -5),(15, -5),(17.5, -5),(22.5, -5),(25, -5),(27.5, -5),(30, -5),(-32.5, -7.5),(-30, -7.5),(-27.5, -7.5),(-22.5, -7.5),(-20, -7.5),(-17.5, -7.5),(-12.5, -7.5),(-10, -7.5),(-7.5, -7.5),(-5, -7.5),(-2.5, -7.5),(15, -7.5),(17.5, -7.5),(25, -7.5),(27.5, -7.5),(30, -7.5),(-32.5, -10),(-30, -10),(-20, -10),(-17.5, -10),(-12.5, -10),(-7.5, -10),(-5, -10),(-2.5, -10),(0, -10),(2.5, -10),(15, -10),(22.5, -10),(25, -10),(27.5, -10),(30, -10),(-30, -12.5),(-20, -12.5),(-12.5, -12.5),(-7.5, -12.5),(-5, -12.5),(-2.5, -12.5),(0, -12.5),(2.5, -12.5),(12.5, -12.5),(15, -12.5),(17.5, -12.5),(22.5, -12.5),(27.5, -12.5),(-30, -15),(-27.5, -15),(-22.5, -15),(-20, -15),(-17.5, -15),(-7.5, -15),(-5, -15),(-2.5, -15),(0, -15),(2.5, -15),(10, -15),(12.5, -15),(15, -15),(17.5, -15),(-20, -17.5),(-17.5, -17.5),(-10, -17.5),(-7.5, -17.5),(-5, -17.5),(0, -17.5),(2.5, -17.5),(10, -17.5),(-17.5, -20),(-12.5, -20),(-10, -20),(-7.5, -20),(-10, -22.5)]
eqfp578_points = [(-10, 30),(-7.5, 30),(-5, 30),(-2.5, 30),(0, 30),(2.5, 30),(5, 30),(7.5, 30),(-15, 27.5),(-12.5, 27.5),(-10, 27.5),(-7.5, 27.5),(-5, 27.5),(-2.5, 27.5),(0, 27.5),(2.5, 27.5),(5, 27.5),(7.5, 27.5),(10, 27.5),(12.5, 27.5),(-20, 25),(-17.5, 25),(-15, 25),(-12.5, 25),(-10, 25),(-7.5, 25),(-2.5, 25),(0, 25),(2.5, 25),(7.5, 25),(10, 25),(12.5, 25),(15, 25),(17.5, 25),(-22.5, 22.5),(-20, 22.5),(-17.5, 22.5),(-15, 22.5),(-12.5, 22.5),(-10, 22.5),(-7.5, 22.5),(-5, 22.5),(-2.5, 22.5),(0, 22.5),(2.5, 22.5),(5, 22.5),(7.5, 22.5),(10, 22.5),(12.5, 22.5),(15, 22.5),(17.5, 22.5),(20, 22.5),(-25, 20),(-22.5, 20),(-17.5, 20),(-15, 20),(-12.5, 20),(-10, 20),(-7.5, 20),(-5, 20),(-2.5, 20),(0, 20),(2.5, 20),(5, 20),(7.5, 20),(12.5, 20),(15, 20),(17.5, 20),(20, 20),(22.5, 20),(-27.5, 17.5),(-25, 17.5),(-22.5, 17.5),(-20, 17.5),(-17.5, 17.5),(-15, 17.5),(-10, 17.5),(-7.5, 17.5),(-5, 17.5),(-2.5, 17.5),(0, 17.5),(2.5, 17.5),(5, 17.5),(7.5, 17.5),(10, 17.5),(12.5, 17.5),(15, 17.5),(17.5, 17.5),(20, 17.5),(22.5, 17.5),(25, 17.5),(-22.5, 15),(-20, 15),(-17.5, 15),(-15, 15),(-12.5, 15),(-7.5, 15),(-5, 15),(-2.5, 15),(0, 15),(2.5, 15),(5, 15),(7.5, 15),(12.5, 15),(15, 15),(17.5, 15),(20, 15),(22.5, 15),(25, 15),(-20, 12.5),(-17.5, 12.5),(-15, 12.5),(-12.5, 12.5),(-10, 12.5),(-5, 12.5),(-2.5, 12.5),(0, 12.5),(2.5, 12.5),(5, 12.5),(7.5, 12.5),(10, 12.5),(15, 12.5),(17.5, 12.5),(20, 