Making 3D Printable Green Man whose Face Fluoresces Green with High CO2 Level
Section 1: Abstract
My project features a Bioart installation that involves expressing a biosensor that fluoresces green in the presence of high acidity. This genetic circuit is used as a proxy for a later, more complex task of creating a biosensor that fluoresces green in the presence of high levels of carbon dioxide. The biosensor will feature E. coli that contain plasmids which are engineered so that a Pars promoter that is triggered by low acidity can fluoresce using sfGFP.
The art work will be created based on creating a 3D printed sculpture of a Green Man, an ancient sculptural motif found in medieval architecture that today is associated with Nature, growth cycles and regeneration. The Green Man is a metaphor for Nature, and the art installation will involve growing a culture of biosensor bacteria onto the sculpture. The project will then capture on video whether carbonic acid dripped onto the Green Man face will cause the bacteria to fluoresce green. The desired glowing effect would help demonstrate the resilient response to rising carbon dioxide with more lush growth of the vegetation the Green Man is associated with.
Significance. The Bioart installation that uses the biosensor is about reflecting on the potential use of biosensors to qualitatively express to people rising levels of climate change gases, and on how plant-life may respond to rising carbon dioxide levels.
Broad objective. The main objective is to encourage discussion about Nature’s response to higher carbon dioxide levels and the prospect of using biosensors to encourage people to become more immediately aware and responsive to climate change gases. Other objectives are about examining how the nature of a sculpture surface could best promote culturing of fluorescing bacteria.
Hypothesis. The project will test whether the plasmid I’m able to engineer can be successfully cultured on a Green Man sculpture and glow green in the presence of carbonic acid made from combining baking soda and vinegar.
Specific Aims. The key steps include:
- 3D print a variety of Green Man sculptures that have different surface properties. Culture E. coli that contain a plasmid which ensures they always glow green. Observe which shape type and shape size (small or large) is best suited to culturing bacteria in the next step.
- Design a plasmid that will fluoresce the most when it is exposed to highly acidic liquid dripped onto it.
- Prepare the art installation and video the results.
Methods. These are the biotechnology techniques I will use:
- designing a plasmid using Benchling and Twist.
- using PCR and the Gibson assembly method in the lab to create E. coli that contain the plasmids that feature the genetic circuit I design
- use heat shock to insert the plasmids into the bacteria
- use lab techniques to efficiently paint agar onto the Green Man figure in ways that can promote culturing the bacteria to spread across its surface.
Section 2: Project Aims
Aim 1: Experimental Aim.
Given the time constraints and potential expense I will incur in making the experiment work, I am limiting myself during course time to the modest goal of using a genetic circuit that can use high acidity as a trigger to turn on fluoresce of the GFP protein found in the plasmid I develop. I am essentially going to follow the Week 2 Homework instructions for submitting a Twist order, but use the well-known promoter ‘P asr’, which will trigger based on exposure to high acidity. I also want to better understand how flourescing bacteria would be best cultured on different sculptural surfaces. This is an important area of exploration to promote Bioart: it needs to fluoresce on the sculpture well enough to register a visual effect of glowing green.
Aim 2: Development Aim.
If I manage to accomplish Aim 1, I will review my resources and attempt to design a genetic circuit that is triggered more on detecting HCO3 rather than simply acidity. I would then try to repeat the same experiment using exposure to carbon dioxide gas, likely produced from just burning air in a test tube until the match blows out.
Aim 3: Visionary Aim.
