Week 2 Lab: DNA Gel Art
Part 1. Benchling & In-silico Gel Art
This week’s lab followed the protocol detailed in “Gel Art: Restriction Digests and Gel Electrophoresis”. The first step was to make a free account at benchling.com and import the Lambda DNA as seen in [Figure 1] below.
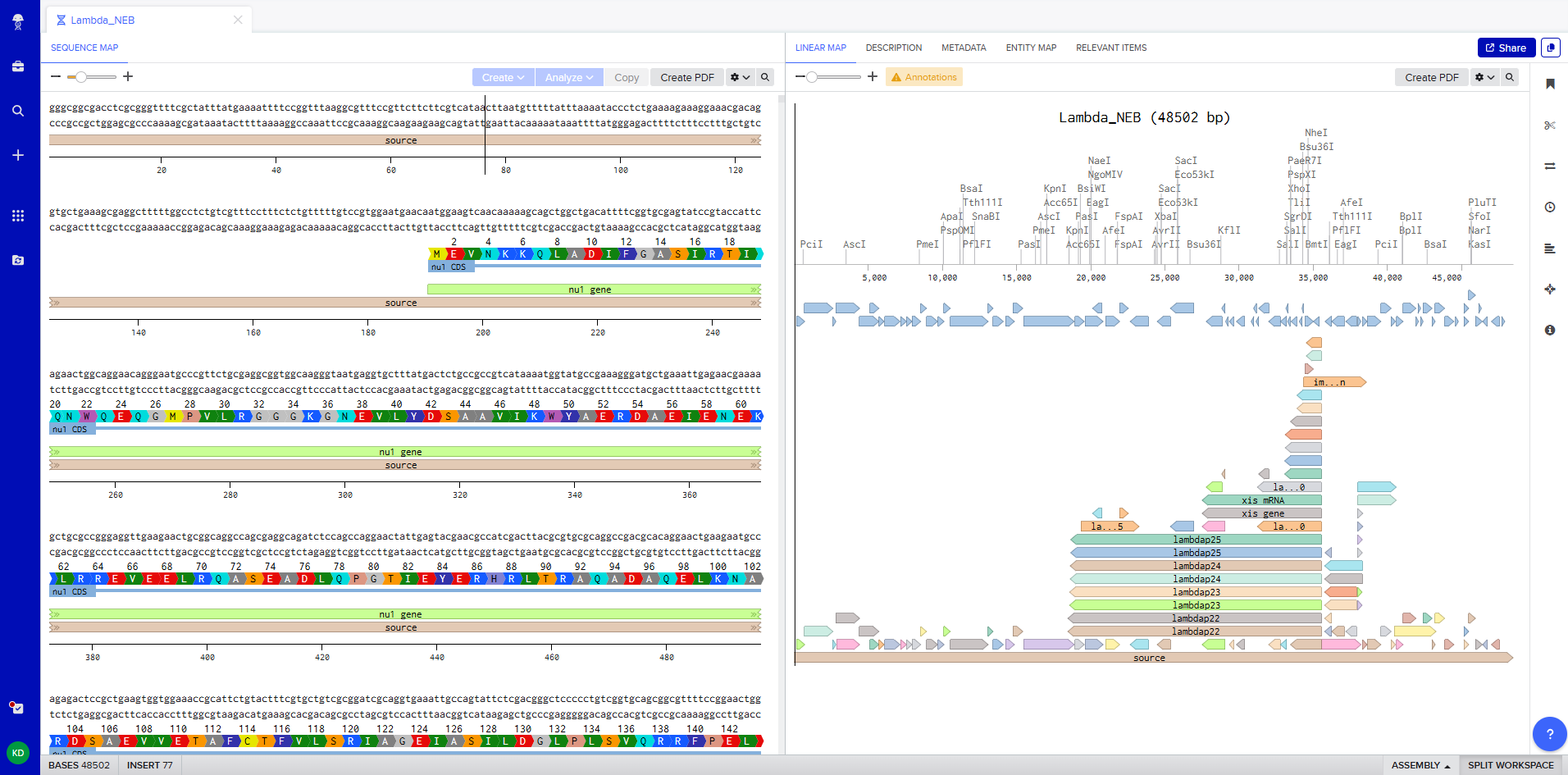
 Figure 1. Imported the Lambda DNA.
