Week 7: Genetic Circuits Part II - Neuromorphic Circuits
Part 1: Intracellular Artificial Neural Networks (IANNs)
- What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
Traditional genetic circuits implement Boolean logic using transcription factors, promoters, and other regulatory elements. Their input/output behavior is inherently digital (ON/OFF), and they often suffer from limited computational capacity, scalability issues, and lack of analog processing. Only simple logic functions (AND, OR, NOT) can be composed. Wiring many gates leads to metabolic burden, crosstalk, and slow response times. They also cannot perform weighted sums or continuous transformations.
Intracellular Artificial Neural Networks (IANNs) overcome these limitations by:
- Using graded expression levels (e.g., of endoribonucleases or transcription factors) to represent continuous values, enabling weighted sums and non-linear activation functions.
- Components (e.g., RNA-binding proteins, ribozymes, or split enzymes) can be combined with less crosstalk than transcription-factor‑based circuits.
- IANNs can be trained (e.g., via directed evolution or feedback control) to approximate complex input‑output mappings, while traditional genetic circuits require hardwired logic. RNA‑based regulation (e.g., Csy4 endoribonuclease) often operates post‑transcriptionally, decoupling from the transcriptional machinery and reducing metabolic load.
- Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
A practical application is an IANN‑based early warning sensor for Nidularia pulvinata, an invasive scale insect, Nidularia pulvinata, currently forcing the felling of dozens of 100‑year‑old holm oaks at Brompton Cemetery, London. The sensor integrates volatile organic compounds (VOCs) from infested trees and honeydew signatures, using a multilayer perceptron to output a fluorescence risk score. This allows targeted intervention before tree health declines irreversibly and addresses a gap where no treatment is currently available. The only current measure is to remove infested trees to slow the spread. Early detection at the edge of infestation zones could enable targeted quarantine and preserve healthy trees.
A deployable biosensor, housed in a microfluidic cartridge, uses engineered bacteria to integrate multiple chemical signatures of infestation. The IANN’s output is a fluorescence intensity that correlates with infestation probability.
Each input is sensed by a specific promoter driving expression of an endoribonuclease (e.g., Csy4):
| Input | Chemical Signature | Sensing Mechanism |
|---|---|---|
| X₁ | (E)-β‑ocimene (stress volatile from infested oaks) | Oak‑responsive promoter (e.g., from Pseudomonas spp.) |
| X₂ | Methyl salicylate (another stress volatile) | Engineered two‑component system |
| X₃ | Honeydew sugars (excreted by scale insects) | Sugar‑binding protein fused to a transcriptional activator |
| X₄ | Ethanol (from early wood decay) | Ethanol‑inducible promoter (e.g., PadhE) |
Each promoter drives production of a distinct endoribonuclease (e.g., Csy4, Cas6, or orthogonal variants). The concentration of each endoribonuclease is proportional to the concentration of the corresponding chemical cue.
IANN Architecture (Multilayer Perceptron):
- Layer 1 – Endoribonucleases are produced. They target specific RNA sequences in the 5’ UTR of genes encoding intermediate transcription factors (layer 2 regulators). Cleavage of those RNAs reduces translation of the intermediate factors, implementing a weighted sum with inhibitory weights.
- Layer 2 – The intermediate transcription factors drive expression of a fluorescent protein (e.g., sfGFP). The combined regulation produces a non‑linear (sigmoidal) output: fluorescence is low under background stress, but rises sharply when the input pattern matches an active infestation.
Fluorescence intensity is read by a simple electronic photodiode or a handheld UV lamp. A positive reading triggers further inspection (e.g., bark peeling or trapping) and allows early quarantine measures.
Limitations:
VOC profiles may overlap with drought or other pests. The IANN must be trained to distinguish Nidularia signatures. This can be done by directed evolution of promoter–endoribonuclease pairs against authentic volatiles.
Engineered bacteria must survive outdoor conditions. Encapsulation in hydrogels or silica, plus integration into a sealed microfluidic chip, protects them from UV, temperature swings, and humidity.
The cartridge must contain the bacteria (e.g., with a kill‑switch or physical separation) to prevent release into the environment.
Transcription, translation, and RNA cleavage take minutes to hours. For a trap‑based or periodic sampling system, this is acceptable; real‑time monitoring would require faster components.
Variability between batches requires pre‑deployment calibration. A constitutively expressed reference dye (e.g., mCherry) can be included to normalise output.