This is where Bioart provides a good support for having a visionary aim. It is meant to encourage more people to become interested in understanding physiological responses to carbon dioxide, in an era where fears of increasing global warming gases are increasing.
Section 3: Background
The theme of this project is not about creating a new novel biotechnology technique for manipulating living matter, nor developing a novel organism using standard techniques such as plasmid engineering. Its innovation is in how an established genetic circuit can be used to make an artistic expression that encourages discussion.
3.1 Synthetic Biology Background
Given the course time constrains and the great breadth of topics in synthetic biology, I decided to ask ChatGPT to help me scope a genetic circuit that would provide the effect I was interested in. I initially asked ChatGPT this prompt: “How could I design a bacteria biosensor that glows green when it is exposed to high concentrations of carbon dioxide? Please include sources.”
Its response suggested three options listed here in increasing levels of difficulty: • Option A: pH-based sensing. This approach uses a crude high acidity as a crude proxy for detecting carbon dioxide dissolved in water (carbonic acid). It has the disadvantage of not specifically triggering on increased exposure to high CO2 levels, but it probably the easiest to do. • Option B: CO2/bicarbonate-responsive regulators. This approach provides a more specific solution but is more complex in that it would involve engineering a transcription factor that is responsive to bicarbonate. • Option C: Synthetic two-component system. This would be the most specific approach, but it is the most complex of the three options. It would involve engineering a CO2-binding regulator and coupling it to a promoter that controlled GFP.
Given that I’m new to the field, there is limited time and that I would need to assess the costs and risks of doing this in the Lifefabs laboratory, I opted for the simplest option A. I leave Option B as something I would like to try after the course finishes. I opted to limit my exploration of the literature to the most common and well tested approaches because my innovation focuses on a new artistic expression than on creating a new innovation in a scientific field I’ve only been exposed to since January.