Figure 1. Imported the Lambda DNA.
The next step was to simulate restriction enzyme digestion with the following enzymes:
- EcoRI
- HindIII
- BamHI
- KpnI
- EcoRV
- SacI
- SalI
Figure 2 shows the virtual digest with EcoRI and Figure 3 shows the virtual digest with all the enzymes mentioned above.
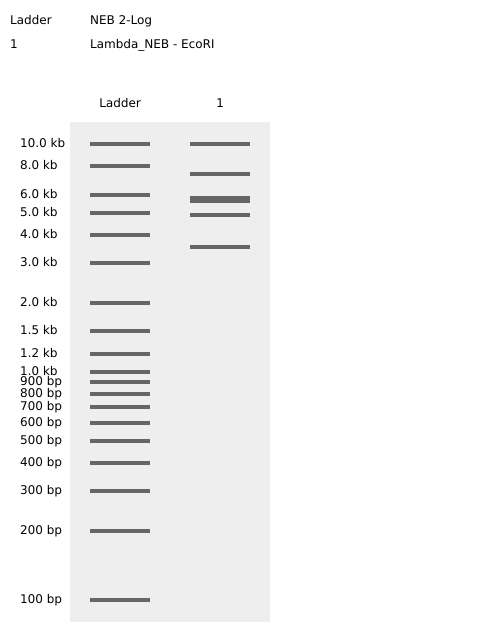
 Figure 2. EcoRI virtual digest.
Figure 2. EcoRI virtual digest.
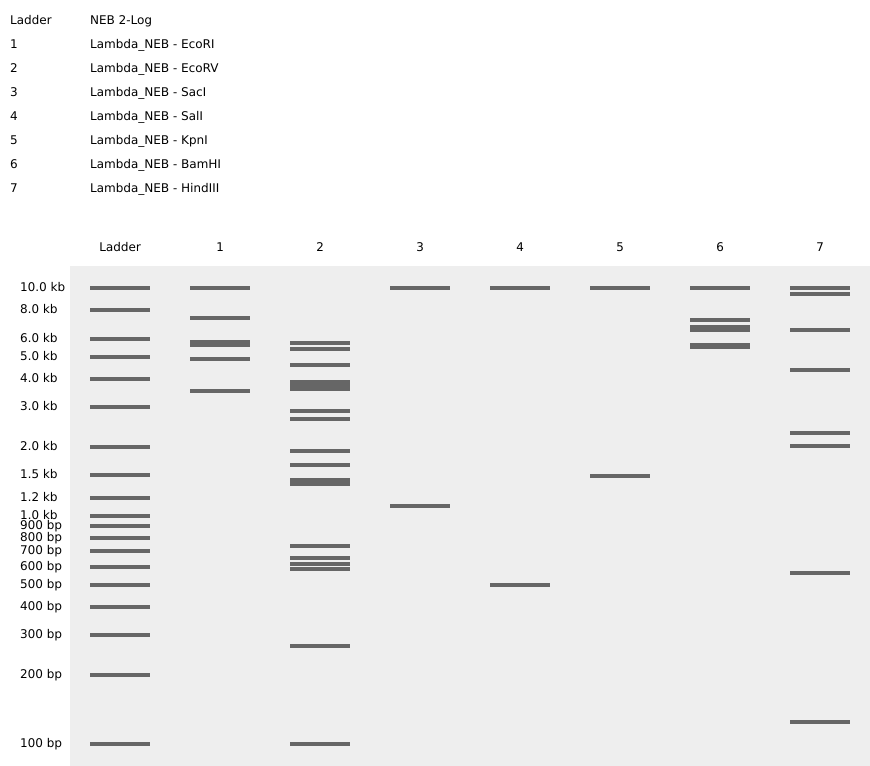

 Figure 3. Full virtual digest.
