Week 2 HW: DNA Read, Write and Edit


Part 1: Benchling & In-silico Gel Art
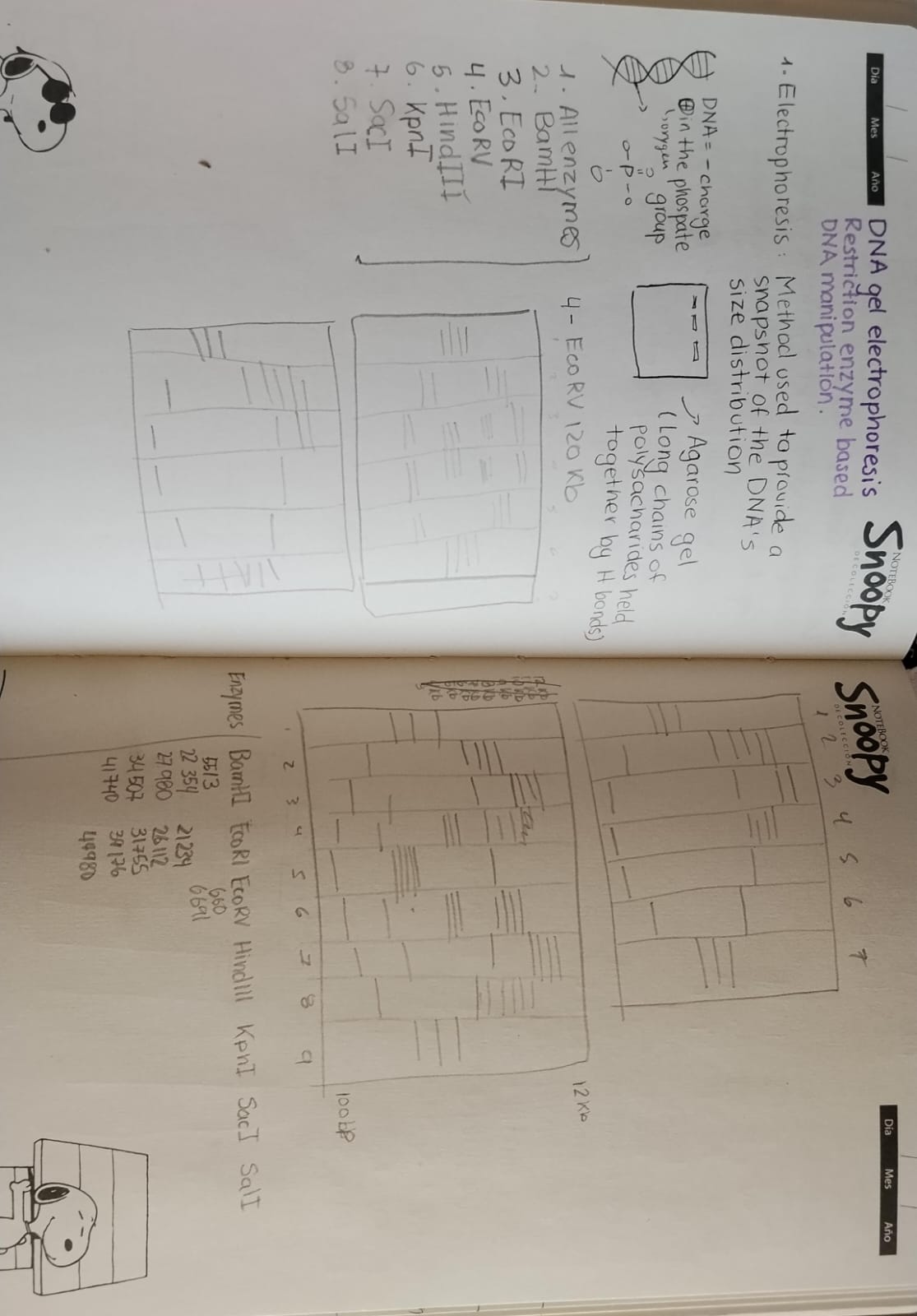
Preliminary notebook sketches illustrating the conceptual design process for the intended latent figure.
