Week 7 HW: Genetic Circuits II, Fungal Materials, and First DNA Twist Order
Week 7 — Genetic Circuits II, Fungal Materials, and First DNA Twist Order
Part 1: Intracellular Artificial Neural Networks (IANNs)
1. What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
IANNs have important advantages over traditional Boolean genetic circuits because they can perform analog computation rather than only binary ON/OFF logic. Classical genetic circuits are useful for implementing logic gates such as AND, OR, and NOT, but they are limited when the biological problem depends on graded signal levels rather than strict binary states.
In contrast, IANNs can assign different weights to different intracellular inputs, combine them through addition or subtraction, and generate a nonlinear output. This makes them more suitable for interpreting real cellular states, where inputs often vary continuously in magnitude. Instead of forcing biology into rigid digital logic, IANNs can classify more subtle and realistic signal combinations.
Another important advantage is that intracellular artificial neurons can be composed into multilayer networks. A single perceptron is limited to linearly separable decision boundaries, but multilayer systems can produce more complex behaviors. In synthetic biology, this is valuable because cellular environments are noisy, multidimensional, and dynamic. An IANN therefore offers a more flexible and tunable framework for state classification than a conventional Boolean circuit.
2. Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
A useful application for an IANN would be the intracellular classification of an infection-like cell state in mammalian cells. Instead of responding to just one biomarker, the circuit could integrate multiple molecular signals that together better represent whether a cell is truly infected or entering a suspicious pathological state.
For example, the system could receive three inputs:
- X1: a signal associated with interferon pathway activation
- X2: a signal associated with inflammatory signaling such as NF-kB activity
- X3: a signal more directly linked to viral infection, such as a viral RNA sensing output
In an IANN, each of these inputs could be assigned a different weight. A viral signal could have the strongest positive weight, a general inflammatory signal could have a moderate weight, and a stress-associated signal could even be assigned a negative influence if it tends to create false positives. The output would behave like a classifier: only when the weighted sum crosses a threshold would the cell activate a fluorescent reporter or another downstream response.
This is more realistic than a strict Boolean circuit because infection-related biology is usually not binary. However, there are important limitations. Different plasmids may enter cells at different copy numbers, creating cell-to-cell variability. Different inputs may also rise and decay at different times, which can distort the intended weighted computation. Additional limitations include molecular burden, leakage in the OFF state, crosstalk between regulatory parts, and the fact that many biological neural-like systems still rely on weights that were optimized offline rather than learned directly inside the cell.
3. Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
3. Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
Below is a conceptual intracellular multilayer perceptron. In this architecture, layer 1 integrates two DNA inputs and produces an intermediate endoribonuclease output. That endoribonuclease regulates the reporter in layer 2.
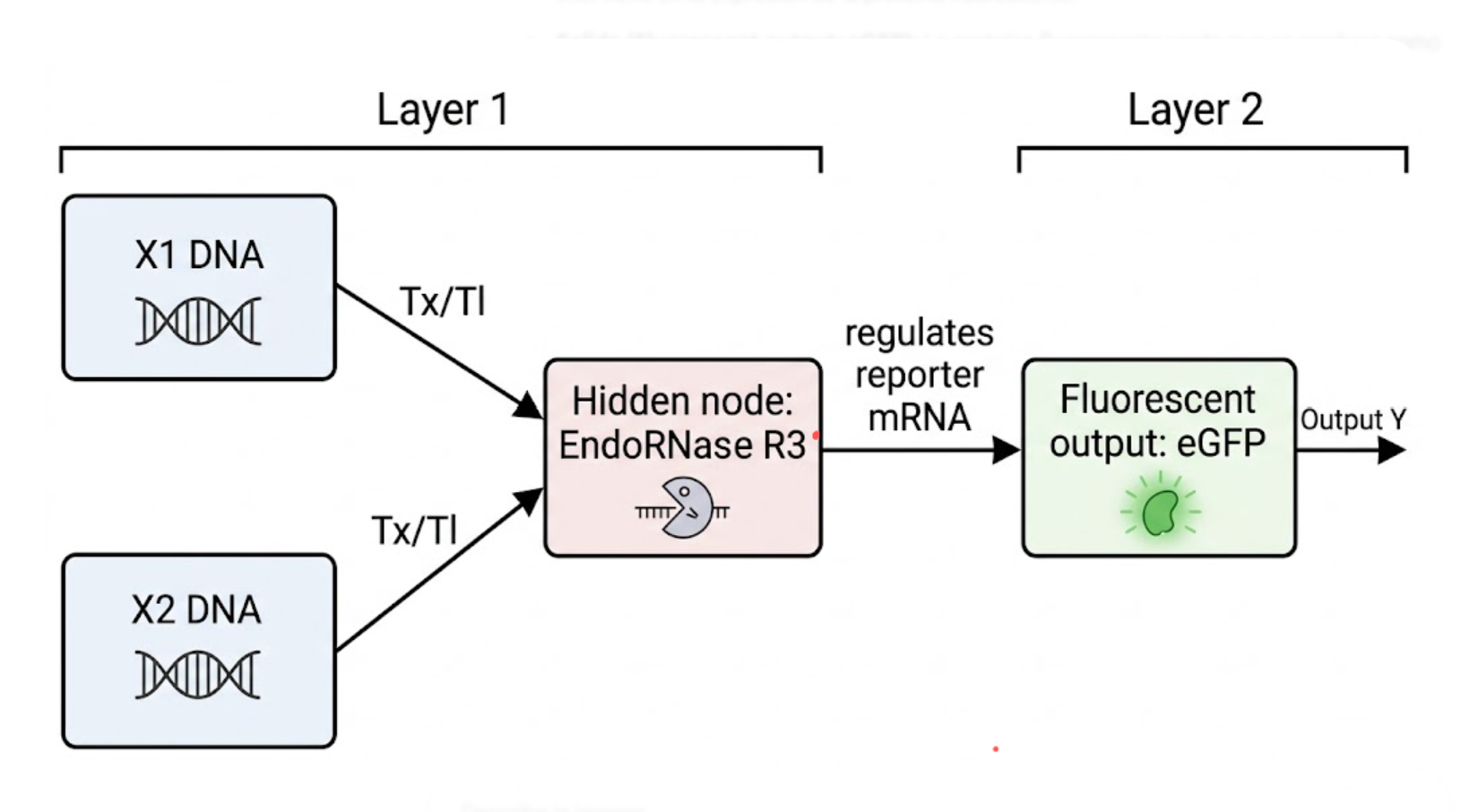

Figure 1. Conceptual intracellular multilayer perceptron in which layer 1 integrates two DNA inputs and produces an intermediate endoribonuclease that regulates fluorescent output in layer 2.
Part 2: Fungal Materials