Figure 3. Full virtual digest.
The last step was to create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks. For this I used Ronan’s website which was a helpful tool to iterate designs, especially since the physical lab experiment could not be carried out due to lab and equipment restrictions.
This was my original idea to make a penguin:
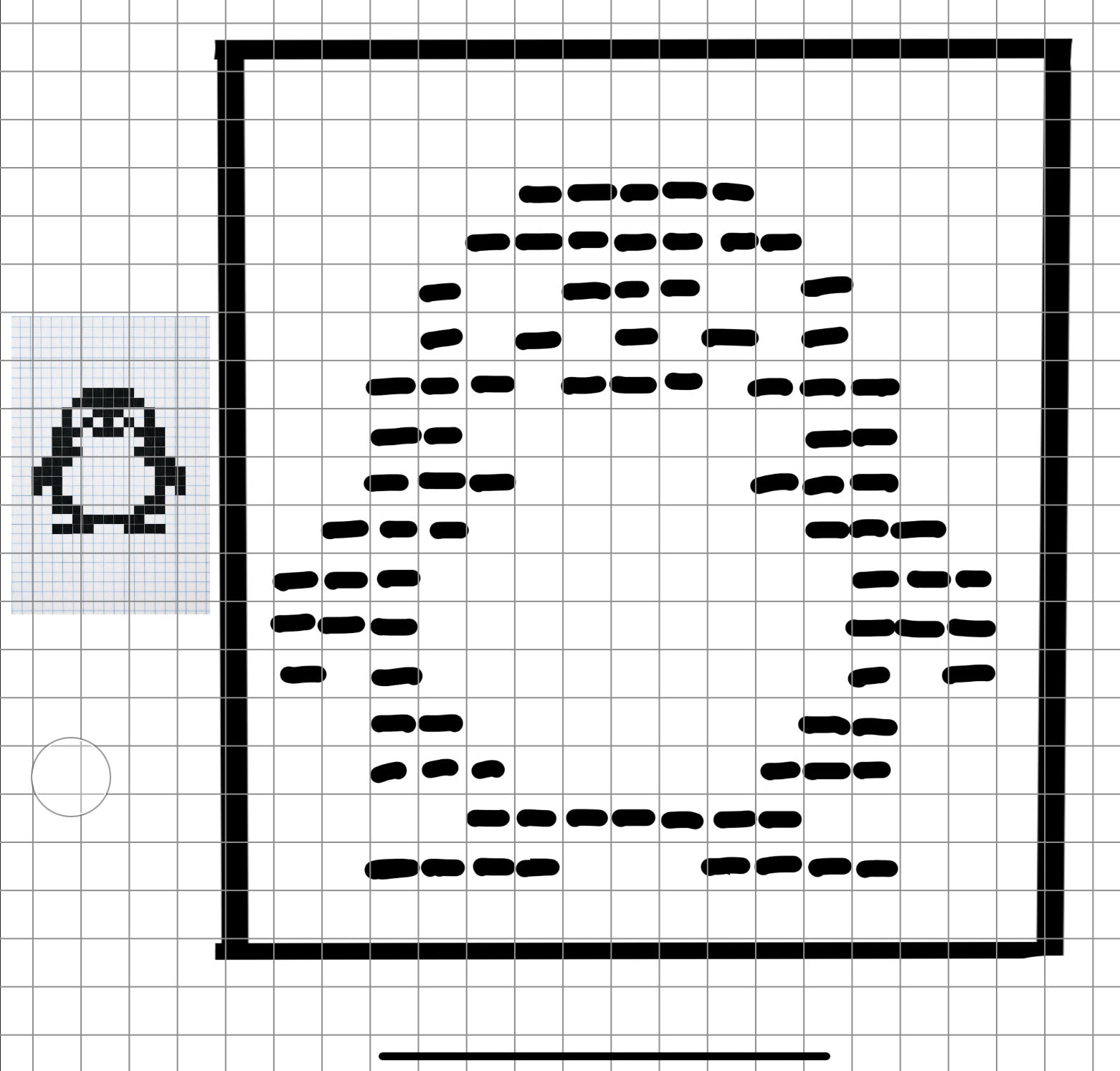
 Figure 4. Pixel penguin sketch.