Final Product


Part 3: DNA Design Challenge
Protein
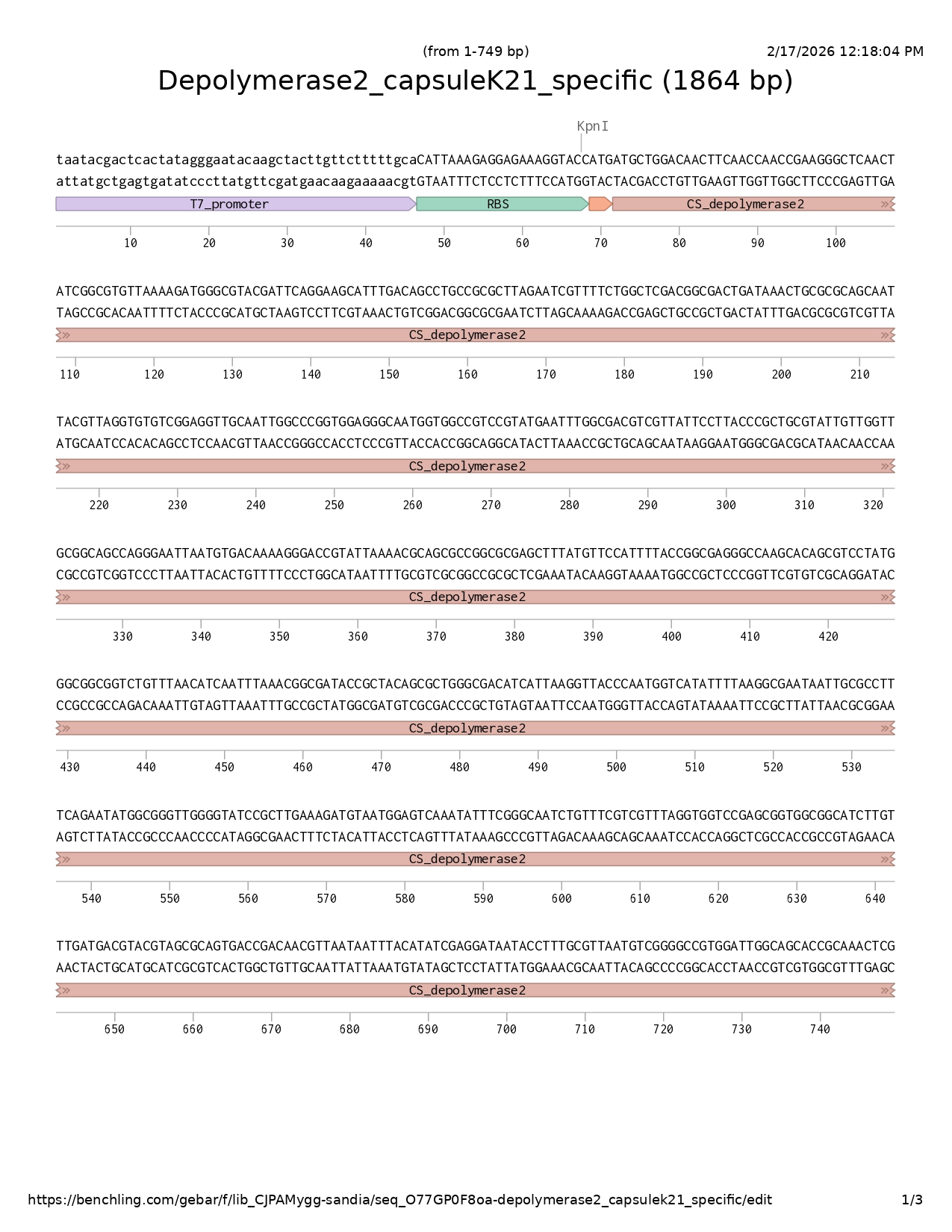
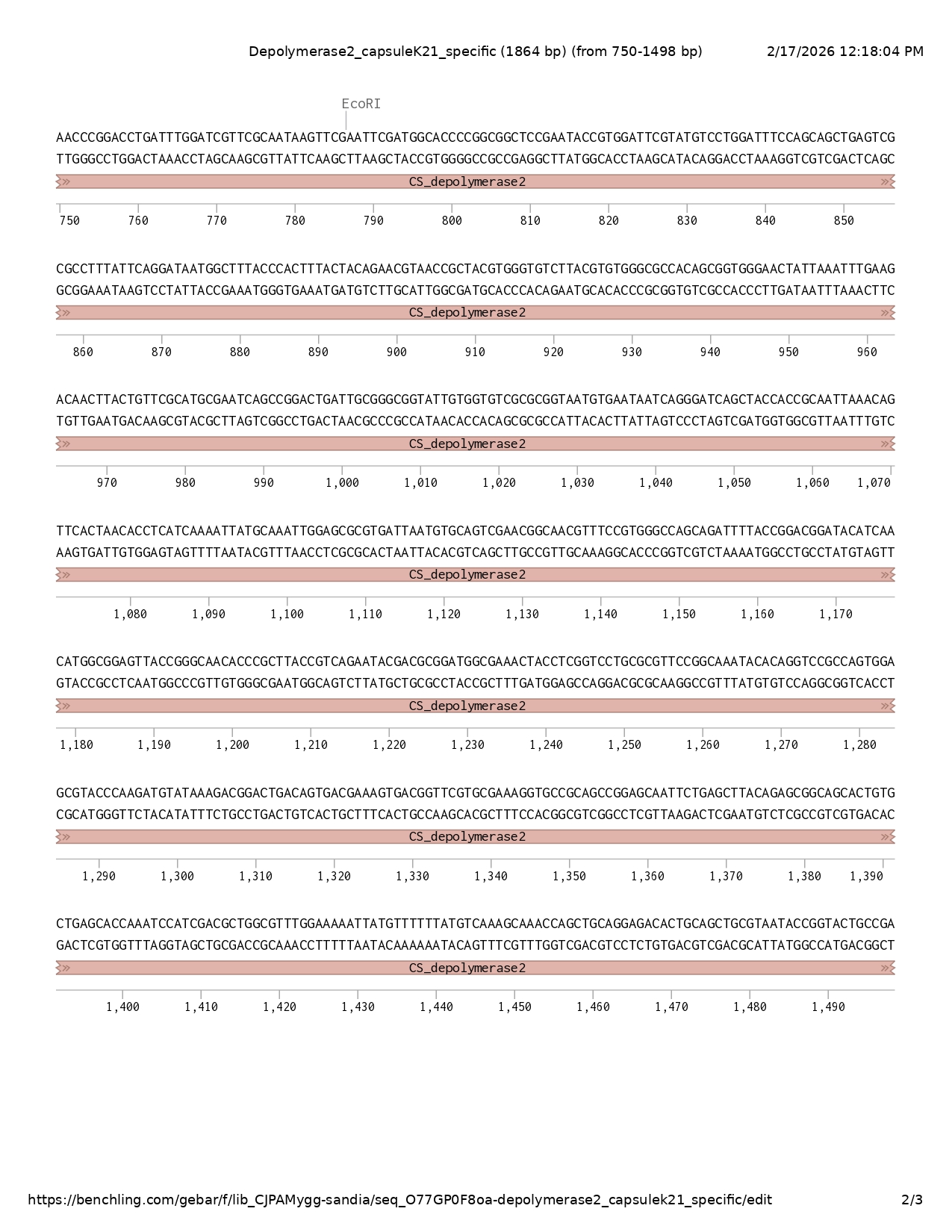
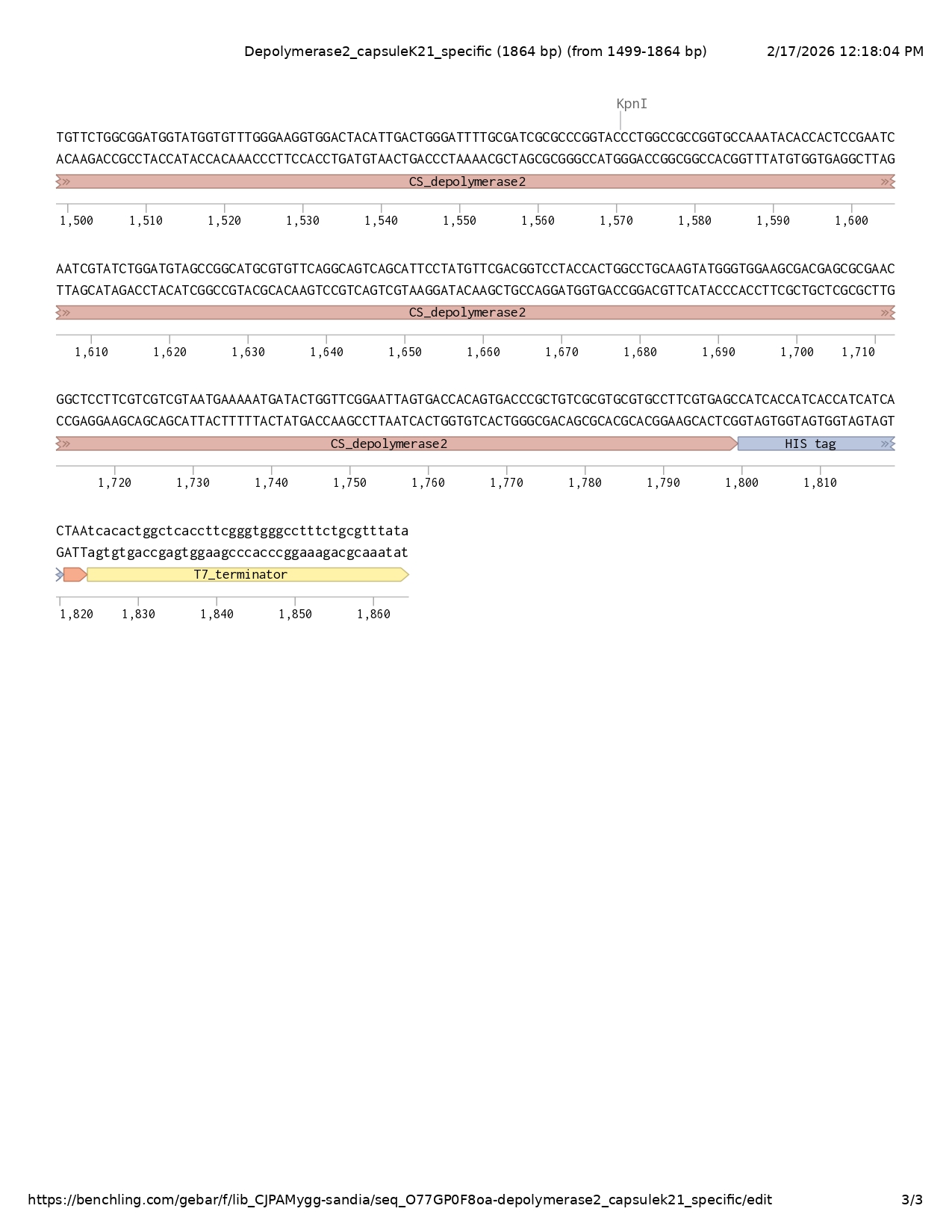
Depolymerase 2 MLDNFNQPKGSTIGVLKDGRTIQEAFDSLPRLESFSGSTATDKLRAAITLGVSEVAIGPVEGNGGRPYEFGDVVIPYPLRIVGCGSQGINVTKGTVLKRSAGASFMFHFTGEGQAQRPMGGGLFNINLNGDTATALGDIIKVTQWSYFKANNCAFQNMAGWGIRLKDVMESNISGNLFRRLGGPSGGGILFDDVRSAVTDNVNNLHIEDNTFALMSGPWIGSTANSNPDLIWIVRNKFEFDGTPAAPNTVDSYVLDFQQLSRAFIQDNGFTHFTTERNRYVGVLRVGATAVGTIKFEDNLLFACESAGLIAGGIVVSRGNVNNQGSATTAIKQFTNTSSKLCKLERVINVQSNGNVSVGQQILPDGYINMAELPGNTRLPSEYDADGETTSVLRVPANTQVRQWSVPKMYKDGLTVTKVTVRAKGAAAGAILSLQSGSTVLSTKSIDAGVWKNYVFYVKANQLQETLQLRNTGTADVLADGMVFGKVDYIDWDFAIAPGTLAAGAKYTTPNQSYLDVAGMRVQAVSIPMFDGPTTGLQVWVEATSANGSFVVVMKNDTGSELVTTVTRCRVRAFVS
Reverse Translate ATGATGTTAGATAATTTTAATCAACCTAAAGGTTCTACTATTGGTGTTTTAAAAGATGGTCGTACTATTCAAGAAGCTTTTGATTCTTTACCTCGTTTAGAATCTTTTTCTGGTTCTACTGCTACTGATAAATTACGTGCTGCTATTACTTTAGGTGTTTCTGAAGTTGCTATTGGTCCTGTTGAAGGTAATGGTGGTCGTCCTTATGAATTTGGTGATGTTGTTATTCCTTATCCTTTACGTATTGTTGGTTGTGGTTCTCAAGGTATTAATGTTACTAAAGGTACTGTTTTAAAACGTTCTGCTGGTGCTTCTTTTATGTTTCATTTTACTGGTGAAGGTCAAGCTCAACGTCCTATGGGTGGTGGTTTATTTAATATTAATTTAAATGGTGATACTGCTACTGCTTTAGGTGATATTATTAAAGTTACTCAATGGTCTTATTTTAAAGCTAATAATTGTGCTTTTCAAAATATGGCTGGTTGGGGTATTCGTTTAAAAGATGTTATGGAATCTAATATTTCTGGTAATTTATTTCGTCGTTTAGGTGGTCCTTCTGGTGGTGGTATTTTATTTGATGATGTTCGTTCTGCTGTTACTGATAATGTTAATAATTTACATATTGAAGATAATACTTTTGCTTTAATGTCTGGTCCTTGGATTGGTTCTACTGCTAATTCTAATCCTGATTTAATTTGGATTGTTCGTAATAAATTTGAATTTGATGGTACTCCTGCTGCTCCTAATACTGTTGATTCTTATGTTTTAGATTTTCAACAATTATCTCGTGCTTTTATTCAAGATAATGGTTTTACTCATTTTACTACTGAACGTAATCGTTATGTTGGTGTTTTACGTGTTGGTGCTACTGCTGTTGGTACTATTAAATTTGAAGATAATTTATTATTTGCTTGTGAATCTGCTGGTTTAATTGCTGGTGGTATTGTTGTTTCTCGTGGTAATGTTAATAATCAAGGTTCTGCTACTACTGCTATTAAACAATTTACTAATACTTCTTCTAAATTATGTAAATTAGAACGTGTTATTAATGTTCAATCTAATGGTAATGTTTCTGTTGGTCAACAAATTTTACCTGATGGTTATATTAATATGGCTGAATTACCTGGTAATACTCGTTTACCTTCTGAATATGATGCTGATGGTGAAACTACTTCTGTTTTACGTGTTCCTGCTAATACTCAAGTTCGTCAATGGTCTGTTCCTAAAATGTATAAAGATGGTTTAACTGTTACTAAAGTTACTGTTCGTGCTAAAGGTGCTGCTGCTGGTGCTATTTTATCTTTACAATCTGGTTCTACTGTTTTATCTACTAAATCTATTGATGCTGGTGTTTGGAAAAATTATGTTTTTTATGTTAAAGCTAATCAATTACAAGAAACTTTACAATTACGTAATACTGGTACTGCTGATGTTTTAGCTGATGGTATGGTTTTTGGTAAAGTTGATTATATTGATTGGGATTTTGCTATTGCTCCTGGTACTTTAGCTGCTGGTGCTAAATATACTACTCCTAATCAATCTTATTTAGATGTTGCTGGTATGCGTGTTCAAGCTGTTTCTATTCCTATGTTTGATGGTCCTACTACTGGTTTACAAGTTTGGGTTGAAGCTACTTCTGCTAATGGTTCTTTTGTTGTTGTTATGAAAAATGATACTGGTTCTGAATTAGTTACTACTGTTACTCGTTGTCGTGTTCGTGCTTTTGTTTCTTAA