3.2 Climate Change: Rising levels of carbon dioxide
TO DO
- indicate how levels of carbon dioxide have been higher and lower than today
3.3 Biology Background: The physiological changes in life forms due to carbon dioxide
- cite papers indicating that humans are experiencing higher levels of CO2 in their blood
- describe CO2 effects on bacteria, fungi, insects, mammals
3.2 Bioart Background
TODO
3.2.1 Green Man: from church architectural motif to modern Nature spirit
3.2.2 Applying bacterial growth cultures onto sculpture surfaces
Section 4: Experimental Design, Techniques, Tools and Technology
For my project, I would have ideally wanted to create a plasmid that would use a promoter to cause sfGFP to fluoresce green in the presence of high concentrations of carbon dioxide. Instead, for the initial part of the project I can do in the current class time frame, I opted to develop a pH sensing genetic circuit that would trigger in the presence of a strong acid to act as a kind of proxy for detecting carbonic acid, which is made from carbon dioxide dissolving in water. In a future phase of my project, beyond the class time frame, I would like to try creating a sensor that detected the presence of HCO3 and which is a more specific indicator of carbon dioxide.
4.1 Identify an appropriate promoter sequence and make a Twist order
There are multiple promoter choices that could be used to detect acids. Based on repeatedly asking both ChatGPT and Gemini, I considered P-asr and P-cadBC. I had trouble trying to obtain the exact promoter sequence for PcadBC, but I had little trouble finding the promoter sequence for P-asr.
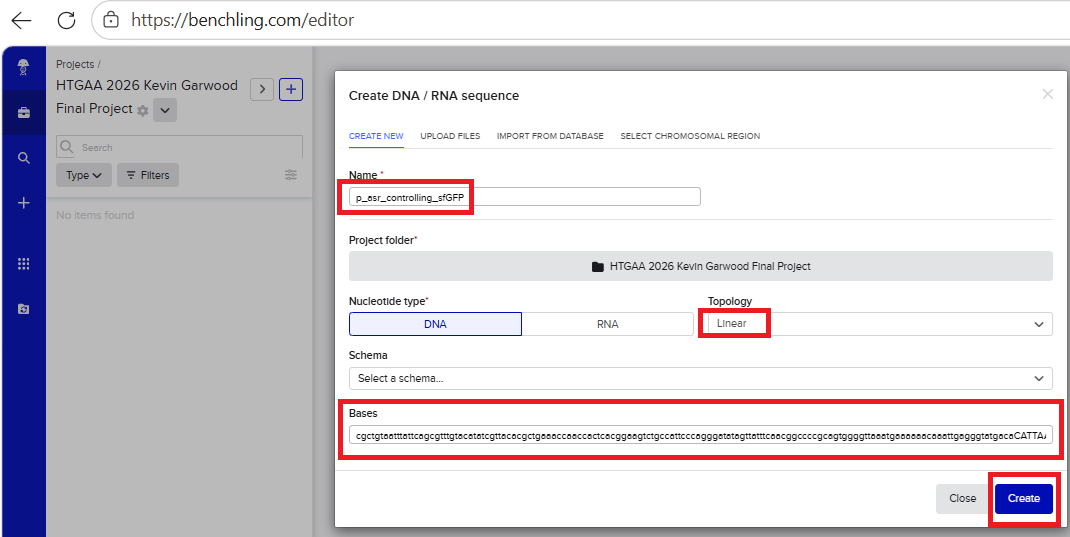
I found a compelling P-asr promoter description on the Registery of Standard Parts web site, under the page asr promoter is a pH-responsive promoter. On that page, I clicked ‘Design’ and then clicked ‘Get part sequence’. I then copied this 140 bp length sequence:
cgctgtaatttattcagcgtttgtacatatcgttacacgctgaaaccaaccactcacggaagtctgccattcccagggatatagttatttcaacggccccgcagtggggttaaatgaaaaaacaaattgagggtatgaca
Borrowing the promoter sequence from BBa_K1231000, I was ready to determine all the parts of the insert sequence:
Promoter: BBa_K1231000 cgctgtaatttattcagcgtttgtacatatcgttacacgctgaaaccaaccactcacggaagtctgccattcccagggatatagttatttcaacggccccgcagtggggttaaatgaaaaaacaaattgagggtatgaca
RBS: CATTAAAGAGGAGAAAGGTACC
Start Codon: ATG
Coding Sequence: Borrowing example from the Homework Week 2 AGCAAAGGAGAAGAACTTTTCACTGGAGTTGTCCCAATTCTTGTTGAATTAGATGGTGATGTTAATGGGCACAAATTTTCTGTCCGTGGAGAGGGTGAAGGTGATGCTACAAACGGAAAACTCACCCTTAAATTTATTTGCACTACTGGAAAACTACCTGTTCCGTGGCCAACACTTGTCACTACTCTGACCTATGGTGTTCAATGCTTTTCCCGTTATCCGGATCACATGAAACGGCATGACTTTTTCAAGAGTGCCATGCCCGAAGGTTATGTACAGGAACGCACTATATCTTTCAAAGATGACGGGACCTACAAGACGCGTGCTGAAGTCAAGTTTGAAGGTGATACCCTTGTTAATCGTATCGAGTTAAAGGGTATTGATTTTAAAGAAGATGGAAACATTCTTGGACACAAACTCGAGTACAACTTTAACTCACACAATGTATACATCACGGCAGACAAACAAAAGAATGGAATCAAAGCTAACTTCAAAATTCGCCACAACGTTGAAGATGGTTCCGTTCAACTAGCAGACCATTATCAACAAAATACTCCAATTGGCGATGGCCCTGTCCTTTTACCAGACAACCATTACCTGTCGACACAATCTGTCCTTTCGAAAGATCCCAACGAAAAGCGTGACCACATGGTCCTTCTTGAGTTTGTAACTGCTGCTGGGATTACACATGGCATGGATGAGCTCTACAAA
7x His Tag: CATCACCATCACCATCATCAC
Stop Codon: TAA
Terminator: Borrowing example from Homework Week 2 CCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATA
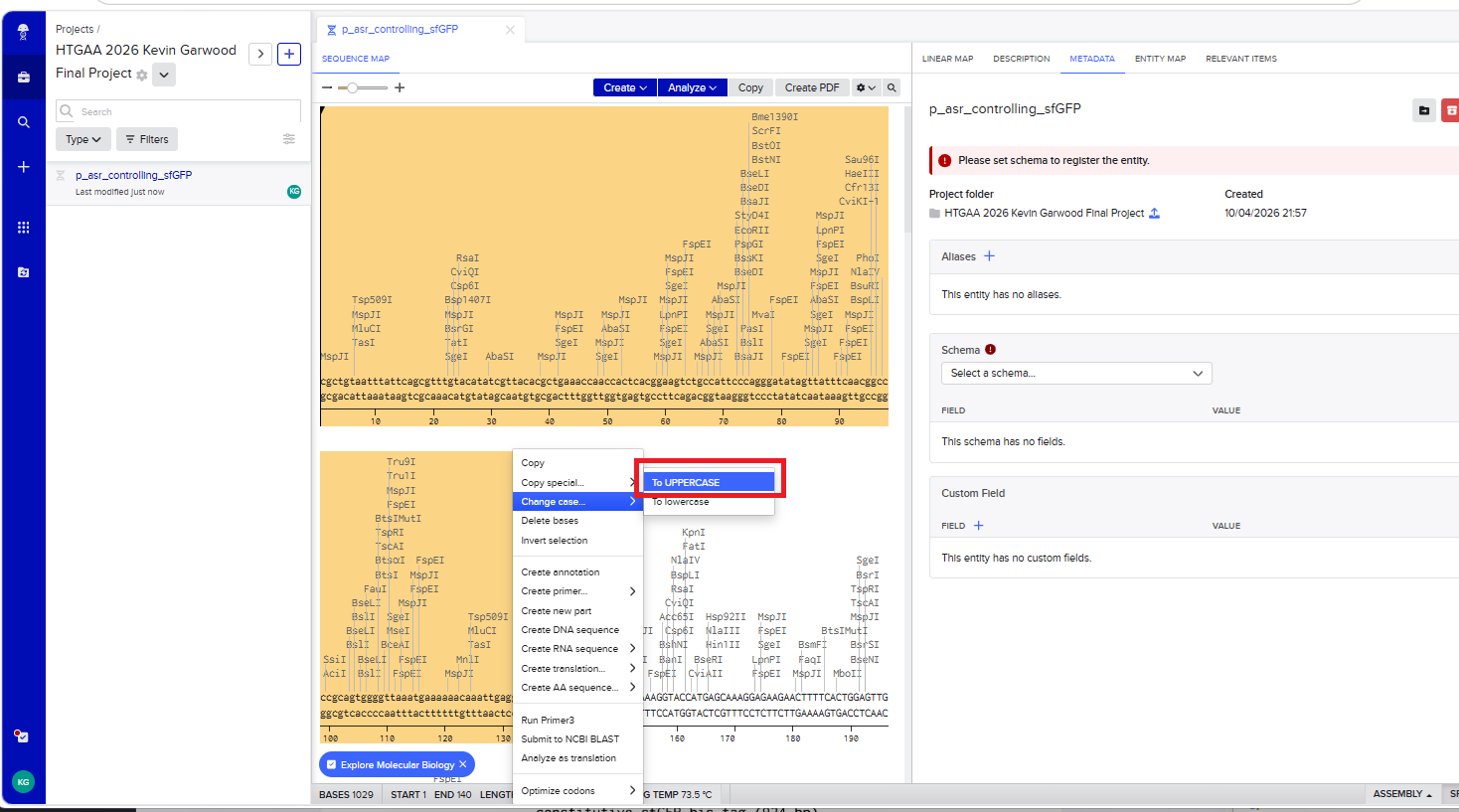
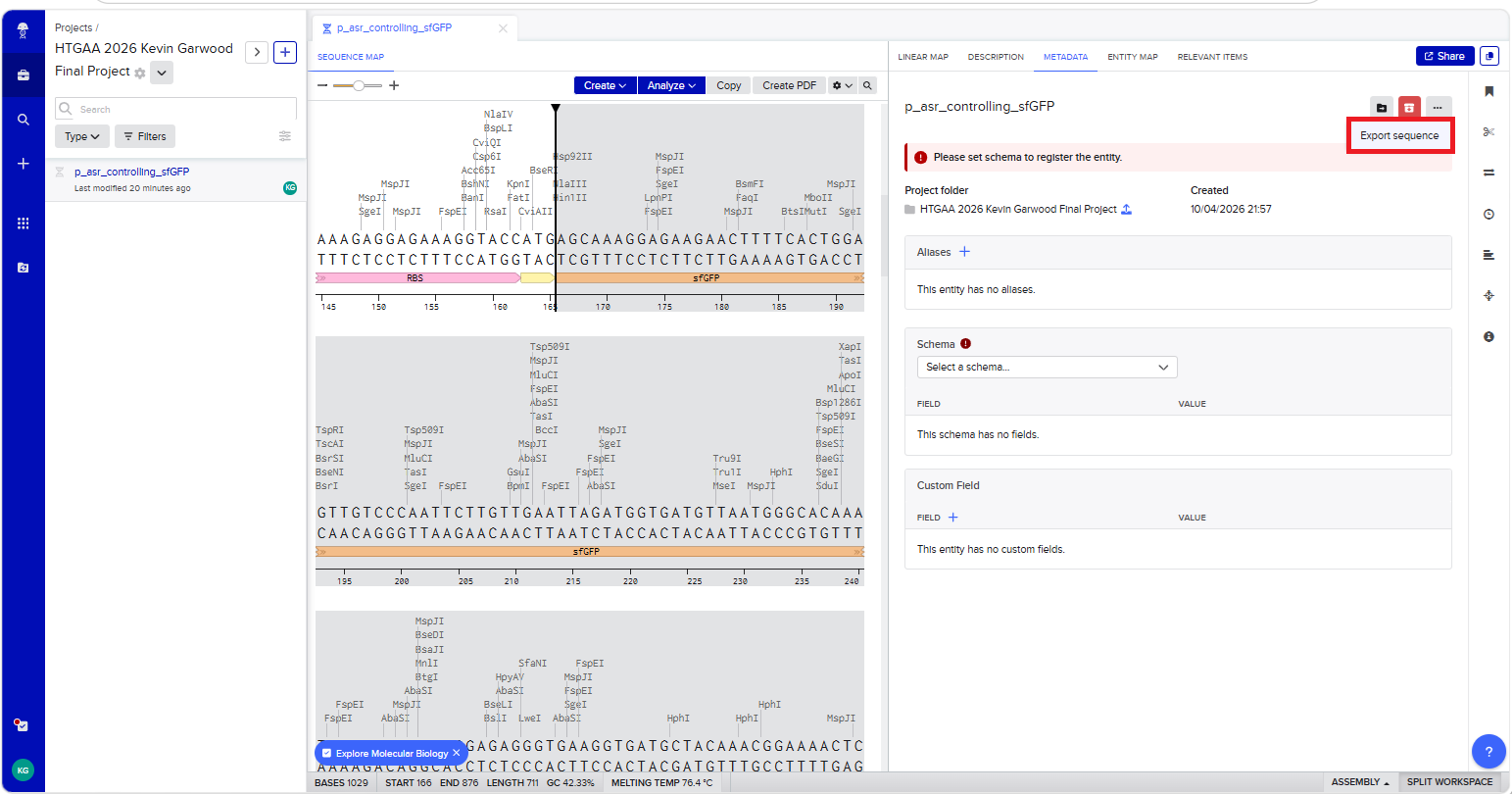
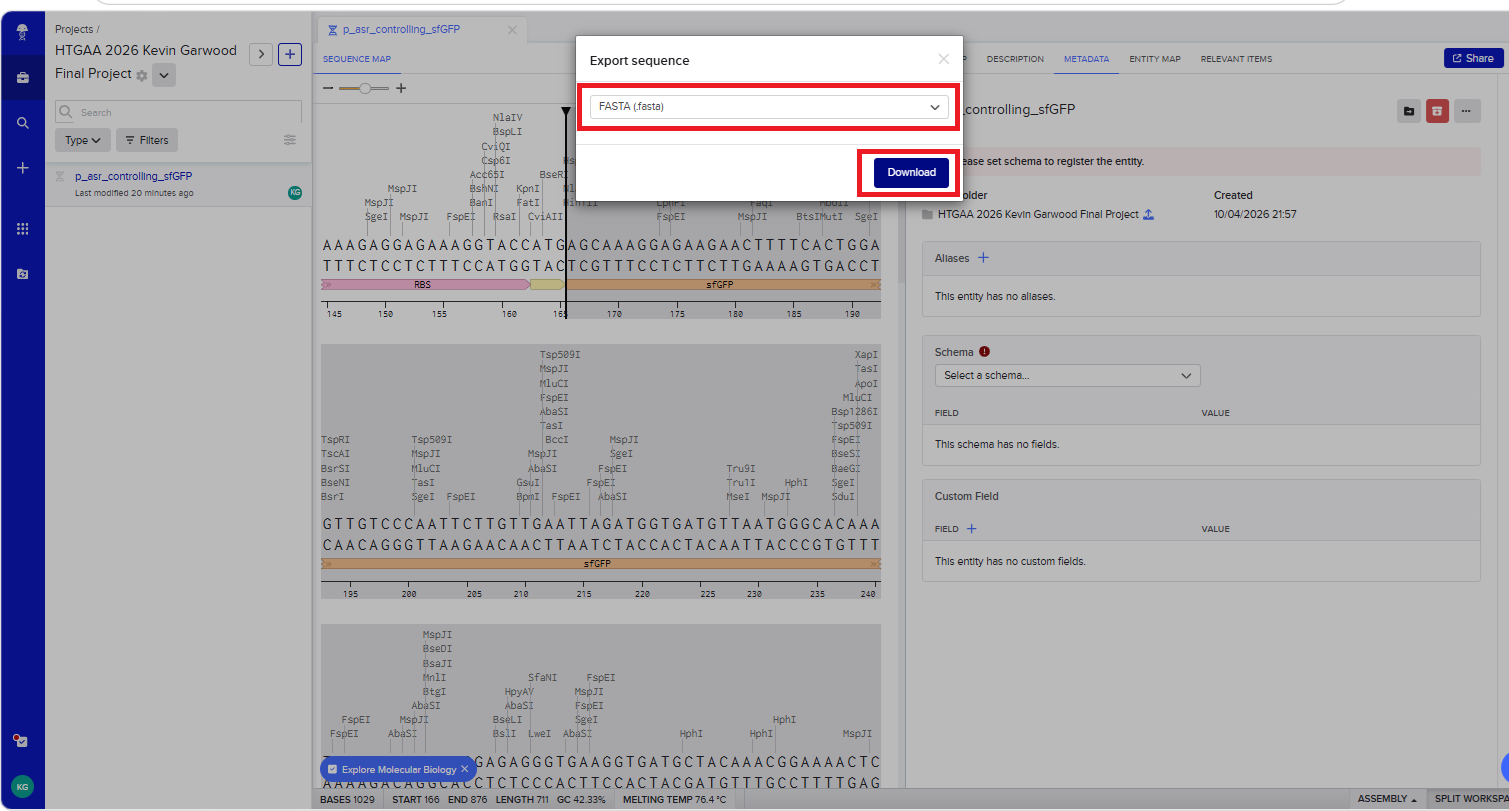
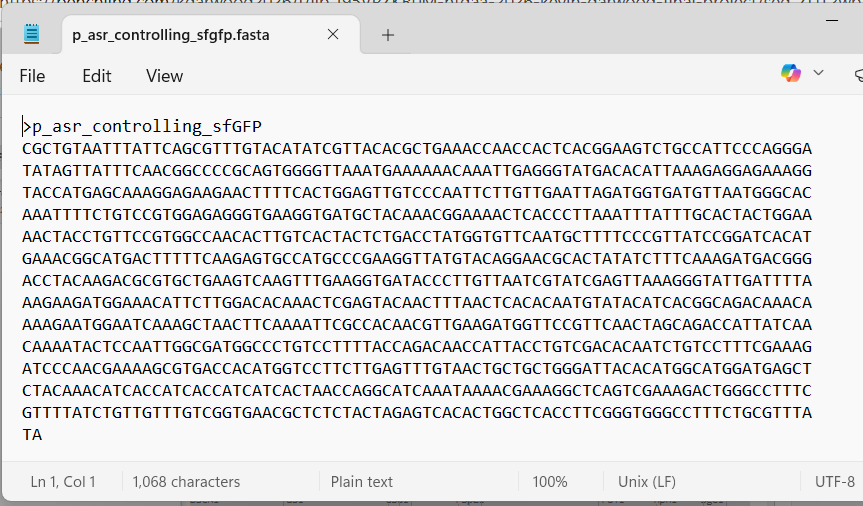
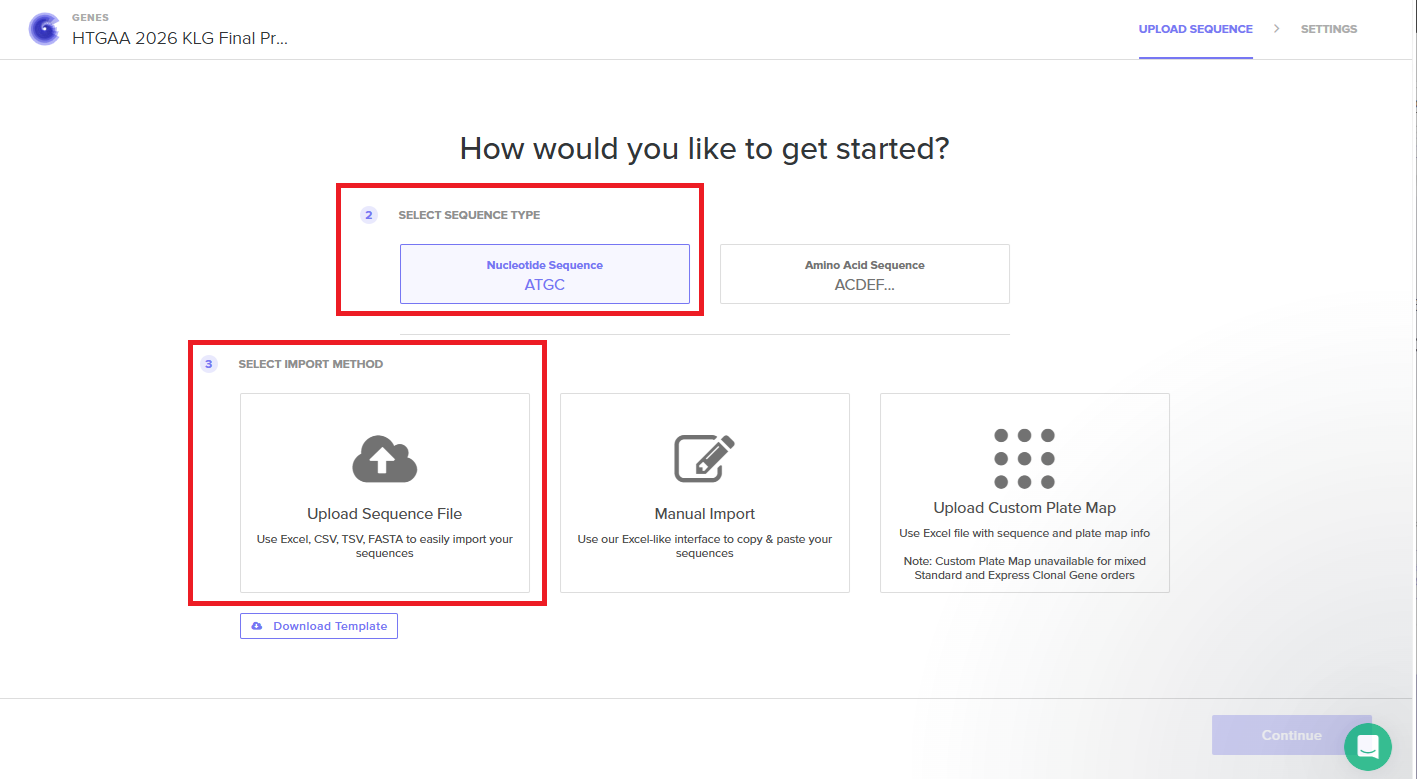
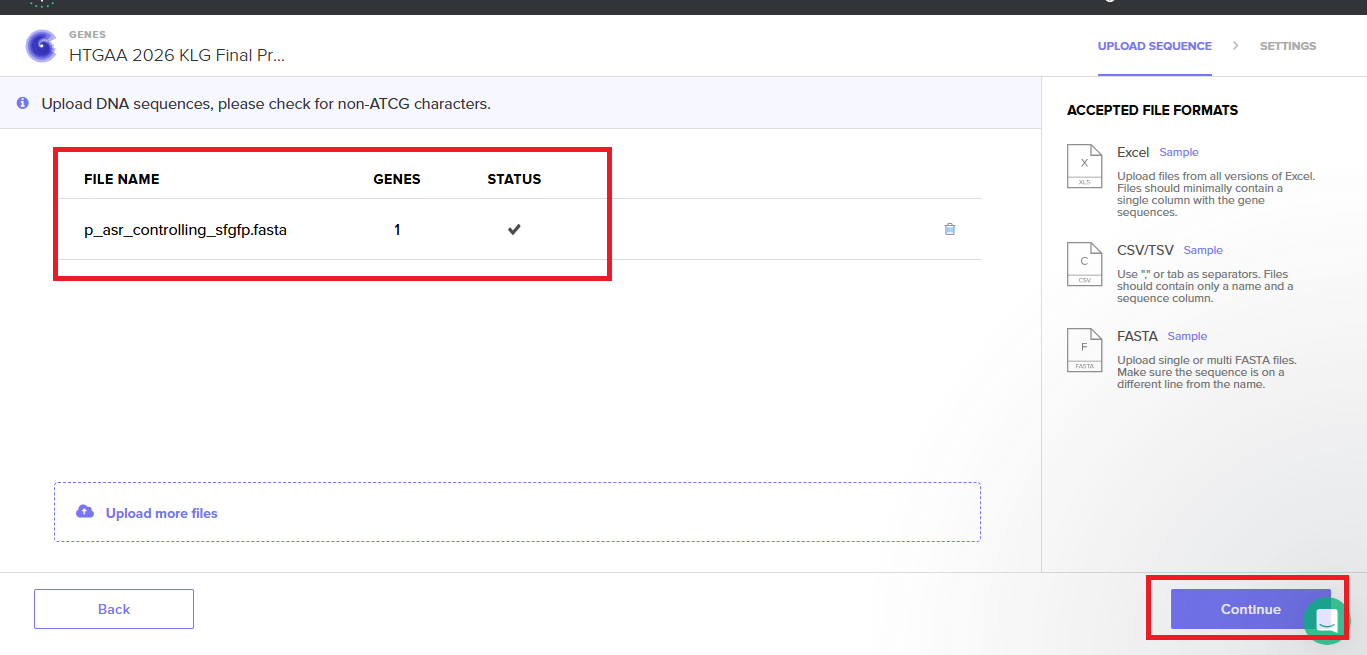
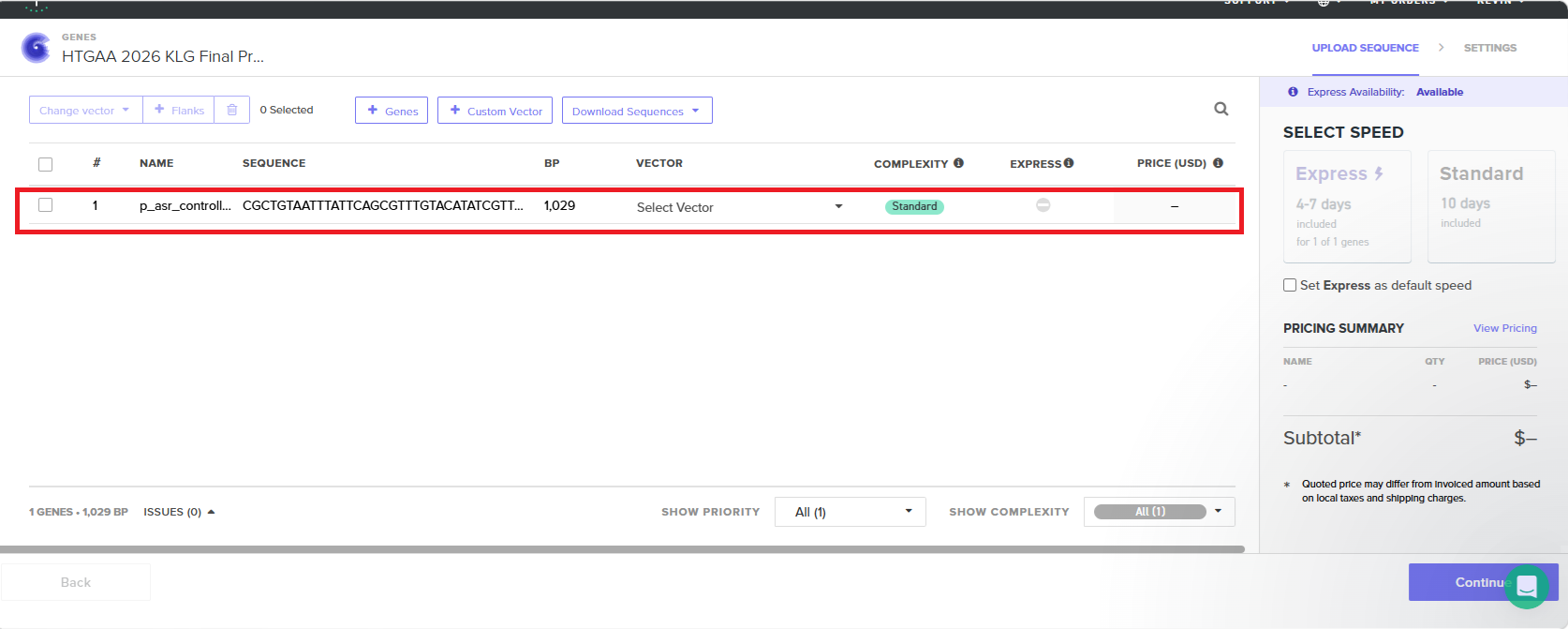
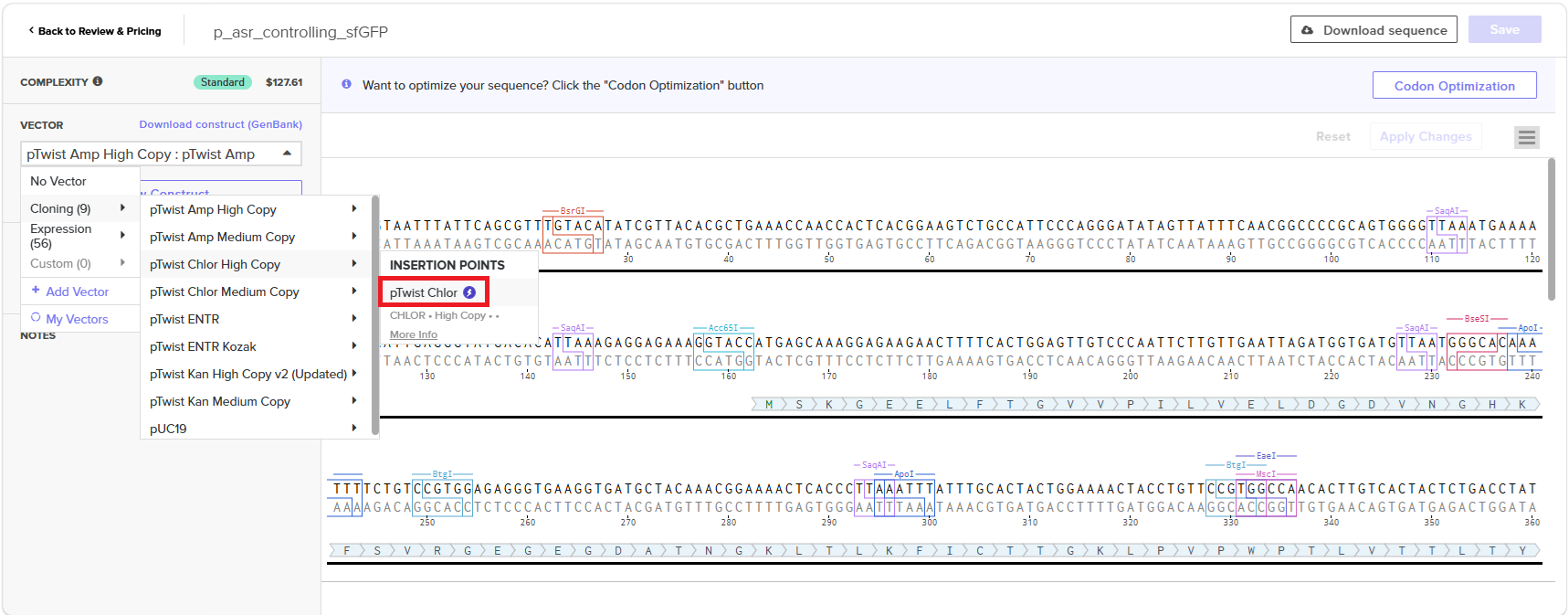
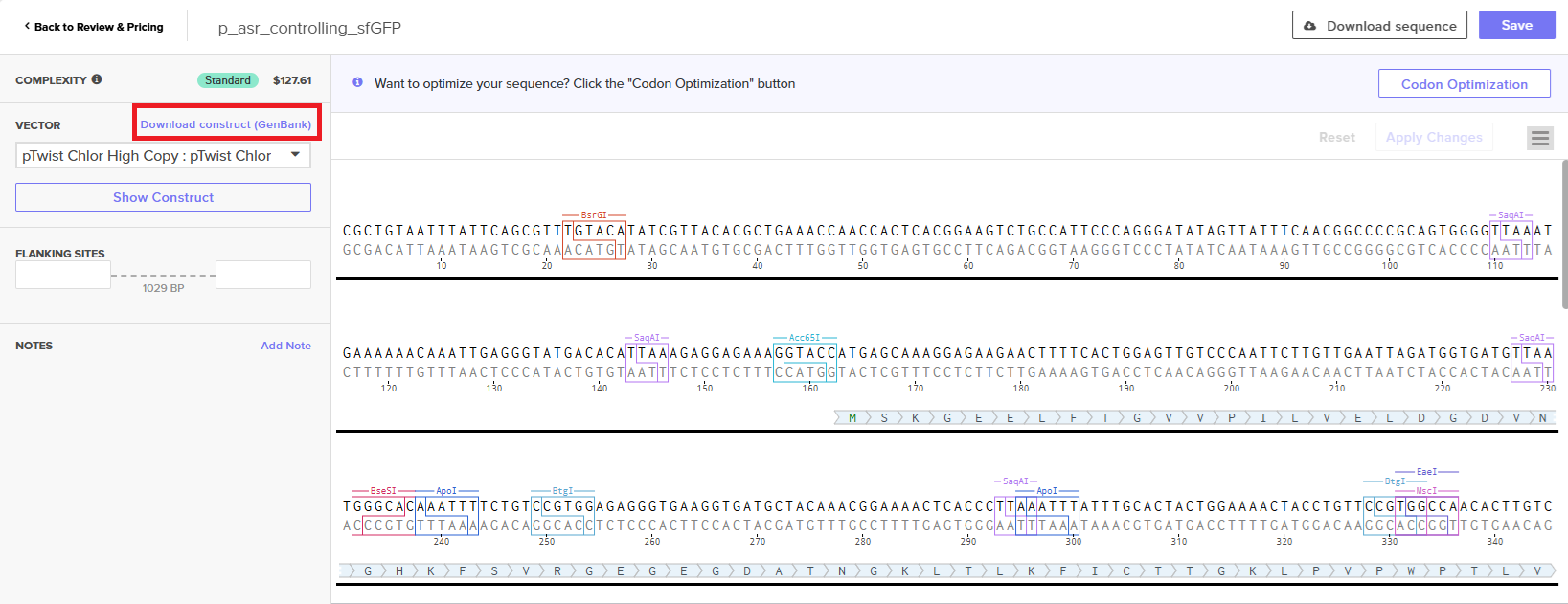
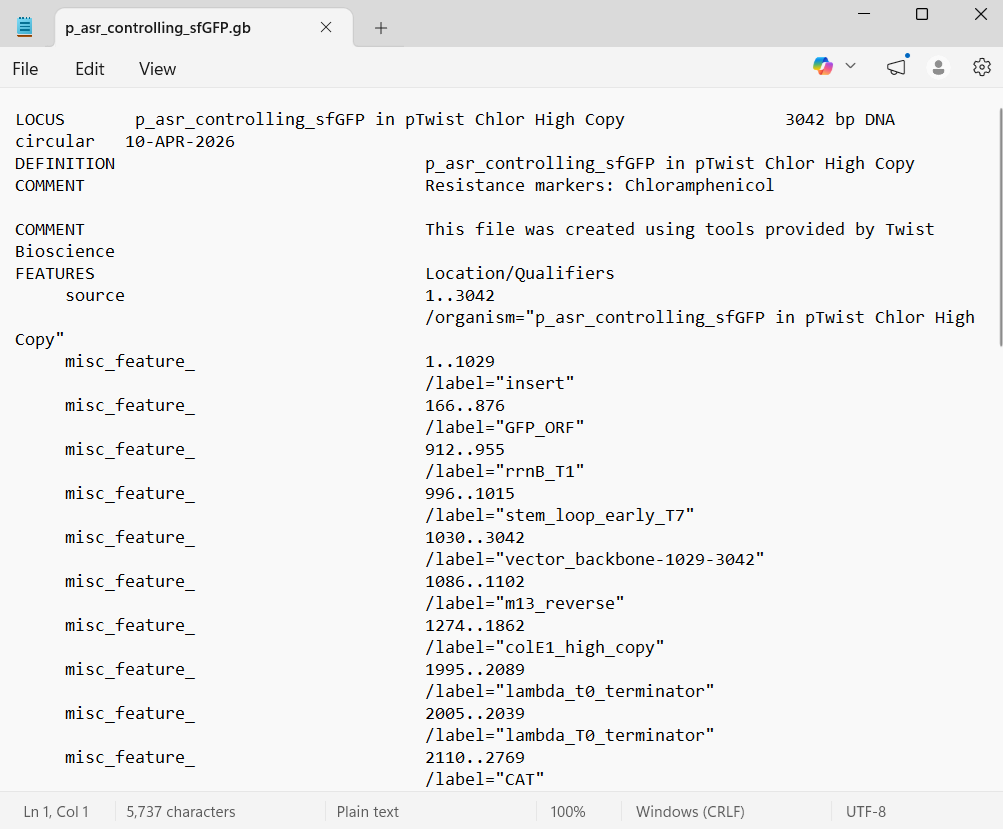
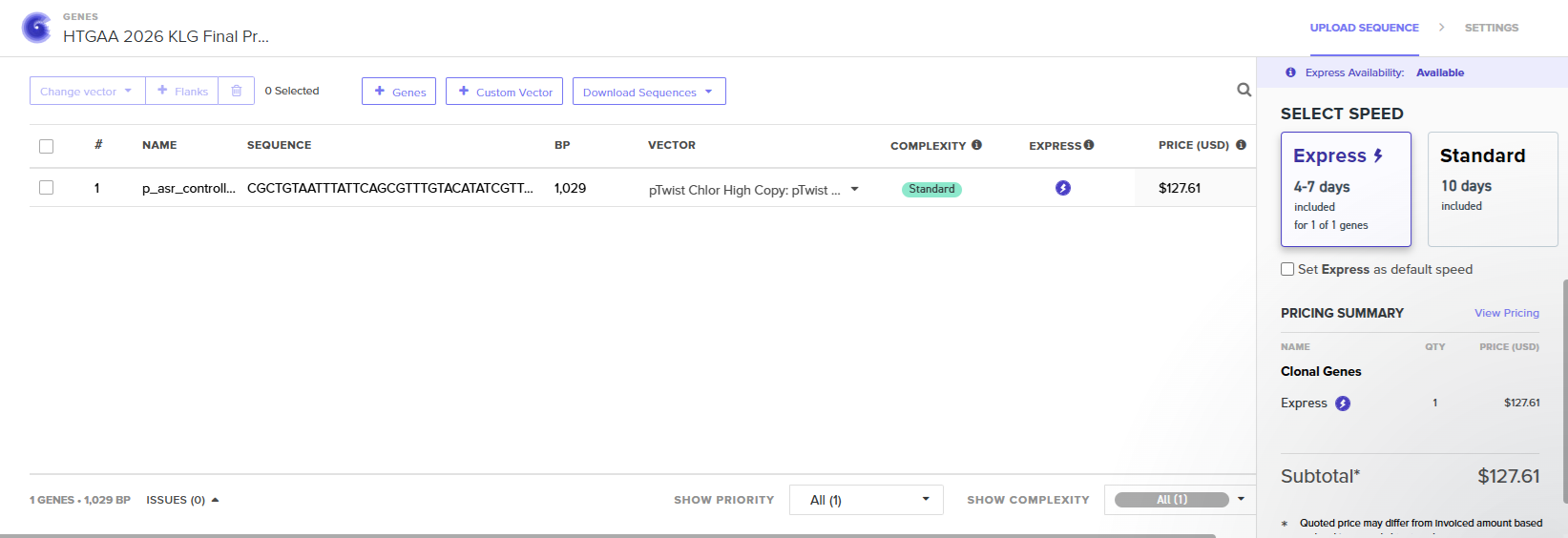
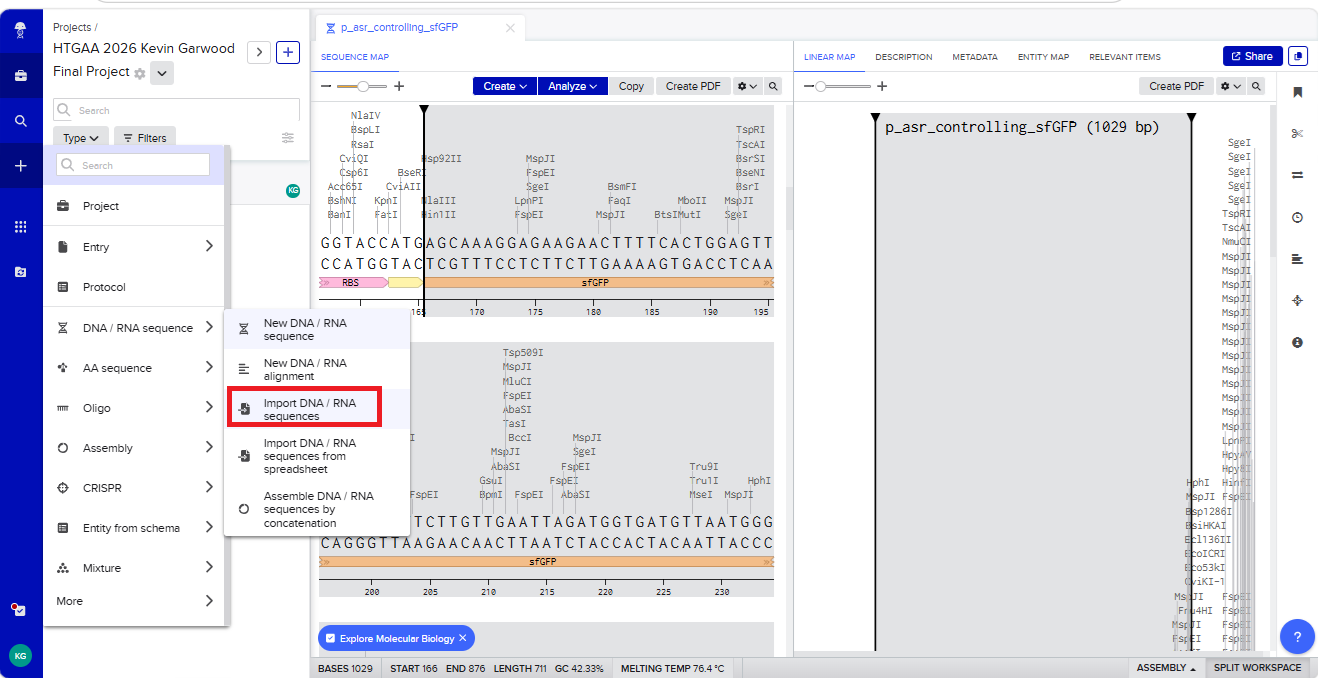
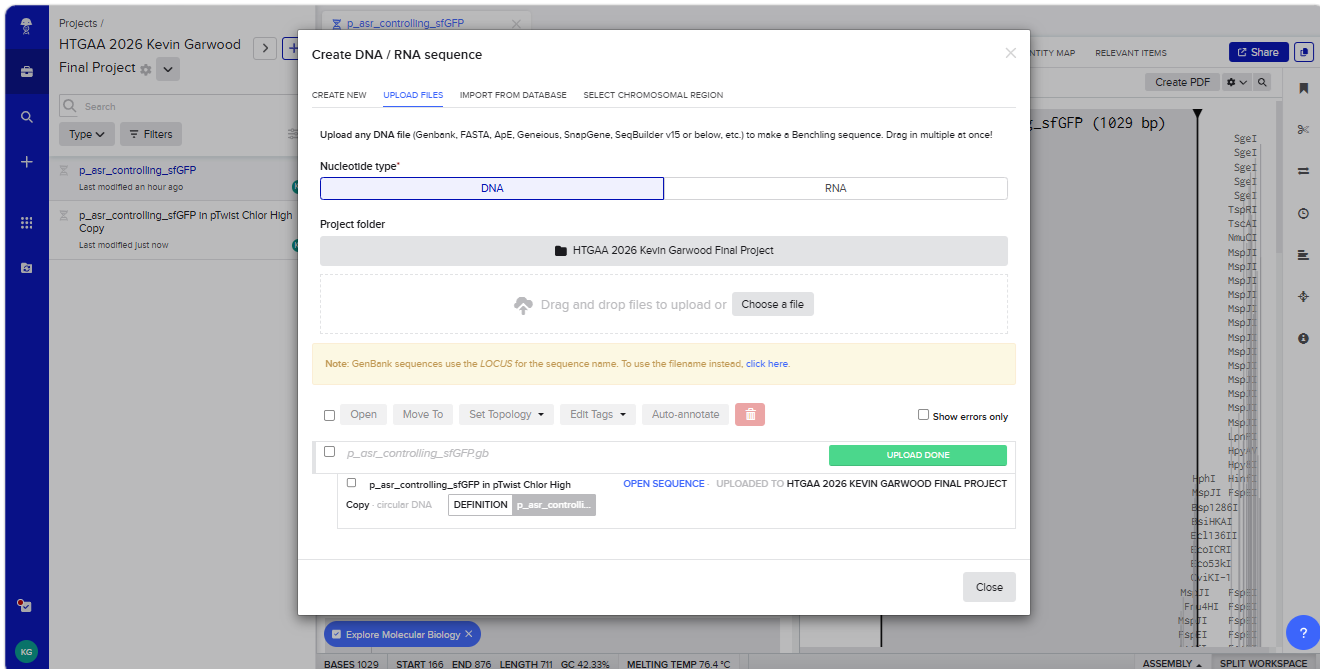
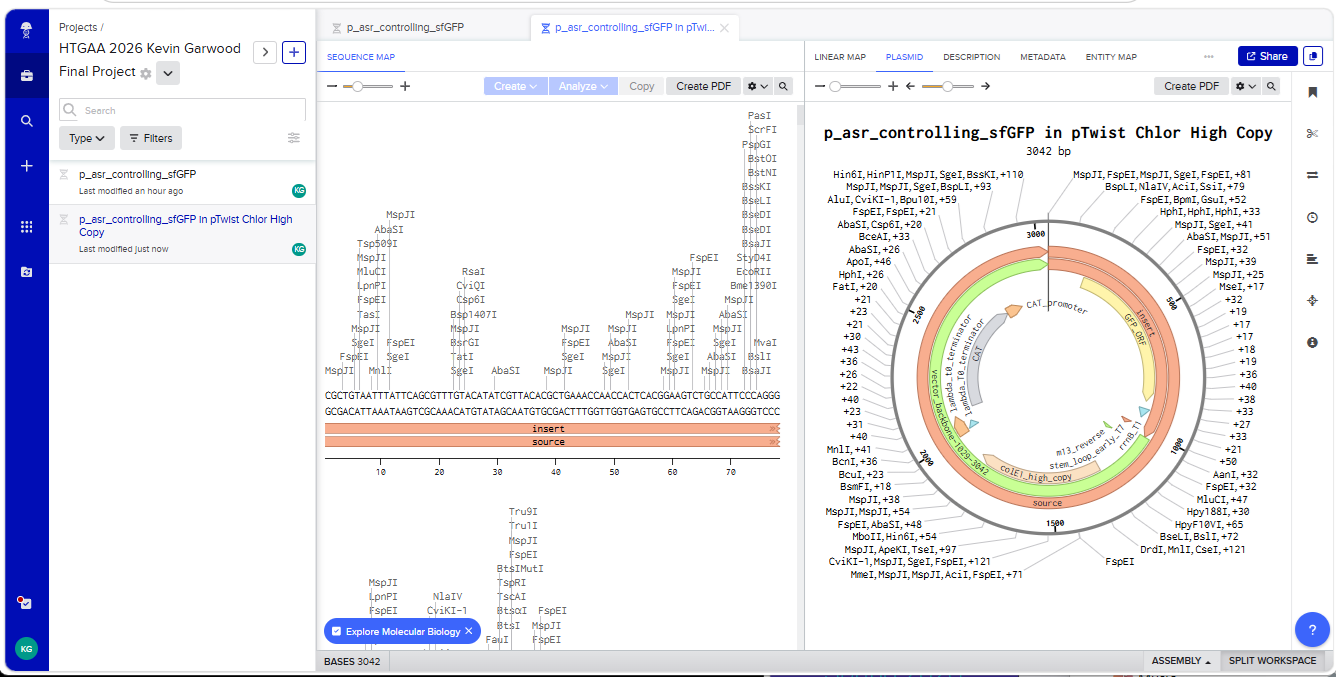
With all of these parts determined, I then followed the Week 2 Homework steps and documented my own progress through it. The order will go to Twist and I will receive the sequence for the engineered insert sequence.