Figure 4. Pixel penguin sketch.
The result using Ronan’s website:
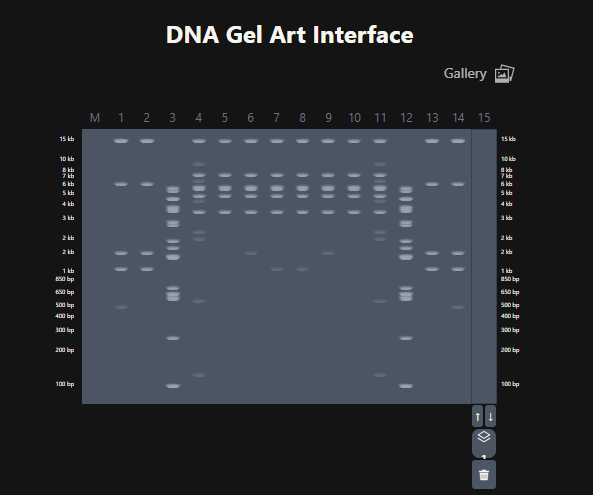

 Figure 5. Attempt at designing a penguin in the style of latent Figure Protocol.
Figure 5. Attempt at designing a penguin in the style of latent Figure Protocol.
These are the gel restriction digest per row:
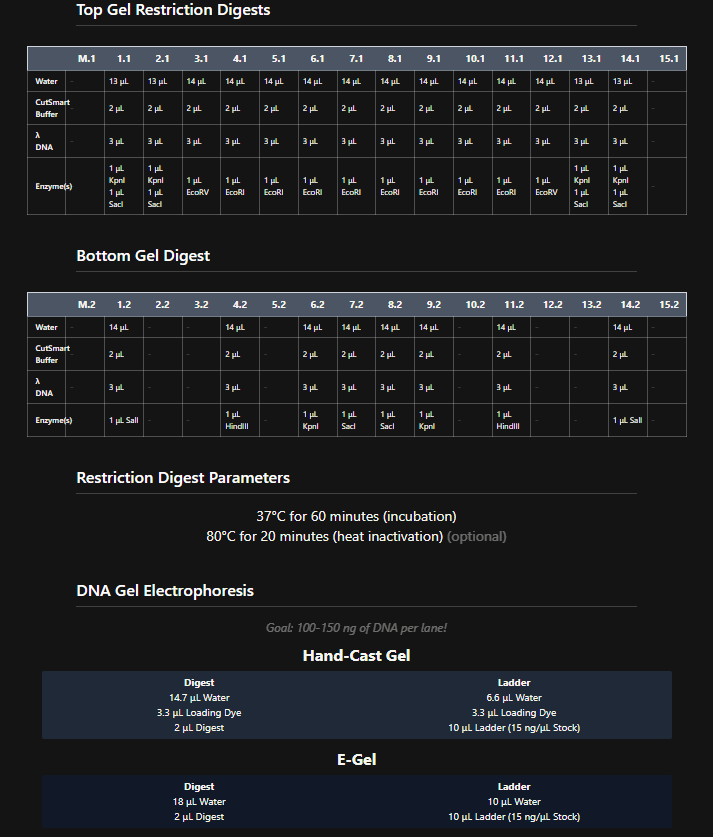
 Figure 6. Tables of restriction enzymes used per row and per layer.