12.5),(22.5, 12.5),(25, 12.5),(-12.5, 10),(-10, 10),(-7.5, 10),(-5, 10),(-2.5, 10),(0, 10),(2.5, 10),(5, 10),(7.5, 10),(10, 10),(12.5, 10),(15, 10),(17.5, 10),(20, 10),(22.5, 10),(-7.5, 7.5),(-5, 7.5),(-2.5, 7.5),(15, 7.5),(17.5, 7.5),(20, 7.5),(-7.5, 5),(5, 5),(7.5, 5),(0, 2.5),(5, 2.5),(7.5, 2.5),(0, 0),(5, 0),(7.5, 0),(12.5, 0),(0, -2.5),(5, -2.5),(7.5, -2.5),(12.5, -2.5),(0, -5),(2.5, -5),(5, -5),(7.5, -5),(12.5, -5),(20, -5),(-25, -7.5),(-15, -7.5),(0, -7.5),(2.5, -7.5),(5, -7.5),(7.5, -7.5),(10, -7.5),(12.5, -7.5),(20, -7.5),(22.5, -7.5),(-27.5, -10),(-25, -10),(-22.5, -10),(-15, -10),(-10, -10),(5, -10),(7.5, -10),(10, -10),(12.5, -10),(17.5, -10),(20, -10),(-27.5, -12.5),(-25, -12.5),(-22.5, -12.5),(-17.5, -12.5),(-15, -12.5),(-10, -12.5),(5, -12.5),(7.5, -12.5),(10, -12.5),(20, -12.5),(25, -12.5),(-25, -15),(-15, -15),(-12.5, -15),(-10, -15),(5, -15),(7.5, -15),(20, -15),(22.5, -15),(25, -15),(27.5, -15),(-27.5, -17.5),(-25, -17.5),(-22.5, -17.5),(-15, -17.5),(-12.5, -17.5),(-2.5, -17.5),(5, -17.5),(7.5, -17.5),(12.5, -17.5),(15, -17.5),(17.5, -17.5),(20, -17.5),(22.5, -17.5),(25, -17.5),(-27.5, -20),(-25, -20),(-22.5, -20),(-20, -20),(-15, -20),(-5, -20),(-2.5, -20),(0, -20),(2.5, -20),(5, -20),(7.5, -20),(10, -20),(12.5, -20),(15, -20),(17.5, -20),(20, -20),(22.5, -20),(25, -20),(-25, -22.5),(-22.5, -22.5),(-20, -22.5),(-17.5, -22.5),(-15, -22.5),(-12.5, -22.5),(-7.5, -22.5),(-5, -22.5),(-2.5, -22.5),(0, -22.5),(2.5, -22.5),(5, -22.5),(7.5, -22.5),(10, -22.5),(12.5, -22.5),(15, -22.5),(17.5, -22.5),(20, -22.5),(22.5, -22.5),(-22.5, -25),(-20, -25),(-17.5, -25),(-15, -25),(-12.5, -25),(-10, -25),(-7.5, -25),(-5, -25),(-2.5, -25),(0, -25),(2.5, -25),(5, -25),(7.5, -25),(10, -25),(12.5, -25),(15, -25),(17.5, -25),(20, -25),(-20, -27.5),(-17.5, -27.5),(-15, -27.5),(-12.5, -27.5),(-10, -27.5),(-7.5, -27.5),(-5, -27.5),(-2.5, -27.5),(0, -27.5),(2.5, -27.5),(5, -27.5),(7.5, -27.5),(10, -27.5),(12.5, -27.5),(15, -27.5),(17.5, -27.5),(-15, -30),(-12.5, -30),(-10, -30),(-7.5, -30),(-5, -30),(-2.5, -30),(0, -30),(2.5, -30),(5, -30),(7.5, -30),(10, -30),(12.5, -30),(-10, -32.5),(-7.5, -32.5),(-5, -32.5),(-2.5, -32.5),(0, -32.5),(2.5, -32.5),(5, -32.5),(7.5, -32.5)]
def draw_points(points, color_string):
"""One tip per color. Aspirates in batches of 20uL, dispenses 1uL per dot."""