Codon optimization ATG TTG GAT AAT TTC AAC CAG CCA AAA GGC TCG ACG ATC GGG GTG CTG AAG GAC GGC CGT ACA ATT CAG GAA GCG TTT GAC AGC CTG CCG CGC CTT GAA TCT TTT TCG GGC AGT ACG GCA ACT GAT AAA CTG CGT GCG GCG ATC ACT CTT GGC GTT AGT GAA GTT GCG ATC GGT CCA GTG GAA GGT AAT GGC GGC CGT CCG TAT GAA TTT GGG GAT GTT GTG ATT CCC TAT CCA TTG CGC ATT GTG GGC TGC GGC AGC CAA GGG ATC AAT GTA ACT AAA GGT ACG GTC TTA AAA CGT AGT GCC GGA GCG TCC TTT ATG TTC CAT TTT ACT GGG GAA GGT CAG GCC CAG CGC CCG ATG GGA GGC GGT CTG TTT AAT ATT AAC CTG AAC GGC GAT ACC GCG ACC GCA CTG GGC GAT ATC ATT AAA GTA ACT CAG TGG AGT TAT TTT AAA GCG AAC AAT TGC GCT TTT CAA AAT ATG GCG GGG TGG GGC ATC CGT CTG AAG GAC GTG ATG GAA AGC AAT ATC AGC GGA AAC TTG TTC CGT CGC CTG GGA GGC CCG TCT GGG GGT GGC ATC TTG TTC GAT GAC GTC CGT AGC GCG GTA ACA GAC AAT GTA AAC AAT TTA CAC ATT GAA GAT AAC ACT TTT GCG TTA ATG AGC GGC CCC TGG ATT GGT AGC ACC GCG AAT AGT AAC CCG GAT CTG ATC TGG ATC GTG CGT AAT AAA TTC GAA TTT GAT GGC ACT CCA GCT GCA CCG AAC ACT GTT GAT AGC TAC GTC CTG GAT TTT CAA CAG CTT AGC CGC GCA TTT ATC CAG GAC AAT GGG TTC ACG CAC TTT ACC ACG GAA CGT AAC CGT TAC GTT GGT GTG TTA CGT GTA GGC GCA ACG GCC GTT GGC ACC ATT AAA TTC GAA GAT AAC CTG CTG TTC GCC TGC GAA AGC GCC GGC CTG ATC GCG GGC GGC ATC GTT GTT AGT CGC GGT AAC