Week 6 — Genetic Circuits Part I: Assembly Technologies
- What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
- Phusion High-Fidelity DNA Polymerase A proofreading polymerase with 3′→5′ exonuclease activity, which ensures very low error rates during DNA synthesis.
- Phusion HF or GC Buffer Provides optimal ionic conditions (Mg²⁺, salts, pH).
- HF buffer: for standard templates
- GC buffer: improves amplification of GC-rich or difficult templates
- dNTPs (400 µM each) Building blocks (dATP, dTTP, dCTP, dGTP) required for DNA strand synthesis.
- Mg²⁺ (within the buffer) Essential cofactor for polymerase activity and influences enzyme fidelity and efficiency.
- What are some factors that determine primer annealing temperature during PCR? The annealing temperature in PCR is determined by several factors:
- Primer melting temperature (Tm) Calculated based on primer sequence (GC content, length). Annealing temperature is typically ~3–5°C below Tm
- Primer length Longer primers = higher Tm
- GC content Higher GC = stronger binding = higher annealing temperature
- Primer sequence composition Secondary structures (hairpins, dimers) affect binding
- Salt concentration Higher salt stabilizes primer-template binding
- Polymerase type Some enzymes (like Phusion) require higher annealing temperatures due to their buffer system
- There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
| PCR | Restriction Enzyme Digests | |
|---|---|---|
| Starting materials | Template DNA and primers | DNA with restriction sites |
| Key reagents | Polymerase, primers and dNTPs | Restriction enzyme and buffer |
| Mechanism | DNA amplification | DNA cutting |
| Temperature profile | Multiple cycles | Single temperature |
| Control of fragment | Defined by primer | Defined by enzyme sites |
| Output | Many copies of a single fragment | Multiple fragments |
| Critical design step | Primer design | Enzyme selection |
| Time | 1 to 3 hours | Roughly 1 hour |
| Flexibility | High | Limited by sequence |
PCR is generally preferable when you need to generate a specific DNA fragment with precise boundaries or added sequences, such as overlaps for Gibson Assembly, because it allows high flexibility through primer design and can amplify even very small amounts of DNA. In contrast, restriction enzyme digestion is preferable when the DNA already contains suitable restriction sites, making it a simpler and faster method for cutting plasmids or generating fragments without the need for amplification. Therefore, PCR is favored for custom design and low DNA availability, while restriction digestion is best for routine cloning tasks where appropriate sites are already present.
- How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning? To ensure that DNA fragments are appropriate for Gibson Assembly, several critical criteria must be met:
- Presence of overlapping regions (20–40 bp) Fragments must share homologous sequences at their ends to allow correct assembly.
- Correct sequence design Overlaps must be: Specific, In the correct orientation, Free of mismatches
- Proper fragment size and integrity Verified by gel electrophoresis
- High purity of DNA Removal of primers, enzymes, and contaminants (e.g., via PCR cleanup)
- Correct concentration (stoichiometry) Balanced molar ratios improve assembly efficiency (this is where your answer fits)
- Absence of unwanted sequences No internal overlaps or conflicting regions
- How does the plasmid DNA enter the E. coli cells during transformation? Plasmid DNA enters E. coli cells through artificially induced membrane permeability:
- Heat shock transformation Cells are treated with CaCl₂ to make membranes more permeable A sudden temperature increase (e.g., 42°C) creates a thermal imbalance This allows DNA to enter the cell
- Electroporation A short electrical pulse creates temporary pores in the membrane DNA enters through these pores After entry the membrane reseals, the cell recovers and begins expressing the plasmid
- Describe another assembly method in detail
SLIC is a DNA assembly method that joins fragments based on short homologous overlaps, similar to Gibson Assembly but with less enzyme requirements. In this method, DNA fragments are first designed with overlapping regions (15–25 bp). A 3′→5′ exonuclease activity (often from T4 DNA Polymerase) is used to chew back the ends of the DNA, generating complementary single-stranded overhangs. These overhangs allow fragments to anneal to each other without ligase initially, forming a stable intermediate. The assembled DNA is then transformed into bacteria, where the host’s repair machinery seals the nicks in the backbone. Unlike Gibson Assembly, SLIC does not require a ligase or multiple enzymes in the reaction mix, making it more cost-effective. However, it can be slightly less efficient and more sensitive to experimental conditions. Therefore, SLIC is a useful low-cost, overlap-based assembly method.
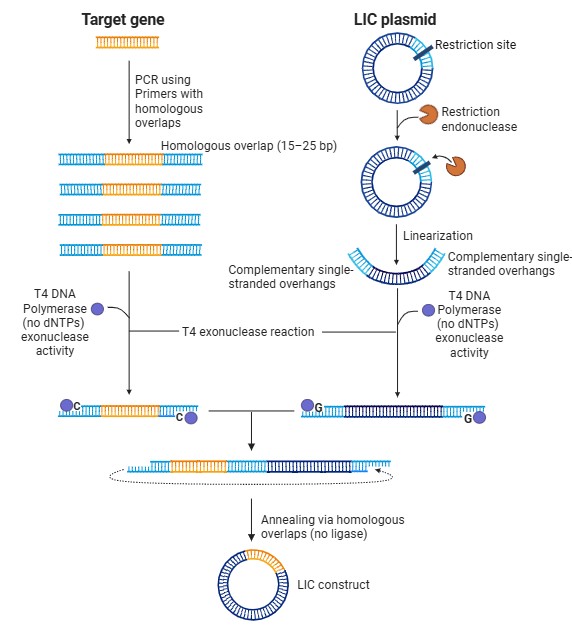

Model this assembly method with Benchling or Asimov Kernel!