pipette_20ul.pick_up_tip()
i = 0
while i < len(points):
batch = min(20, len(points) - i)
pipette_20ul.aspirate(batch, location_of_color(color_string))
for _ in range(batch):
x, y = points[i]
loc = center_location.move(types.Point(x=x, y=y))
dispense_and_detach(pipette_20ul, 1, loc)
i += 1
pipette_20ul.drop_tip()
# Draw order: blue background first, then purple (mCherry), then pink (TagRFP) on top
draw_points(eqfp578_points, 'navy') # EqFP578 → blue
draw_points(mcherry_points, 'darkorchid') # mCherry → purple
draw_points(tagrfp_points, 'hotpink') # TagRFP → pink
# drop_tip() is called inside draw_points() for each color
This idea is ba continuation on the research I did for the homework in Weeks 1 and 2.
This idea came about after reading about the recent wildfires in Patagonia, Argentina and further researching recent wildfires across the world including Chile, Spain and Portugal. I started thinking about the ways that already exist to prevent wildfires and thought if there were any methods that use modified living organisms.
Because my background is in biomedical engineering I thought I should have at least one project idea that was related healthcare. Before starting my degree I did some research into bandaids made with hydrogels which act like stitches and prevent infection. Last summer, I did an internship which used microfluidics for blood glucose monitoring which made me think if there was any way I could combine microfluidics and something to improve wound care. That led me to chronic wounds and I began researching monitoring methods for these wounds that led me to biosensors for chronic wound management.
I read the following paper: “Semiautomated Production of Cell-Free Biosensors” (Brown DM, Phillips DA, Garcia DC, et al. ACS Synthetic Biology. 2025;14(3):979-986.)
The researchers used an Opentrons OT-2 to automate the assembly of cell-free biosensors. They compared manual vs. robotic assembly of biosensors that produce colorimetric (LacZ) or fluorescent (GFP) signals. Using the robot, they successfully constructed an entire 384-well plate of fluoride-sensing biosensors with consistent performance. They noted that manual assembly leads to quality control and performance variability issues. Their robotic workflow produced biosensors that performed close to expected detection outcomes proving that Opentrons can manufacture point-of-care diagnostic devices at scale.
For my Smart Scab project, I will use automation tools to manufacture and optimize cell-free biosensors that detect bacterial quorum sensing molecules. My plan has four main or potential components. Inspired by Brown et al.’s Semiautomated Production of Cell-Free Biosensors, I could use the OT-2 to assemble 384-well plates of cell-free reactions with different concentrations of quorum sensing molecules (AHLs for Pseudomonas, AIPs for S. aureus). This would allow me to generate full dose-response curves in 15 minutes rather than 3 hours manually. This is an example Pytho pseudocode:
APEX’s spreadsheet-based configuration means I could run this without advanced coding skills, I simply fill CSV files with source wells and transfer volumes.
I would also design and 3D print several custom adapters, including an agar plate adapter, a hydrogel mold, and deck riser. The agar plate adapter positions 90 mm Petri dishes on the OT-2 deck for automated colony sampling (following APEX’s design). The hydrogel mold creates uniform 8 mm diameter, 2 mm thick alginate discs that fit into the patch housing, ensuring consistent sensor deposition. And the deck riser allows stacking multiple 384-well plates to increase throughput beyond the OT-2’s standard 4-plate capacity.
Finally, once my OT-2 protocols identify the best-performing protein variants, I would scale up using Ginkgo Cloud Lab. That would give me access to over 70 automated instruments including liquid handlers, incubators, and plate readers.