And special thanks to Juan Diego for helping me make some slight modifications…
JD studied my insert sequence and suggested a few modifications:
- some nucleotides to provide space on either side of the RBS so that the RBS had room to bind
- an extra stop codon to help ensure transcription ends
- restriction enzymes to make it easier later one if I wanted to snip out sfGTP and put in another colourful protein!
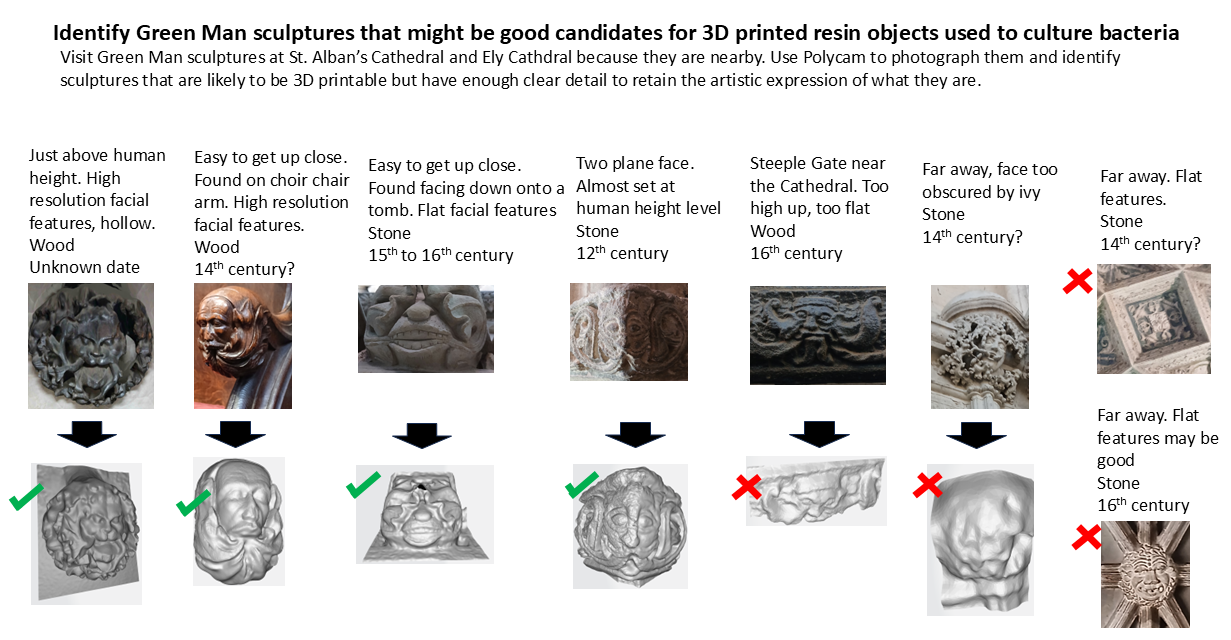
4.2 Identify candidate Green Man sculptures that would be suitable for 3D printing for the project
I used the phone application Polycam to capture multiple examples of Green Man sculptures that I found at both Ely Cathedral and St. Albans Cathedral. A number of sculptures there were not appropriate for 3D printing because they were too far away from my camera to provide much detail.


4.3 Perform bacterial transformation to insert engineered plasmids into competent bacteria
I will follow the AddGene protocol for Bacterial Transformation described at this link. For ease of reading, I’ll copy the steps here.
Take competent cells out of -80°C and thaw on ice (approximately 20-30 mins).
Remove agar plates (containing the appropriate antibiotic ) from storage at 4°C and let warm up to room temperature and then (optional) incubate in 37°C incubator.
Mix 1 - 5 μl of DNA (usually 10 pg - 100 ng) into 20-50 μL of competent cells in a microcentrifuge or falcon tube. GENTLY mix by flicking the bottom of the tube with your finger a few times.
Note: Transformation efficiencies will be approximately 10-fold lower for ligation of inserts to vectors than for an intact control plasmid.Incubate the competent cell/DNA mixture on ice for 20-30 mins.
Heat shock each transformation tube by placing the bottom 1/2 to 2/3 of the tube into a 42°C water bath for 30-60 secs (45 secs is usually ideal, but this varies depending on the competent cells you are using).
Put the tubes back on ice for 2 min.
Add 250-1,000 μl LB or SOC media (without antibiotic) to the bacteria and grow in 37°C shaking incubator for 45 min. Note: This outgrowth step allows the bacteria time to generate the antibiotic resistance proteins encoded on the plasmid backbone so that they will be able to grow once plated on the antibiotic containing agar plate. This step is not critical for Ampicillin resistance but is much more important for other antibiotic resistances.
Plate some or all of the transformation onto a 10 cm LB agar plate containing the appropriate antibiotic. Note: We recommend that you plate 50 μL on one plate and the rest on a second plate. This gives the best chance of getting single colonies, while allowing you to recover all transformants. Note: If the culture volume is too big, gently collect the cells by centrifugation and resuspend in a smaller volume of LB so that there isn’t too much liquid media on the agar plates. If the agar plate doesn’t dry adequately before the cells begin dividing, the bacteria diffuse through the liquid and won’t grow in colonies.
Incubate plates at 37°C overnight.
4.4 Create mix of LB agar with chloramphenicol to pour into five petri plates.
For this part, I will follow the protocol for making the mix we followed in the Lifefabs labs. For convenient reading, I will copy the steps here:

- Clean the fume hood area with isopropyl alcohol.
- Put on labcoat, goggles and a face mask. Put on blue gloves.
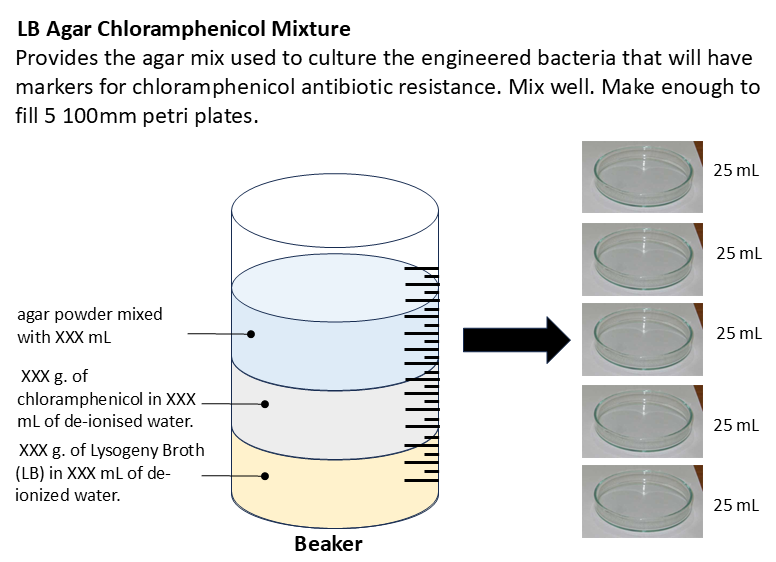
- In the fume hood, prepare a mixture of LB, agar, and chloramphenicol (How exactly again??). Make enough to pour 125mL for five plates.
- Dispose of any waste in the Clinical Waste bin.
- Clean fume hood surface with isopropyl alcohol.
- Thoroughly wash hands.
4.5 Culture engineered bacteria in petri plates filled with LB agar chloramphenicol
In this part of the protocol I will culture the engineered bacteria into LB agar petri plates that have been prepared with chloramphenicol. These will be used in the next step, when they will be exposed to acid.

- Clean the fume hood area with isopropyl alcohol.
- Put on labcoat, goggles and a face mask. Put on blue gloves.
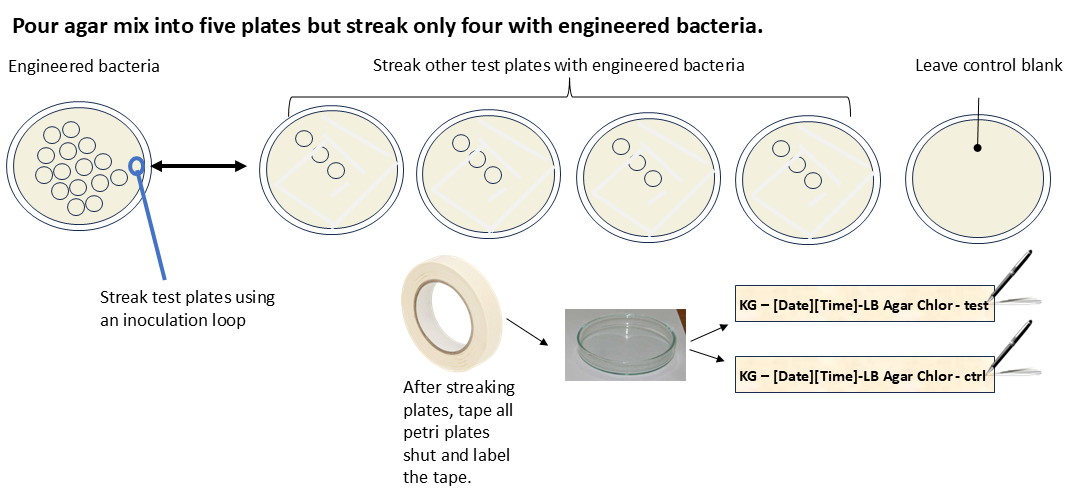
- In the fume hood and using the mix prepared in (4.4), pour 25 mL into each of five sterile petri plates.
- Obtain the petri plate that has been cultured from (4.3).
- Using a large innoculation loop, streak four petri plates.
- Leave the fifth petri plate empty.
- Put the petri plate from (4.3) back into storage to save for any future batches.
- Seal all the petri plates and use plate tape to seal them shut.
- Write “KG - [date][time] - LB agar chlor” on the side of each one.
- Place them in a cupboard that will be kept at room temperature.
- Dispose of any waste in the Clinical Waste bin.
- Clean fume hood surface with isopropyl alcohol.
- Thoroughly wash hands.