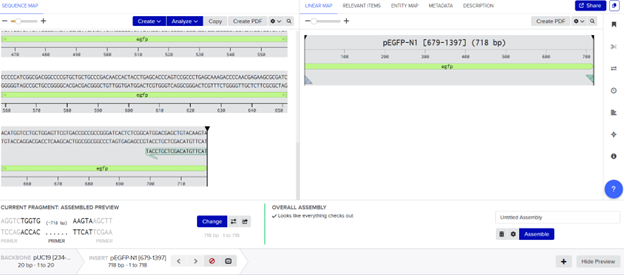
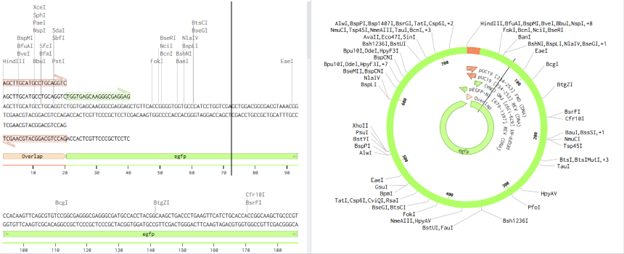
SLIC was modeled in Benchling by designing 20 bp homologous overlaps via PCR primers and assembling fragments using an overlap-based assembly tool (Gibson simulation), since exonuclease processing and in vivo repair cannot be directly simulated.
ASIMOV kernel
Build three of your own Constructs using the parts in the Characterized Bacterials Parts Repo
- Explain in the Notebook Entry how you think each of the Constructs should function
- Run the simulator and share your results in the Notebook Entry
- If the results don’t match your expectations, speculate on why and see if you can adjust the simulator settings to get the expected outcome
Construct 1
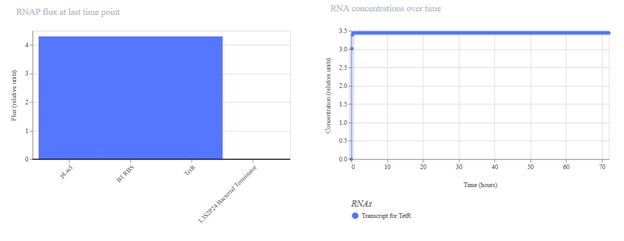
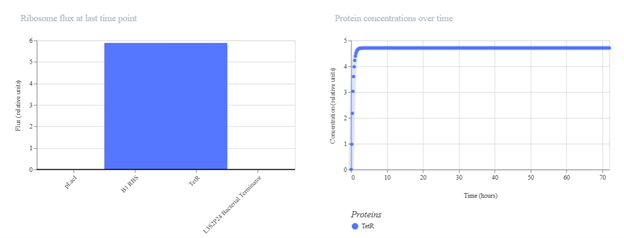
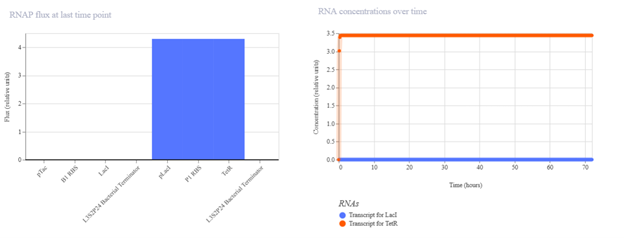
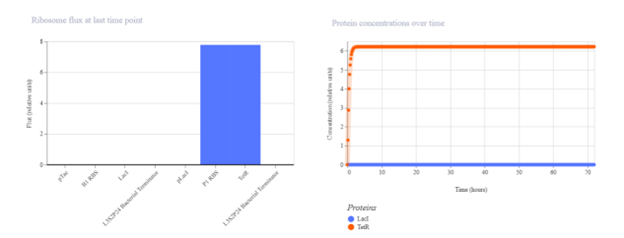
pTac → B1 RBS → TetR → Terminator I expected pTac to constitutively drive TetR production. In simulation, TetR should increase over time and possibly stabilize at a plateau. If expression is lower than expected, it may be due to weak promoter activity, translation efficiency, or high degradation settings.
Construct 2
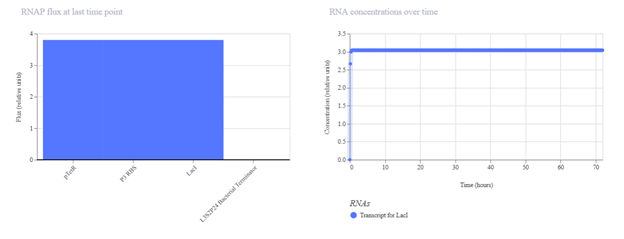
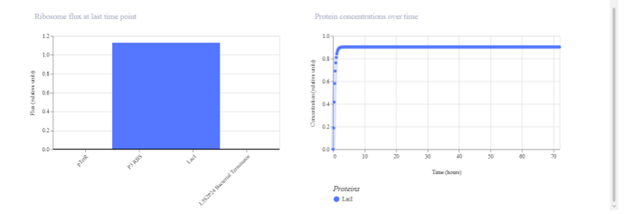
pTetR → P3 RBS → LacI → Terminator I expected this construct to produce LacI continuously. The simulation output should show rising LacI concentration, although with dynamics that may differ from Construct 1 due to promoter and RBS differences. If the observed signal is weak, it may reflect lower promoter strength or insufficient simulation time.
Construct 3
pTac → B1 RBS → LacI → Terminator + pLacI → P1 RBS → TetR → Terminator I expected the first cassette to express LacI, which would repress the pLacI promoter in the second cassette and keep TetR expression low. In simulation, LacI should increase while TetR should remain low compared with an unrepressed expression construct. If TetR remains high, possible causes include promoter leakiness, delayed LacI accumulation, or insufficient repression strength.