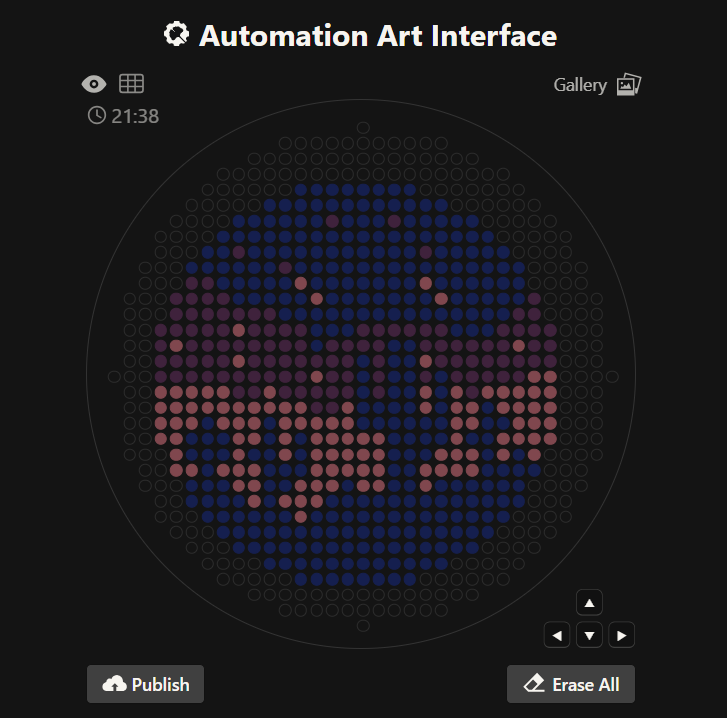
 Figure 1. Night sky in a desert (Automation Art Interference GUI).
Figure 1. Night sky in a desert (Automation Art Interference GUI).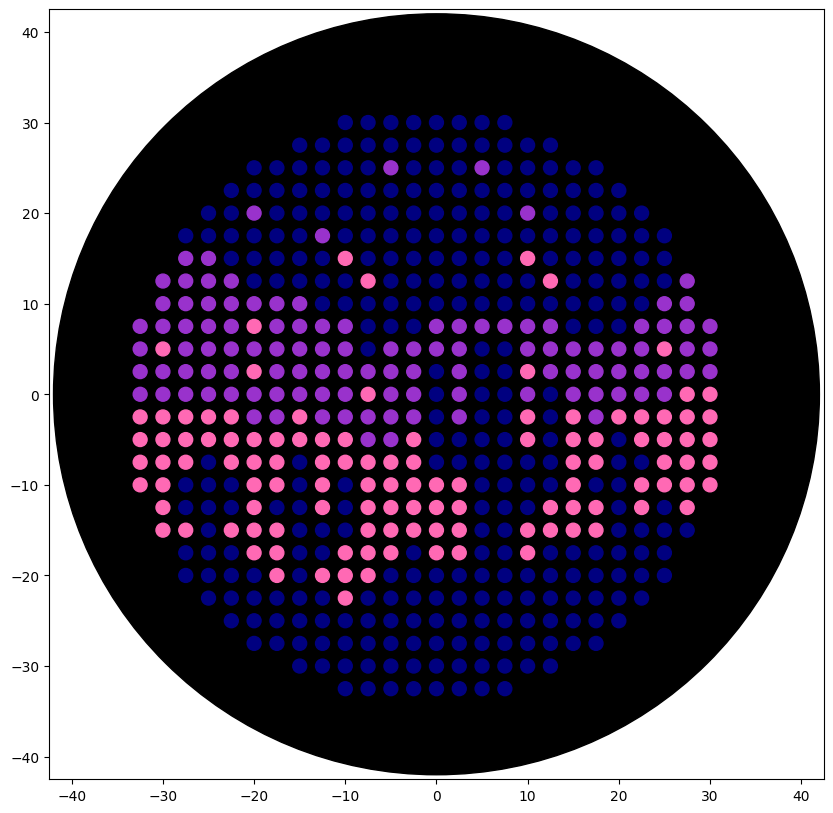
 Figure 2. Night sky in a desert (Opentrons Colab).
Figure 2. Night sky in a desert (Opentrons Colab).