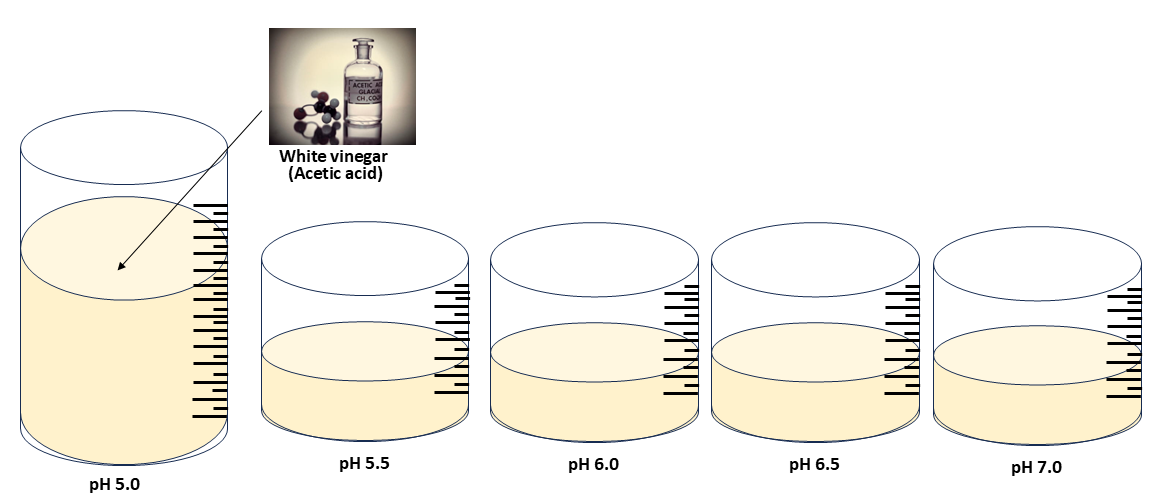
4.6 Apply solutions of acid pH 5.0, 5.5, 6.0, 7.0 onto plates cultured with the engineered bacteria.
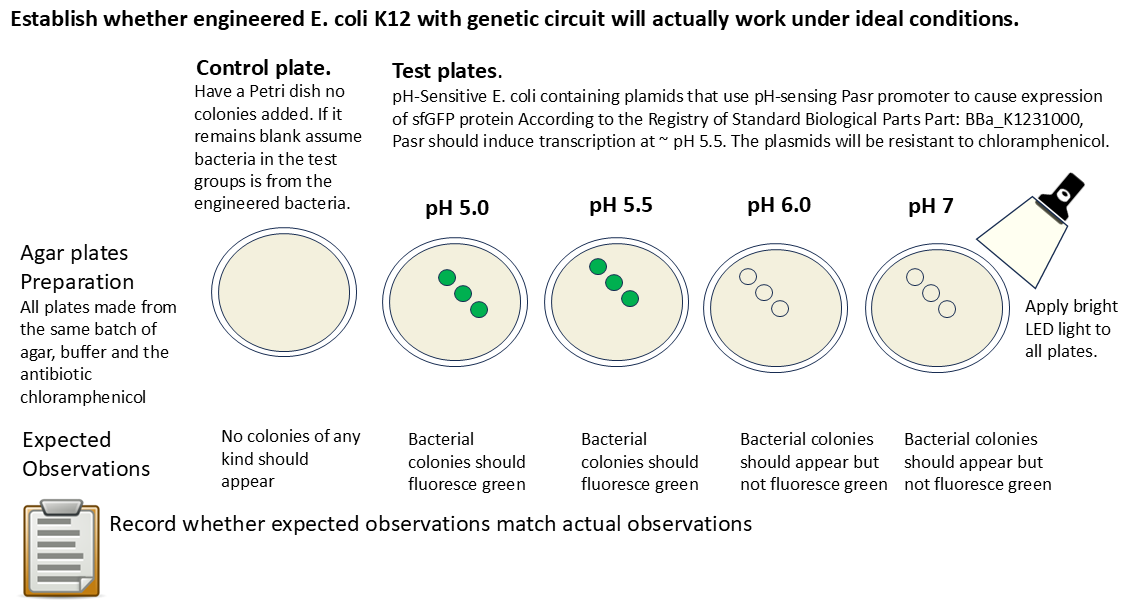
In this part, I will test to make sure that the genetic circuit I ordered through the Twist plasmid order actually works. The circuit involves the promoter P-asr sensing acidic conditions and inducing the production of sfGFP at ~ pH 5.5. The expected pH sensing activity is described in Viral Patel’s protocol.
In the installation phase, the Green Man figure will be exposed to carbonic acid because the Bioart is meant to emphasis the Green Man’s reaction to increased carbon dioxide. However, to show that the plasmids will work well at the expected pH level, I will need to use an acid that has a stable pH and carbonic acid quickly changes into water and carbon dioxide. Therefore, for this part, I will just use acetic acid made from white vinegar to create solutions at different pH. Unlike solutions of carbonic acid, the solutions of diluted acetic acid should remain at a stable pH.
- Obtain a bottle of white vinegar.
- Clean laboratory bench space with isopropyl alcohol.
- Put on labcoat, goggles and a face mask. Put on blue gloves.
- Prepare a 500mL solution of citric acid that has pH 5.0 in a beaker.
- Obtain four other beakers, one for each pH of 5.0, 5.5, 6.0, 7.0.
- From this solution, make dilution calculations to make solutions of pH 5.5, pH 6.0, pH 7.0. Verify pH values with a pH meter.
- Obtain the petri plates made in (4.5). Remove the petri tape. Open
- Use a pipetter to pipette 10mL of each solution into a plate. Leave the fifth plate alone. Replace 20 mL pipette tip each time.
- Wait a couple of minutes.
- Darken the lab area and shine a flashlight onto the plates.
- Record observations. The control should show no bacteria, indicating that the storage of all of them did not incur any contamination. The plates with pH 5.0 and 5.5 should fluoresce. The plates with pH 6.0 and 7.0 should not. The control should not show any bacterial
- Dispose of any waste in the Clinical Waste bin.
- Clean laboratory bench space with isopropyl alcohol.
- Thoroughly wash hands.
4.7 3D print four Green Man sculptures.
In this part, I will export *.STL files from Polycam and submit them to one of the Lifefabs 3D printers. Each model will be scaled such that the longest side will be 100mm, the diameter of a typical petri plate.
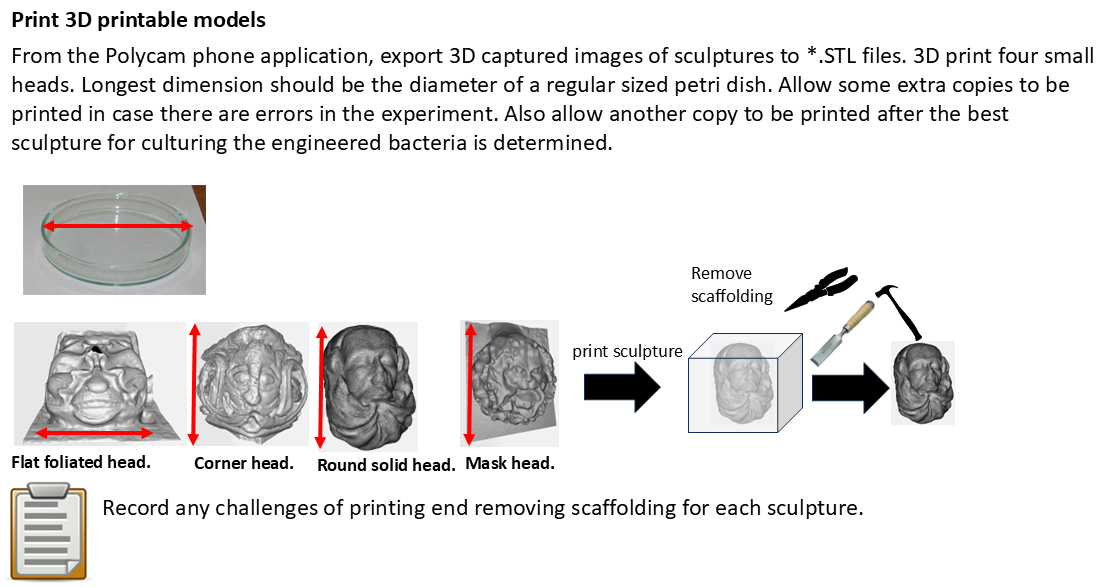


- Obtain enough resin to make four 100 mm3 resin cubes.
- Submit each *.STL model to the 3D printer
- Remove each 3D printed object and use pliers to remove any scaffolding produced within each cube.
4.8 Build a small simple bioreactor that can accommodate the Green Man sculptures.
The final installation will involve culturing the engineered bacteria onto the surface of the sculpture that will be best able to help them spread and show the best definition of the Green Man features. Applying the bacteria directly to resin surfaces likely will not work and the resin itself might be slightly bactericidal. Painting a thin agar layer onto shapes will not work if exposed to the open air because they will dry out. Also, the bacterial colonies need frequent nourishment to continue growing.
This step requires me to build a very small bioreactor which will have space to accommodate a 100mm x 100 mm x 100mm Green Man sculpture. Make the dimensions 150mm x 150 mm x 150 mm.
The protocol for building a small bioreactor involves these steps: … …
4.9 Culture ‘Always On’ GFP-bearing E. coli Strain K12.
This part is meant to grow a supply of E. coli strain K12 bacteria that just contain GFP. The stock should already be obtainable from Lifefabs, and the bacteria will have plasmids where the GFP will always fluoresce.


- Obtain a petri plate of GFP-bearing E. coli Strain K12.
- In a fume hood, follow (4.4) to create enough LB agar with chloramphenicol to pour 25mL in one petri plate.
- Using an innoculating loop, streak GFP-bearing E. coli strain K12.
- Seal the petri plate and use plate tape to seal them shut.
- Write “KG - [date][time] - GFP LB agar chlor” on the side of each one.
- Place them in a cupboard that will be kept at room temperature.
- Dispose of any waste in the Clinical Waste bin.
- Clean fume hood surface with isopropyl alcohol.
- Thoroughly wash hands.
4.10 Observe how GFP-bearing E. coli strain K12 grows on each of the four sculptures put in a bioreactor
This step will likely take the most time to do. For each sculpture, I will cover it in culture of ‘growth broth’ and lower it into a small bioreactor whose dimensions will accommodate the sculptures.