1. What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Existing fungal materials are mainly based on mycelium, the filamentous vegetative structure of fungi. One major category is mycelium-based composites, in which fungi grow through agricultural or industrial waste and bind the substrate into a lightweight solid material. These are being explored or used for protective packaging, thermal insulation, acoustic panels, and interior design elements.
Another important category is pure mycelium materials, which are produced with less dependence on a bulky plant substrate and can be processed into leather-like sheets, foam-like materials, and paper-like materials.
Their main advantages are related to sustainability. They can be grown from agricultural residues, usually require lower energy inputs than many conventional materials, and are often biodegradable or compostable. In addition, fungal materials can show useful properties such as low density, thermal insulation, acoustic absorption, and, in some cases, favorable fire-related behavior.
Their disadvantages are also important. Many fungal materials still have lower and more variable mechanical strength than conventional plastics, foams, or structural composites. They can absorb moisture, which may weaken performance over time. Long-term durability, reproducibility, and large-scale manufacturing consistency remain major challenges. For that reason, fungal materials are currently more realistic for packaging, insulation, acoustics, and leather alternatives than for demanding structural applications.
2. What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
One application I find especially interesting would be to engineer fungi to create smart building materials that not only provide insulation or structure, but also sense environmental changes. For example, I would like to engineer a fungal material that could detect persistent moisture inside walls and respond with a visible color change or another easy-to-read signal.
This would be useful because hidden water damage is often detected too late, after microbial growth, structural problems, or health risks have already started. A fungal material that acts both as a material and as a living sensor could support more sustainable and safer buildings.
Fungi offer important advantages over bacteria for this type of application. Fungi naturally grow as extended hyphal networks, allowing them to form cohesive three-dimensional materials directly on solid substrates. Many fungi also grow on lignocellulosic or waste-derived feedstocks, which is attractive for low-cost and sustainable manufacturing. In addition, fungi are naturally well suited to material formation because their biology already supports macroscopic structure generation.
Compared with bacteria, fungi may therefore be better chassis for engineered living materials when the goal is to build a physical object rather than only produce a soluble molecule. However, fungi also have drawbacks: they often grow more slowly, can be harder to genetically manipulate than standard bacterial hosts, and may introduce variability in morphology and performance. Even so, they are especially promising for material-oriented synthetic biology.
Quick comparison
| Material type | Example applications | Main advantages | Main limitations |
|---|---|---|---|
| Mycelium-based composites | Packaging, insulation, acoustic panels | Low energy, biodegradable, waste-based feedstocks | Variable strength, moisture sensitivity |
| Pure mycelium / myco-leather | Leather alternatives, flexible sheets | Animal-free, potentially biodegradable, tunable processing | Durability and scale-up still challenging |
Part 3: First DNA Twist Order
Final project selection snapshot
For my individual final project, I selected the concept of an Automated Optimization of a DNAzyme–Cas12a Amplified Lead Sensor. The project is based on coupling a Pb²⁺-responsive DNAzyme to a CRISPR-Cas12a amplification step, so that substrate cleavage releases a trigger capable of activating Cas12a and generating a fluorescent signal.