Below is a diagram depicting an intracellular single-layer perceptron where the X1 input is DNA encoding for the Csy4 endoribonuclease and the X2 input is DNA encoding for a fluorescent protein output whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation. Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
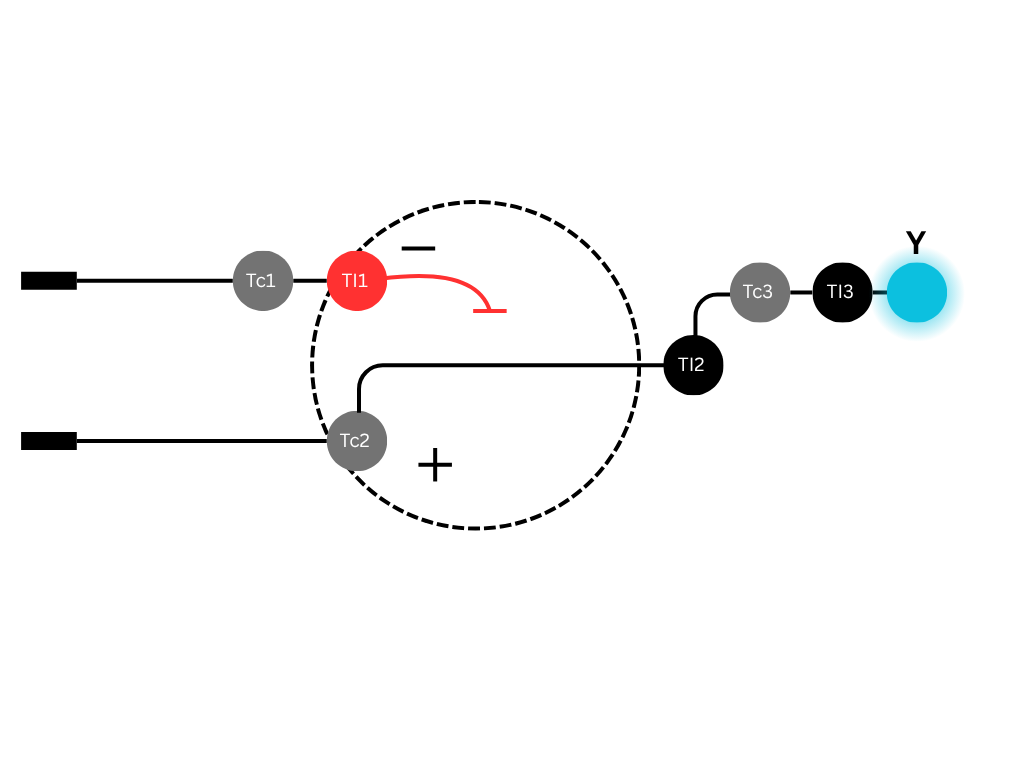

The diagram shows two DNA strands (Layer 1 and Layer 2) with transcription (Tc) and translation (TI) symbols, and the regulatory interaction between layers. Some annotations:
- Grey “Tc” circles represent transcription (RNA polymerase binding).
- Red “TI” circles represent translation (ribosome binding).
- A dashed circle encloses the endoribonucleases with a “−” sign indicating repressive regulation (RNA cleavage).
- The second‑layer transcription factor (TF) is produced only if the endoribonucleases are below a threshold; it then drives output.
- This implements a two‑layer perceptron where layer 1 performs weighted summation (via cleavage rates) and layer 2 applies a non‑linear activation (via TF‑mediated transcription).
The bottom DNA strand has two parts, a transcription factor (TF) gene that is regulated by the endoribonuclease, and a fluorescent protein (Y) gene that is activated by the TF. TI2 is the translation for the TF, and the mRNA of that TF gene is the target of the endoribonuclease.
Part 2: Fungal Materials
- What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Fungi are being used to create sustainable alternatives to conventional materials. An example are mycelium composites. These are dried, heat-treated mycelium grown on agricultural waste such as hemp and sawdust. These are used for packaging, insulation, furniture and even building blocks. ITs advantages include being biodegradable, they are grown from waste feedstocks, are fire-retardant and offer good insulation. Additionally, it can be moulded into complex shapes. Some disadvantages include a lower mechanical strength than plastic or wood, sensitivity to moisture, and slower production than injection moulding. Another example is fungal building materials which are similar in that they are mycelium-brick composites, such as Ganoderma lucidum with rice hulls, and self-repairing fungal-infused concrete. This is mainly used in construction, insulation and temporary structures, but it can also be used in self-healing concrete for infrastructure. Some advantages include being able to heal cracks in concrete, offering excellent insulation value, and being renewable. However, as before, these materials require careful moisture control, and currently the structural strength is insufficient for load-bearing wall.
- What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
You might want to genetically engineer fungi as fungal biosensors for environmental monitoring. For example, you could engineer a mushroom to bioluminesce in response to specific pollutants. The mushrooms would act as a living sentinel in forests, farms or urban parks. Anyone passing by might see this glow and report it, which would make it an early-warning system. In general, fungi offer eukaryotic proteing-processing power, natural secretion, and the ability to form structured materials. So, they can be used to create functional lviing materials, biosensors, and sustainable products. Bacteria are easier to engineer for fast, intracellular tasks but for complex proteins they’re not ideal.
There are several categories we can use to compare fungi and bacteria. The first is the secretion capacity. Fungi naturally secrete large amounts of enzymes and proteins which is ideal for producing materials or enzymes extracellularly. Bacteria have some secretion systems but they often require engineering. Additionally, fungi have filamentous growth which allows them to colonize solid substrates like wood, soil, and textiles, and form 3D structures, whereas bacteria typically grow in liquid or as surface biofilms. Bacteria are mostly unicellular and less suited for forming macroscopic materials directly. Also, fungi naturally degrade lignocellulose, produce antibiotics, and perform complex secondary metabolism. They can be engineered for novel metabolic pathways using existing backbones. Bacteria also have rich metabolism, but for plant‑based feedstocks, fungi often have an edge. Furthermore, Many fungi are GRAS (e.g., Aspergillus oryzae, Saccharomyces cerevisiae) and can be engineered with built‑in kill switches. Filamentous fungi are less likely to spread uncontrollably compared to some bacteria. However, some bacteria are pathogenic, which means containment can be more challenging in open environments. Lastly, Fermentation and solid‑state cultivation are well established in fungi and it can be grown on agricultural waste, reducing cost. Bacterial fermentation is also scalable but often requires purified sugars.