GTG AAC AAC CAG GGC TCC GCT ACG ACG GCC ATT AAA CAG TTC ACG AAT ACG TCC AGC AAA TTG TGT AAA CTG GAA CGT GTT ATT AAC GTG CAG AGT AAT GGC AAT GTG TCG GTG GGC CAA CAA ATC CTG CCG GAC GGG TAT ATC AAT ATG GCT GAG CTG CCT GGC AAC ACC CGC TTA CCG AGC GAA TAT GAC GCA GAT GGT GAA ACT ACC AGT GTA TTA CGC GTG CCA GCA AAC ACC CAG GTC CGC CAG TGG TCG GTG CCT AAA ATG TAT AAA GAC GGC TTG ACC GTA ACG AAA GTG ACG GTC CGT GCA AAA GGG GCA GCC GCC GGT GCC ATC CTG AGC TTG CAG AGC GGC TCG ACC GTG CTG TCT ACG AAA AGC ATT GAT GCT GGC GTG TGG AAG AAT TAT GTT TTC TAT GTT AAA GCG AAT CAG CTT CAG GAA ACT CTG CAG CTT CGC AAT ACA GGT ACT GCA GAC GTA CTT GCG GAC GGT ATG GTT TTT GGC AAG GTG GAT TAT ATC GAC TGG GAT TTC GCG ATT GCC CCG GGG ACC CTG GCG GCC GGT GCG AAA TAT ACG ACC CCT AAT CAG TCG TAC CTG GAT GTC GCG GGC ATG CGT GTG CAA GCG GTC TCG ATT CCC ATG TTT GAT GGC CCT ACG ACT GGA TTA CAG GTC TGG GTA GAA GCC ACC AGC GCG AAC GGT AGT TTC GTG GTG GTC ATG AAA AAC GAC ACG GGT TCA GAA TTG GTC ACC ACC GTG ACT CGC TGC CGT GTG CGC GCG TTT GTA TCA
Build Your DNA Insert Sequence
Choose Your Vector

What DNA would you want to sequence (e.g., read) and why?