In the short term, the project focuses on in-silico design and kinetic modeling. In the medium term, the goal is to optimize the assay experimentally using automated liquid handling. In the long term, the platform could be translated into a modular and portable environmental sensing format.
Aim 1 draft
The first aim of my final project is to computationally design and prioritize a modular DNAzyme–Cas12a lead sensor by optimizing nucleic acid architecture, assessing structural plausibility of the Cas12a activation complex, and building an ODE-based kinetic model to predict signal amplification, leakage, and theoretical sensitivity before wet-lab testing.
DNA design strategy for this assignment
For this first DNA synthesis design exercise, I chose to build a constitutive sfGFP expression cassette as a workflow control. Although my individual final project is focused on a DNAzyme–Cas12a amplified lead sensor, this Week 7 design is intended to document the full sequence design and cloning workflow in a simple and robust way.
The insert was designed as a linear expression cassette containing:
- a constitutive promoter
- an RBS
- a start codon
- the sfGFP coding sequence
- a 7xHis tag
- a stop codon
- a terminator
Insert documentation


Backbone documentation
Backbone vector: pTwist Amp High Copy
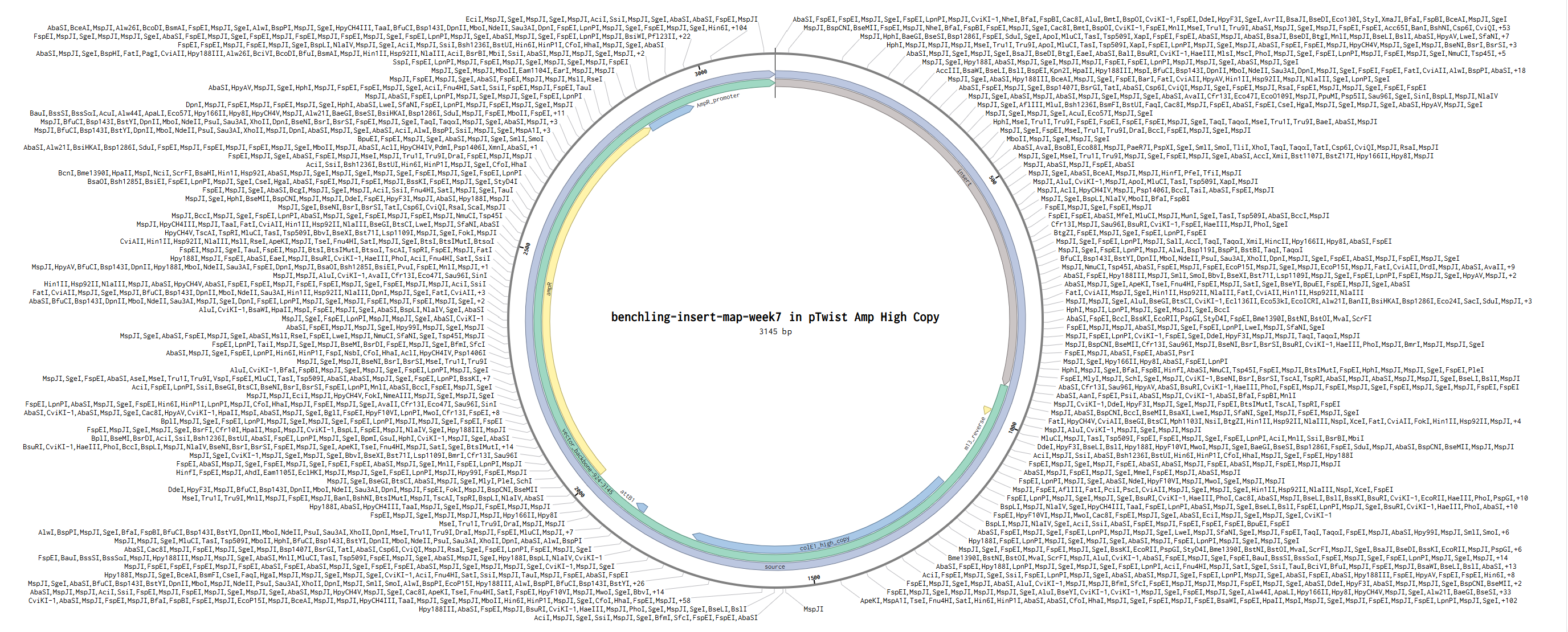

Reflection
This exercise helped me connect sequence design, annotation, synthesis planning, and plasmid-level documentation into one workflow. In future iterations, I plan to replace the generic reporter cassette with a project-relevant construct connected to my DNAzyme–Cas12a sensing platform.
References
- HTGAA 2026 Genetic Circuits II Lab Protocol.
- Vasle, A. H., & Moškon, M. (2024). Synthetic biological neural networks: From current implementations to future perspectives. BioSystems, 237, 105164.
- HTGAA Spring 2026 — Week 2: DNA Read, Write, & Edit.
- HTGAA 2026: Final Project Selection.
- HTGAA 2026: Individual Final Project Documentation.