Week 6 Genetic Circuits Part I: Assembly Technologies
Genetic Circuits Part I: Assembly Technologies
1 What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose? A product page for Fisher Scientific indicate: “Phusion High-Fidelity PCR Master Mix is convenient 2X mix containing Phusion DNA Polymerase, nucleotides, and optimized reaction buffer including MgCl2. Two master mix formulations are available: with HF Buffer (F-531S and F-531L) and with GC Buffer (F-532S and F-532L).”1
The optimised reaction buffer is a mix of chemicals that support the best environment to allow DNA polymerase to work during PCR amplification.2 The DNA polymerase are enzymes that are used to duplicate the genetic information stored in DNA, generating a faithful copy.3
- Phusion DNA Polymerase
- nucleotides
- optimised reaction buffer
2 What are some factors that determine primer annealing temperature during PCR? The annealing temperature, which is the optimal temperature at which the primers can bind to the DNA, is heavily related to the primer melting temperature and the guanine citosine (GC) content of the primer. The melting temperature of the primer is the temperature at which 50% of the DNA duplex separates and becomes single stranded. The melting temperature in turn is dependent on the length of the oligonucleotide sequence and the composition of the DNA molecule. Generally, the annealing temperature should be no more than 5 degrees lower than the melting temperature of the primers. If the annealing temperature is set too low, the the result will feature partial annealing with mismatched bases that produces non-specific amplifications. If it is set too high, it will reduce the PCR yield.
3 There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR focuses on a specific region of DNA defined by primers, whereas restriction enzymes focus on a specific sequence that may occur multiple times. Whereas PCR is used for DNA sequencing and gene isolation, restriction digests are used in cloning. Generally you would use PCR to multiply the number of sequences of interest to replicate and then use restriction digestion to prepare those sequences for cloning and to insert DNA fragments into plasmids.
4 How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning? The main consideration should focus on the overlap length the of the forward and reverse primers. The longer the insert and the larger the number of insert fragments, the longer the primers need to ensure stable annealing. According to a SnapGene video -for a single insert sequence of 0.1 to 0.5kb, the overlapping tails of between 15 to 30 nucleotides should be sufficient. For an insert sequence between 8 and 10 kb, the tail overlap should increase to about 40 bp in length.4
5 How does the plasmid DNA enter the E. coli cells during transformation? Plasmid DNA enters the E. coli cells by passing through temporary pores that appear in the bacteria when it is stressed (typically by heat)
6 Describe another assembly method in detail (such as Golden Gate Assembly)
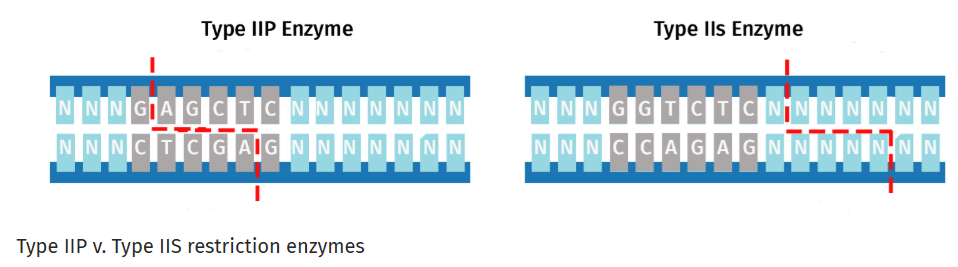
6.1 Explain the other method in 5 – 7 sentences plus diagrams (either handmade or online). Golden Gate Assembly is a DNA assembly method that relies on Type IIS restriction enzymes such as Bsal that cleave DNA outside their recognition sequences. It has two steps that occur within the same reaction:
- Type IIS restriction enzyme digestion
- DNA ligation
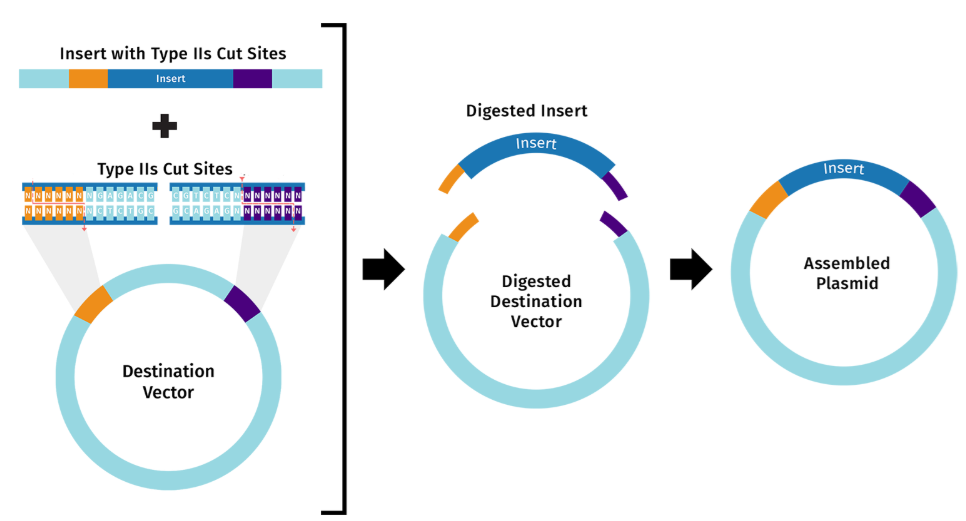
Whereas Gibson Assembly relies on melting overlaps and filling gaps, Golden Gate uses Type IIS restriction enzymes to cleave outside their recognition sequences to produce unique overhangs. Those unique overhangs help make Golden Gate better able to assemble DNA that involves many many insert fragments.
These two diagrams borrowed from the SnapGene web site best describe the key parts of this approach.5


6.2 Model this assembly method with Benchling or Asimov Kernel!
Thermo Scientific™ Phusion High-Fidelity PCR Master Mix with HF or GC Buffer, Fisher Scientific, https://www.fishersci.co.uk/shop/products/phusion-high-fidelity-pcr-master-mix-gc-buffer/10369537 ↩︎
PCR buffers. CliniSciences, https://www.clinisciences.com/en/buy/cat-pcr-buffers-2087.html#:~:text=PCR%20buffers,-Polymerase%20Chain%20Reaction&text=Tris%2DHCl%20is%20the%20primary,and%20activity%20throughout%20thermal%20cycling. ↩︎
DNA Polymerase, ScienceDirect, https://www.sciencedirect.com/topics/neuroscience/dna-polymerase ↩︎
SnapGene, A Detailed Look at Gibson Assembly, Youtube, https://www.youtube.com/watch?v=etPiygqEv5E ↩︎
SnapGene, What is Golden Gate Assembly?, https://www.snapgene.com/guides/golden-gate-assembly ↩︎