I would like to sequence viral metagenomic DNA (environmental virome) collected from aquatic ecosystems, such as wastewater effluent, hospital discharge sites, and agricultural runoff. Rather than focusing exclusively on bacterial genomes, this approach prioritizes complete bacteriophage genomes present in these environments.
Phages are major drivers of bacterial evolution, influencing antimicrobial resistance dissemination, virulence modulation, and horizontal gene transfer. By sequencing environmental phage DNA, it becomes possible to identify functional genetic modules such as receptor-binding proteins, depolymerases, integrases, and transducing elements that shape bacterial populations.
This intends shifts surveillance from reactive pathogen detection to a predictive aproach. Environmental virome sequencing could serve as an early-warning system, detecting emerging resistance dynamics or novel virulence-associated genetic elements before they become clinically dominant.
In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
To sequence environmental viral metagenomic DNA, It would be ideal to use a combination of third-generation long-read sequencing (Oxford Nanopore or PacBio) and second-generation high-throughput short-read sequencing (Illumina) in a hybrid strategy.
I think that this approach would leverage the strengths of both platforms: long reads enabling assembly of complete phage genomes and the resolution of structural variants, while short reads provide high accuracy for polishing and variant correction.
Input: purified viral DNA extracted from environmental water samples.
Essential preparation steps:
viral particle enrichment (filtration and DNase treatment to remove non-viral DNA)
viral DNA extraction
library preparation
fragmentation (for short-read platforms)
end repair and adapter ligation
quality control and quantification
Illumina (second generation):
DNA fragments bind to a flow cell.
Bridge amplification creates clusters.
Sequencing by synthesis occurs using fluorescently labeled reversible terminator nucleotides.
After each nucleotide incorporation, fluorescence is detected.
Base calling is determined by the emitted fluorescent signal.
Nanopore (third generation):
Single DNA molecules pass through a protein nanopore.
Each nucleotide alters ionic current differently.
Electrical signal changes are recorded in real time.
Machine learning algorithms convert signal patterns into base calls.
The output consists of:
FASTQ files containing sequence reads with quality scores
Assembled phage genomes
Annotated functional gene predictions
Comparative genomic datasets for surveillance
What DNA would you want to edit and why?