Figure 6. Tables of restriction enzymes used per row and per layer.
Part 4: Prepare a Twist DNA Synthesis Order
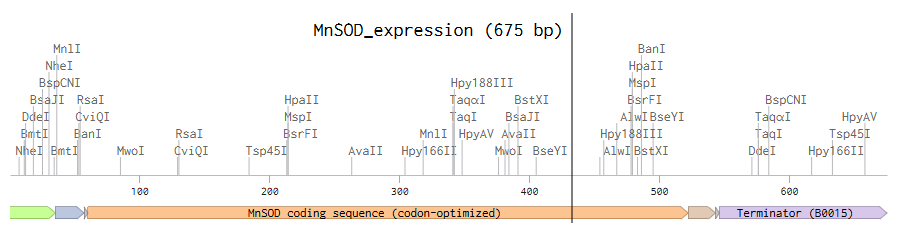
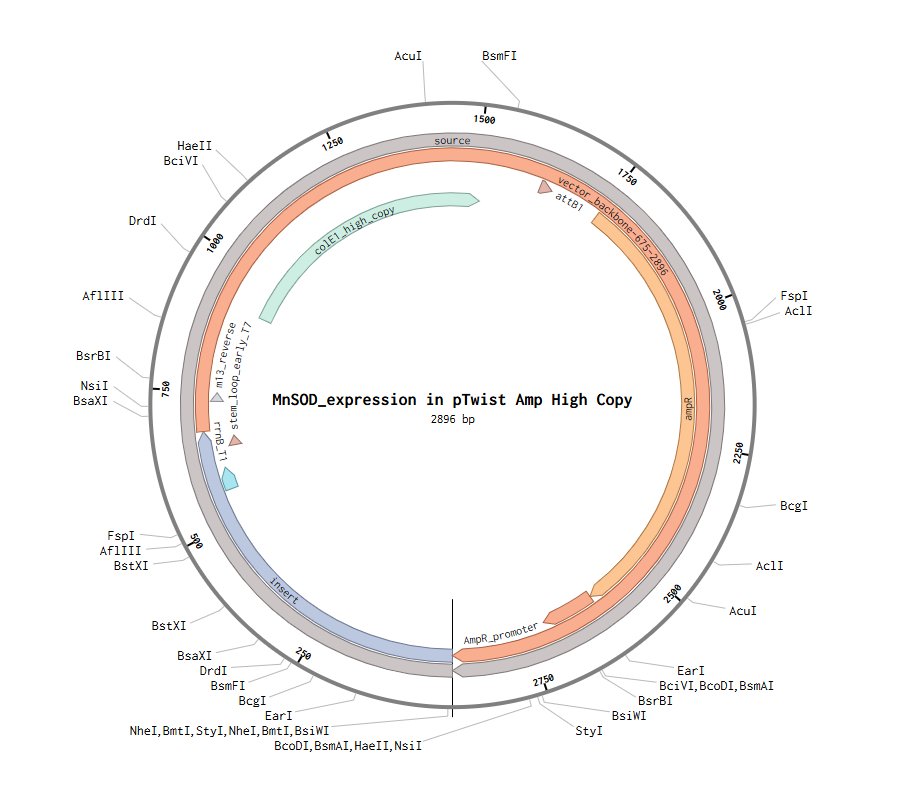
After creating my Twist and Benchling account, I built my DNA insert sequence to make MnSOD (see [Figure 7,8]). I went through each piece of the DNA sequence and annotated the parts and finally got a Linear Map of the entire sequence [Figure 9]. Then on Twist, I imported the sequence by uploading the FASTA file from Benchling. Since the order is for a clonal gene, I had to then select a cloning vector like pTwist Amp High Copy. I then proceeded to download construct (GenBank) to get the full plasmid sequence and imported this to Benchling to see the plasmid with the expression cassette [Figure 10].

 Figure 7. Annotations for MnSOD.
Figure 7. Annotations for MnSOD.