Week 5 Protein Design Part 2
Part 1: Generate Binders with PepMLM
Begin by retrieving the human SOD1 sequence from UniProt (P00441) and introducing the A4V mutation. I visited the Uniprot page for (P00441)[https://www.uniprot.org/uniprotkb/P00441/entry#sequences] and the normal sequence is:
012345… MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
A4V represents one mutation where the ‘A’ changes to a ‘V’ at position 4: MATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
I found additional information about the AV4 mutation on the ALS Association’s page
Using the PepMLM Colab linked from the HuggingFace PepMLM-650M model card: Generate four peptides of length 12 amino acids conditioned on the mutant SOD1 sequence. To your generated list, add the known SOD1-binding peptide FLYRWLPSRRGG for comparison. Record the perplexity scores that indicate PepMLM’s confidence in the binders.
I opened my copy of the PepMLM script and entered the sequence. I tried adjusting the slider to change the peptide length from 15 to 12, but the slider appears fixed. So I opened the code and just changed the peptide length to ‘12’. I set the number of peptides to 4. ![Adjusting the PepMLM script]](sod1_a4v_pepMLM1.png)
I then ran the code for ‘Load Model’. Next, I modified ‘Generate Peptides’ so that it would include the known binder FLYRWLPSRRGG I obtained one binder result:
Binder Pseudo Perplexity
FLYRWLPSRRGG 20.63523127
WHYYAAALEHGX 11.53622912
WLYYVTAVRWGX 22.31654743
HRYPAAAVAHKX 8.720199577
WRYPVAAAEWGE 14.43921585'
Part 2: Evaluate Binders with AlphaFold3
Navigate to the AlphaFold Server: alphafoldserver.com. For each peptide, submit the mutant SOD1 sequence followed by the peptide sequence as separate chains to model the protein-peptide complex. Record the ipTM score and briefly describe where the peptide appears to bind.
For each peptide sequence, I used the alphafold query form as shown:
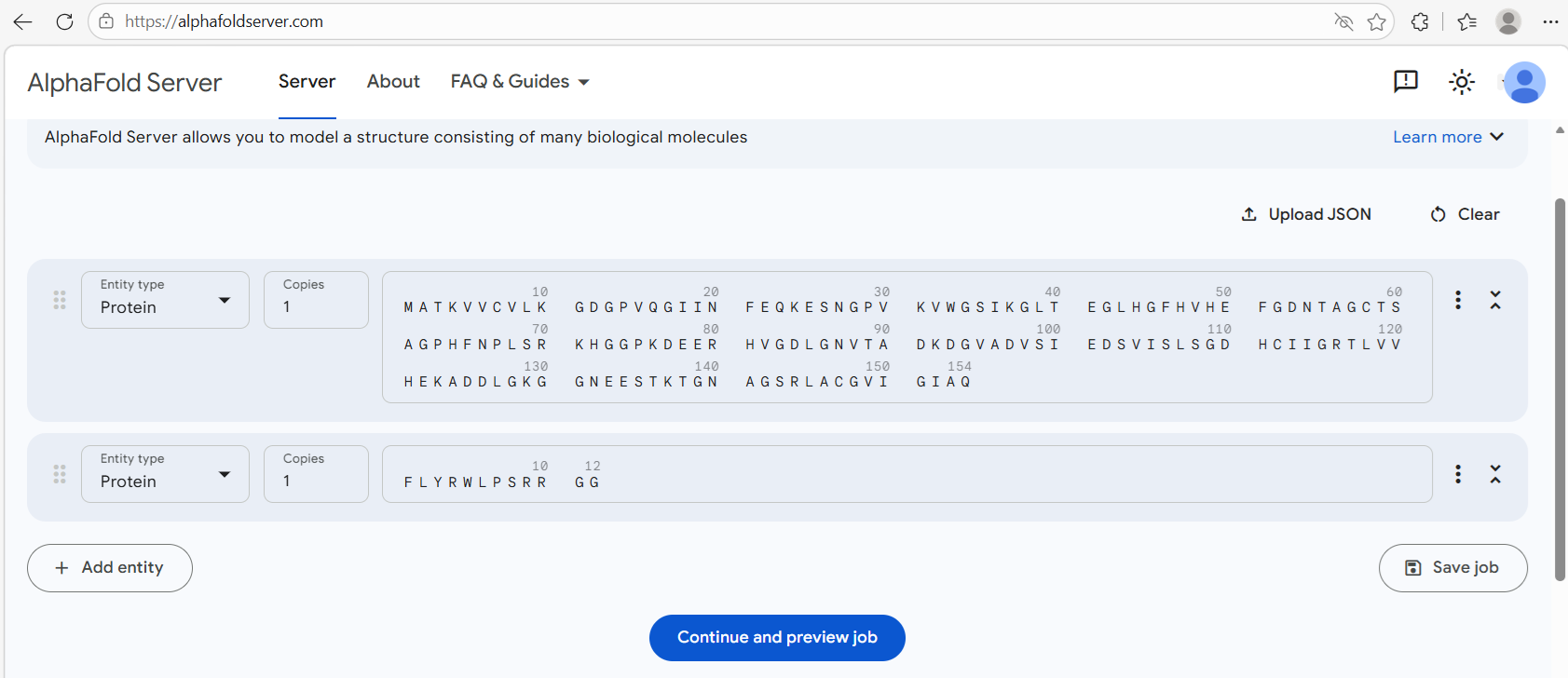

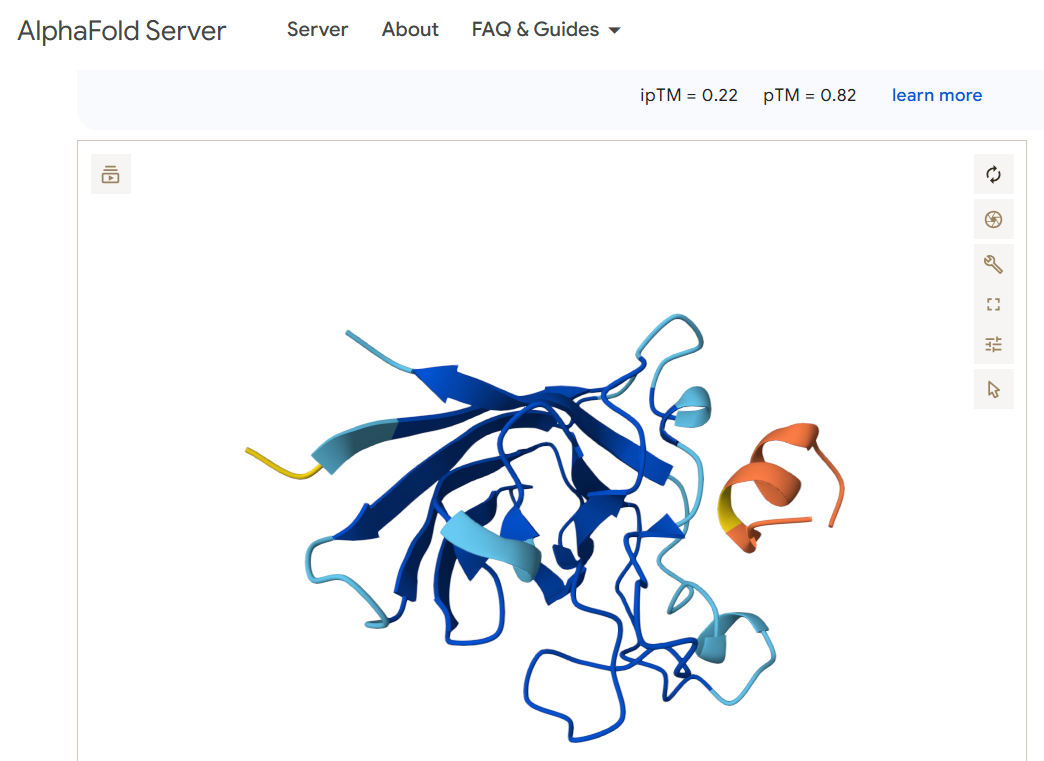
For the first candidate binder FLYRWLPSRRGG, I obtained this image:

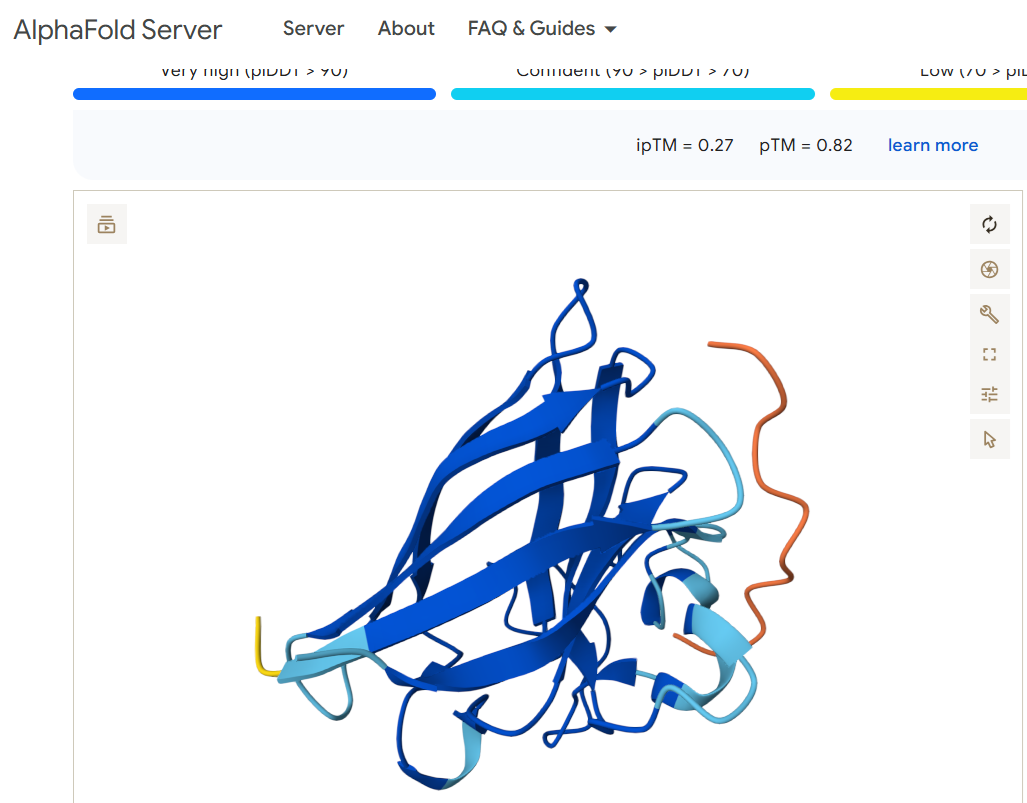
For the binder WRYPVAAAEWGE, I obtained this image:

For the other three binder candidates, I got an error:


These are the results I could gather:
Binder ipTM pTM
FLYRWLPSRRGG 0.22 0.82
WHYYAAALEHGX Error
WLYYVTAVRWGX Error
HRYPAAAVAHKX Error
WRYPVAAAEWGE 0.27 0.82'
Does it localize near the N-terminus where A4V sits? Neither of the two viable binding candidates FLYRWLPSRRGG or WRYPVAAAEWGE appears to bind anywhere near the N terminus (shown in the left part of the diagrams)
Does it engage the β-barrel region or approach the dimer interface? The peptides appear to engage the beginning of the beta sheet.
Does it appear surface-bound or partially buried? Both of the peptides appear to hover over the other protein and seem to me like they may be more surface bound than becoming buried or entangled.
In a short paragraph, describe the ipTM values you observe and whether any PepMLM-generated peptide matches or exceeds the known binder. The two binder candidates that are able to work have a similar pTM score. Both are above 0.5 which suggests the overall predicted fold for the complex might be similar to the true structure. Both also have a very low ipTM, which suggests they are low-confidence predictions of relative positions of subunits within the complex. The candidate WRYPVAAAEWGE seems slightly better a candidate than FLYRWLPSRRGG, but not by much. Both seem like poor candidates as they don’t appear to bind anywhere near the N terminus where the mutation exists.