For this project, I would love to synthesize a phage-derived receptor-binding domain (RBD) fused to a fluorescent reporter, essentially creating a highly specific bacterial detection module inspired by bacteriophages.
Phages are incredibly precise when it comes to recognizing their bacterial hosts — their tail fibers or tailspikes bind very specific surface structures like capsules or LPS. Instead of synthesizing a whole phage genome (which would be unnecessary and unsafe), I would isolate just the receptor-binding domain of a phage tail fiber that targets a clinically relevant bacterium, such as Klebsiella pneumoniae. Then I would fuse that domain to a reporter protein like GFP.
The idea is that this synthetic gene would encode a fusion protein that binds specifically to its bacterial target and produces a fluorescent signal. So instead of using antibodies for detection, we would be using phage specificity as a biosensing tool. I think that’s incredibly powerful because phage receptor-binding proteins are often more specific than antibodies and can distinguish even subtle differences like capsule types.
To synthesize the phage receptor-binding domain–GFP fusion construct, I would use commercial gene synthesis based on phosphoramidite solid-phase DNA synthesis, followed by enzymatic DNA assembly (such as Gibson Assembly).
Phosphoramidite chemistry is the standard method used to chemically synthesize short DNA oligonucleotides. These oligos can then be assembled enzymatically into a full-length gene construct. This approach is highly accurate and allows complete sequence customization, including codon optimization and addition of regulatory elements.
I would choose this method because it enables precise design of non-replicative, modular constructs without needing a natural template, which is ideal for synthetic biology applications.
What DNA would you want to edit and why?
I would want to edit bacteriophage genomes, specifically lytic phages that infect clinically relevant bacteria such as Klebsiella pneumoniae or other multidrug-resistant pathogens.
Phages naturally evolve to recognize and infect bacteria, but their host range is often narrow and their therapeutic use can be limited by bacterial resistance mechanisms. By editing phage DNA, we could enhance desirable properties such as host specificity, lytic efficiency, or anti-virulence activity, while maintaining safety.
The types of edits I would focus on include:
Modifying tail fiber or receptor-binding protein genes to expand or retarget host range.
Inserting capsule depolymerase genes to improve penetration of protective bacterial capsules.
Deleting lysogeny-related genes (if present) to ensure strictly lytic behavior.
Optimizing regulatory elements to increase stability and predictability of infection dynamics.
The goal would not be to make phages more harmful, but rather more precise and controllable as therapeutic or ecological tools. In a One Health context, engineered phages could be used to reduce pathogenic bacteria in clinical, agricultural, or environmental settings without relying solely on antibiotics.
To edit bacteriophage genomes, I would use CRISPR-Cas–based genome editing combined with homologous recombination in bacterial host cells.
CRISPR-Cas systems are precise and programmable, making them ideal for modifying specific genes such as tail fiber or depolymerase genes. This approach allows targeted edits without randomly mutating the phage genome. CRISPR-Cas works by using a guide RNA to direct the Cas nuclease to a specific DNA sequence. The Cas enzyme creates a cut at that location. If a repair template containing the desired modification is provided, the cell’s natural DNA repair machinery incorporates the new sequence.
For phage editing, the general process would involve:
Designing a guide RNA targeting the phage gene of interest.
Designing a donor DNA template containing the desired modification.
Introducing the CRISPR system and donor template into a bacterial host.
Infecting the bacteria with the phage.
Selecting for phages that incorporate the desired edit.
Preparation includes:
Designing guide RNAs targeting specific phage genes.
Designing a donor DNA repair template containing the edited sequence.
Cloning CRISPR components into plasmids.
Preparing competent bacterial host cells.
There are several limitations:
Editing efficiency can vary depending on the phage and target gene.
Some phages may escape editing due to rapid replication.
Off-target effects are possible if guide RNAs are not carefully designed.
Delivery of editing components must be optimized.