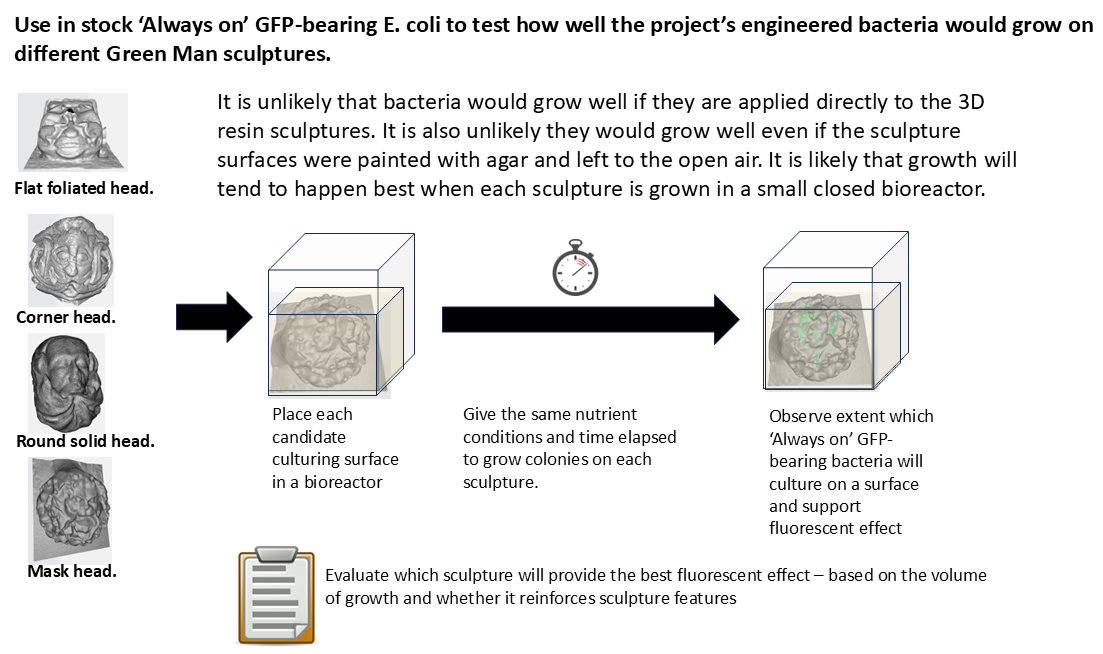

For each sculpture
- Clean the fume hood area with isopropyl alcohol.
- Put on labcoat, goggles and a face mask. Put on blue gloves.
- Clean the bioreactor, spray with isopropyl alcohol and then wipe dry with paper towel.
- Prepare a culture of growth broth.
- Obtain the GFP culture from (4.9)
- Prepare a mix of LB agar with choramphenicol that can be painted onto the sculpture surface. Assume about 25 mL painted in a thin layer with a paint brush.
- Paint the sculpture with a layer of the agar mix. Let the mix gel and then streak the GFP culture onto the gel.
- Lower the sculpture into the bioreactor.
- Pour enough ‘growth broth’ into the bioreactor until it covers the edges of the sculpture face.
- Seal the bioreactor and ‘start’ it. Let it run for a day.
- Place the bioreactor into a cupboard.
- Wait a day.
- Take the bioreactor out and observe the results. Evaluate the performance of the growth by how well it glows and how well it highlights the definition of the sculpture features.
4.11 Create the scenery that will surround the Green Man sculpture
The goal of this is to help produce a set prop that will provide some atmosphere for the Green Man to fluoresce.
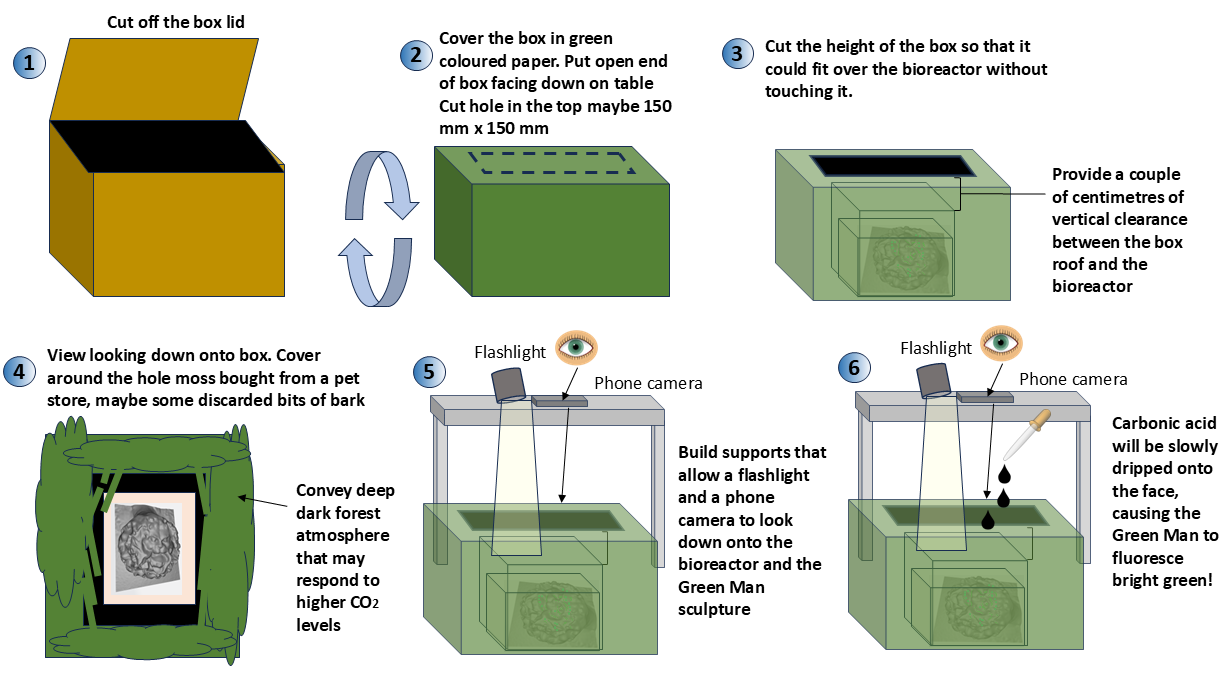

- Obtain a cardboard box. Remove the lid. Cover it in a green piece of coloured paper obtained from an art supply store. Use a scissors to cut as necessary.
- Assume the video will be shot from above onto a dish that contains the sculpture cultured with the engineered bacteria. Turn the box upside down onto a table and cut a hole in the top of the box.
- Cut the sides of the box so they will be about 400 mm high on the table. It should include enough clearance height to not touch the maximum dimension of the Green Man sculpture, which should be at most 100 mm. The height of the box should darken the area where the Green Man sculpture will appear.
- Surrounding the hole, cover the lid with spaghnum moss bought from a pet store and maybe some discarded bark. Glue moss to cardboard.
- Develop some kind of scaffolding which will allow a video to be filmed above the sculpture. It should also be able to hold a narrow beamed light that can shine onto the face of the sculpture.
4.12 Prepare a solution of carbonic acid that is initially about pH 5.0
Having established that the engineered bacteria will fluoresce under acidic conditions, I can now be confident that I can make a solution of carbonic acid that is initially about ph 5.0. Note that it cannot be too low, or it will likely disable the genetic circuit and/or kill the bacteria.
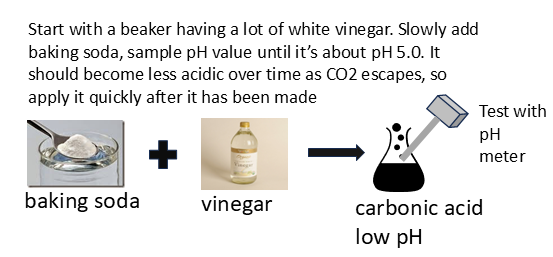

- Obtain baking soda.
- Obtain a bottle of white vinegar
- Carefully combine both in a beaker and measure the pH using a pH meter.
- Obtain a dropper or a pipetter. If using a pipette device, allocate one disposable pipette tip.
4.13 Make the Bioart installation
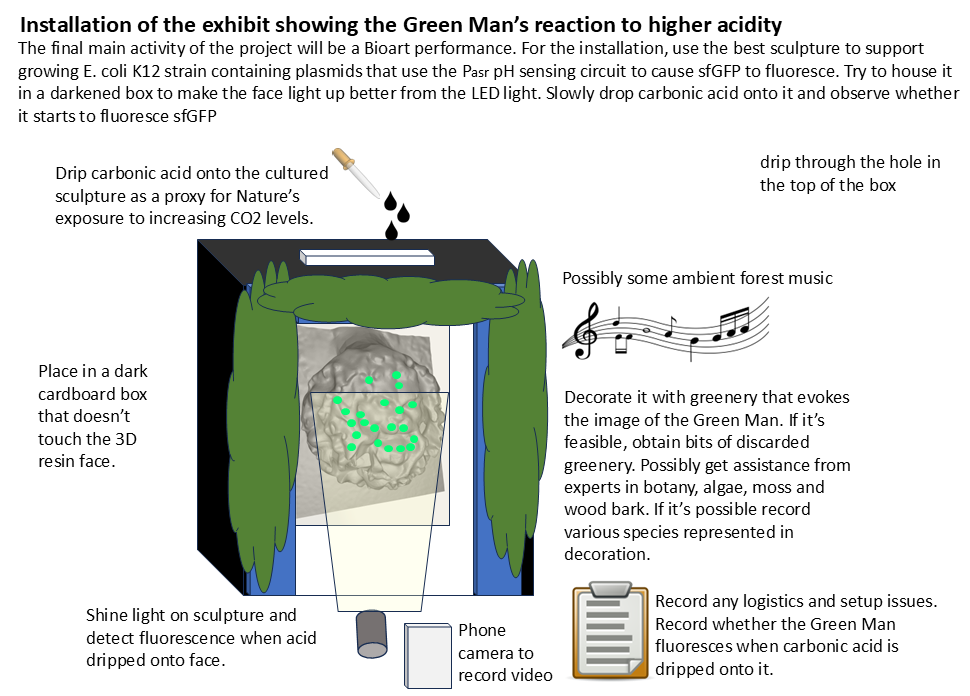

- Repeat (4.10) but this time use the engineered bacteria culture obtained in (4.3).
- Clean laboratory bench space with isopropyl alcohol.
- Put on labcoat, goggles and a face mask. Put on blue gloves.
- Take the bioreactor out of the cupboard. Open the lid of bioreactor
- Use a pipette device with a disposable pipette tip to pipette out most of the nutrient broth, which should leave the sculpture looking wet and covered in the bacteria colony.
- Cover the bioreactor with the scenery made from the cardboard box.
- Set up the video camera on my phone.
- Shine light through the hole onto the sculpture in the bioreactor.
- Start filming. Initially there should be no green fluorescence. Using a dropper or pipetter, slowly drip about 10 to 15 mL of carbonic acid onto the bacterial colony growing on the Green Man sculpture. Pause occassionally and observe whether it begins to fluoresce.
- Leave the video camera to run for awhile to capture what is expected to be a maximal level of fluorescence.
- Record observations.
- Dispose of the cardboard box prop. It should not touch the bioreactor at all and therefore should be safe to dispose of in normal waste.
- Dispose of all of the broth and the Green Man sculpture in the clinical waste bin.
- Clean out the bioreactor, spray it with isopropyl alcohol, then wipe clean.
- End of Bioart exhibit!
Section 5 : Results & Quantitative Expectations
5.1 Findings from identifying potential Green Man sculptures to 3D print
5.2 Findings about which sculpture surface provided the best option for culturing GFP-bearing bacteria.
TO DO
5.3 Findings about how well the genetic circuit worked
TO DO