Part 3: Evaluate Properties of Generated Peptides in the PeptiVerse
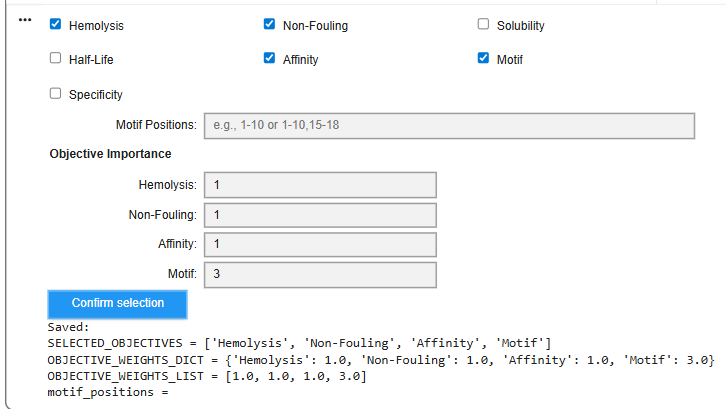
Structural confidence alone is insufficient for therapeutic development. Using PeptiVerse, let’s evaluate the therapeutic properties of your peptide! For each PepMLM-generated peptide: Paste the peptide sequence. Paste the A4V mutant SOD1 sequence in the target field. Check the boxes Predicted binding affinity Solubility Hemolysis probability Net charge (pH 7) Molecular weight
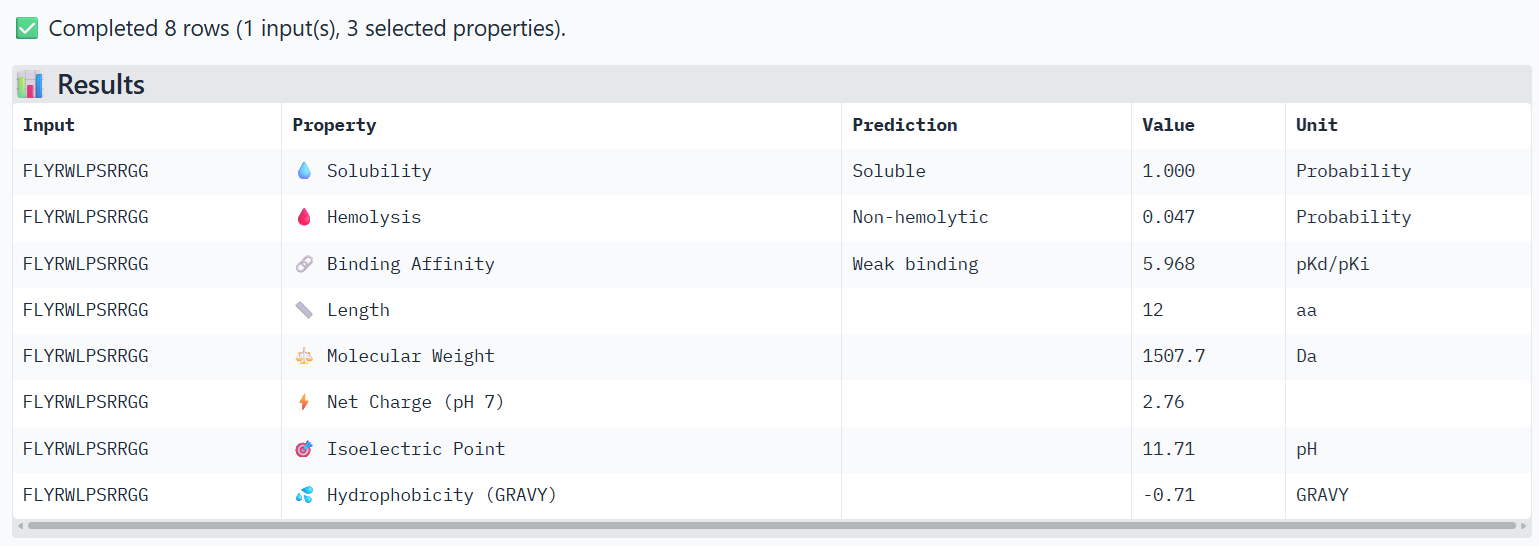
For the main suggested candidate FLYRWLPSRRGG, I obtained these results:

For the other binder candidate WRYPVAAAEWGE, I obtained these results:
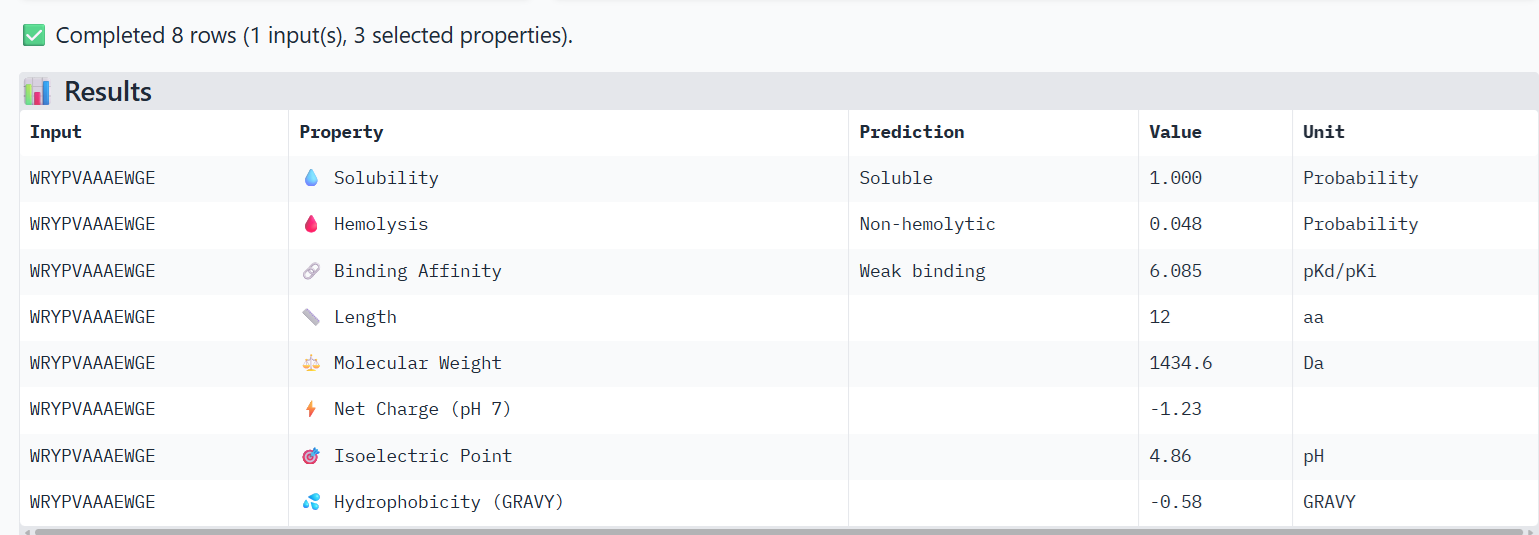

Compare these predictions to what you observed structurally with AlphaFold3. In a short paragraph, describe what you see. Do peptides with higher ipTM also show stronger predicted affinity? Are any strong binders predicted to be hemolytic or poorly soluble? Which peptide best balances predicted binding and therapeutic properties?
I observe that WRYPVAAAEWGE, which has a slightly higher ipTM value than FLYRWLPSRRGG, also shows very slightly higher hemolysis and binding affinity properties, but still not much. Both are considered soluble, non-hemolytic and weak binding.
Choose one peptide you would advance and justify your decision briefly. I’m not sure I’d advance either, because their ipTM scores were so low. Yet, it appears that WRYPVAAAEWGE has slightly better properties than FLYRWLPSRRGG.
Part 4: Generate Optimized Peptides with moPPIt
Now, move from sampling to controlled design. moPPIt uses Multi-Objective Guided Discrete Flow Matching (MOG-DFM) to steer peptide generation toward specific residues and optimize binding and therapeutic properties simultaneously. Unlike PepMLM, which samples plausible binders conditioned on just the target sequence, moPPIt lets you choose where you want to bind and optimize multiple objectives at once.
Open the moPPit Colab linked from the HuggingFace moPPIt model card Make a copy and switch to a GPU runtime. In the notebook: Paste your A4V mutant SOD1 sequence. Choose specific residue indices on SOD1 that you want your peptide to bind (for example, residues near position 4, the dimer interface, or another surface patch). Set peptide length to 12 amino acids. Enable motif and affinity guidance (and solubility/hemolysis guidance if available). Generate peptides. After generation, briefly describe how these moPPit peptides differ from your PepMLM peptides. How would you evaluate these peptides before advancing them to clinical studies?
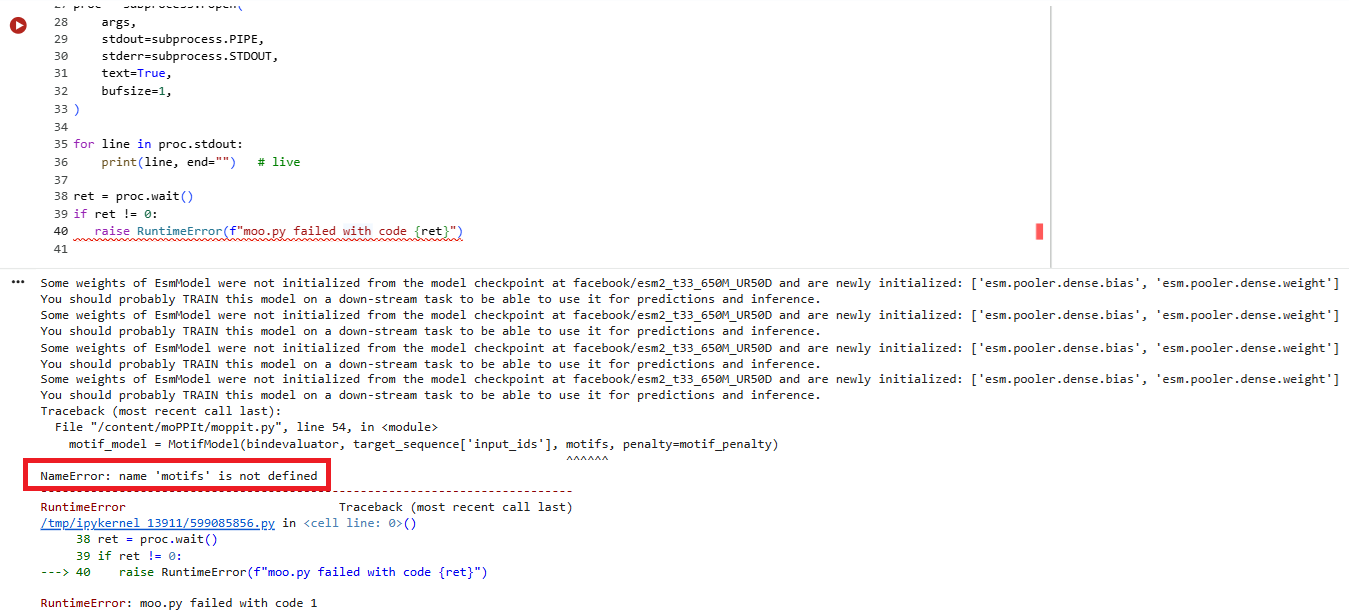
I ran the script and tried to specify a motif value of ‘2’ or ‘3’ - but the next script which generated the peptides complained of an error.
I then unselected ‘motif’ and tried to run it. I got the following sets of results:
'YKQYKHKQLCPI' 0.9623022861778736, 0.7793857455253601, 6.442235946655273
'KKEKNKKKCGLS' 0.9733861181885004, 0.9757760167121887, 7.327633857727051
'KDKKKDKYYCTI' 0.9739618953317404, 0.9253293871879578, 7.365080833435059
I would try to insert A4V mutant SOD1 into animal and then human cell lines and see whether using the peptides reduced the problems associated with A4V.