Week 9 HW: Cell-free Systems
Week 9 — Cell-free Systems
Homework Part A: General and Lecturer-Specific Questions
General homework questions
1. Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
Cell-free protein synthesis offers major advantages over traditional in vivo expression because the reaction occurs outside living cells, in a simplified and highly controllable environment. Instead of relying on cell growth, viability, and intracellular regulation, the experimenter can directly tune DNA concentration, salts, cofactors, energy source, reaction time, temperature, and inducer concentration. This makes the system highly flexible for rapid prototyping, mechanistic studies, and controlled optimization of genetic constructs. Unlike cell-based production, cell-free systems do not require maintaining living hosts and reduce interference from the host’s own physiology and background protein production. This is one of the reasons they are widely used in synthetic biology, protein engineering, biosensing, and CRISPR-related research.
Cell-free expression is especially more beneficial than cell production in at least two important cases. First, it is very useful for rapid testing of synthetic circuits, because constructs can be evaluated without transformation, colony growth, and cellular induction. Second, it is advantageous for proteins that are toxic or difficult to express in vivo, since production is no longer tied to cell survival. A third strong case is portable biosensing, especially with freeze-dried reactions that can be rehydrated on demand in low-resource settings or even spaceflight contexts.
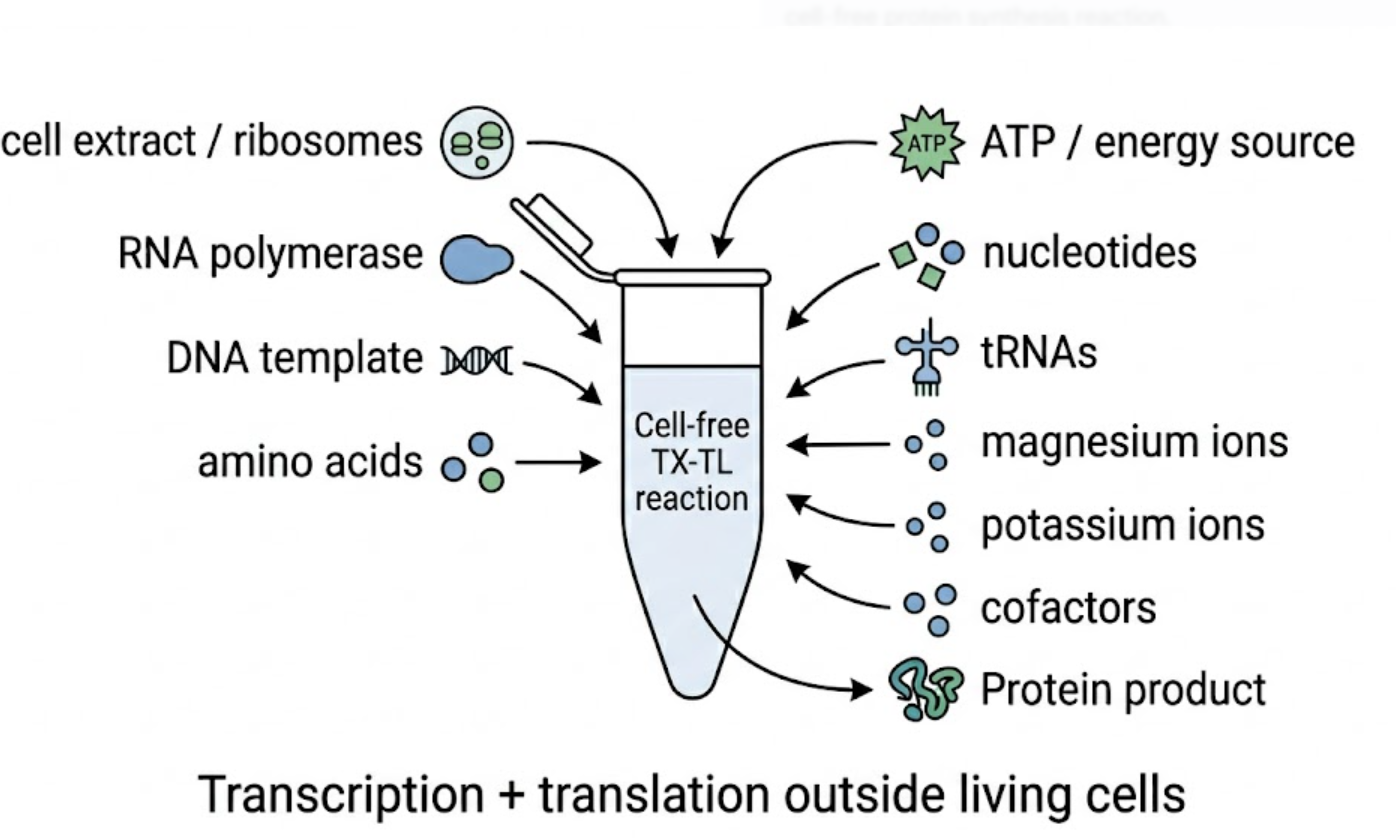

2. Describe the main components of a cell-free expression system and explain the role of each component.
A cell-free expression system contains the molecular machinery needed for transcription and translation but outside living cells. At the core of the system is either a whole-cell extract or a reconstituted PURE system. The extract or purified system provides ribosomes, translation factors, enzymes, and supporting biochemical machinery required for protein synthesis. In whole-cell extract systems, many metabolic enzymes and auxiliary cellular components are still present, while PURE systems contain only essential purified components.
The reaction also needs a buffering system, such as HEPES, to maintain stable pH and preserve enzyme activity. It requires nucleotides (ATP, GTP, CTP, UTP) for transcription and tRNAs for translation. It also needs amino acids, which are the building blocks of the protein product. Additional cofactors help maintain a productive biochemical environment. These include folinic acid, NAD, coenzyme A, spermidine, sodium oxalate, and salts such as magnesium glutamate and potassium glutamate. Magnesium is especially important because it acts as a cofactor for many enzymes involved in transcription and translation. DTT helps maintain reducing conditions and protects sensitive biomolecules.
The system also requires an energy source and a way to maintain energy availability during the reaction. Common energy substrates include 3-PGA or PEP. Finally, the system needs a template, usually DNA or RNA, that encodes the protein or biosensor of interest. In T7-based systems, T7 RNA polymerase may also be included, and RNase inhibitors can be added to protect transcripts from degradation. Together, these components support transcription, translation, RNA stability, enzymatic activity, and sustained protein production.
3. Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
Energy provision and regeneration are critical in cell-free systems because transcription and translation are highly energy-demanding processes. ATP is required directly for biosynthesis, and the reaction also depends on a stable biochemical environment to sustain RNA synthesis, protein synthesis, and associated enzymatic steps over time. Because there are no living cells continuously regenerating metabolites, the reaction can stall quickly if ATP and related energy intermediates are depleted. The lab notes explicitly include 3-PGA or PEP as energy-supporting substrates and explain that they help provide energy and intermediate metabolites for reaction stability.
One practical method to ensure continuous ATP supply is to include an energy regeneration substrate such as phosphoenolpyruvate (PEP) or 3-phosphoglycerate (3-PGA) in the reaction mixture. These compounds help sustain ATP production through the metabolic capability retained in the extract. In practice, I would test at least two energy conditions in parallel, for example PEP versus 3-PGA, and compare final yield and expression kinetics to determine which formulation better supports prolonged protein synthesis.
4. Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
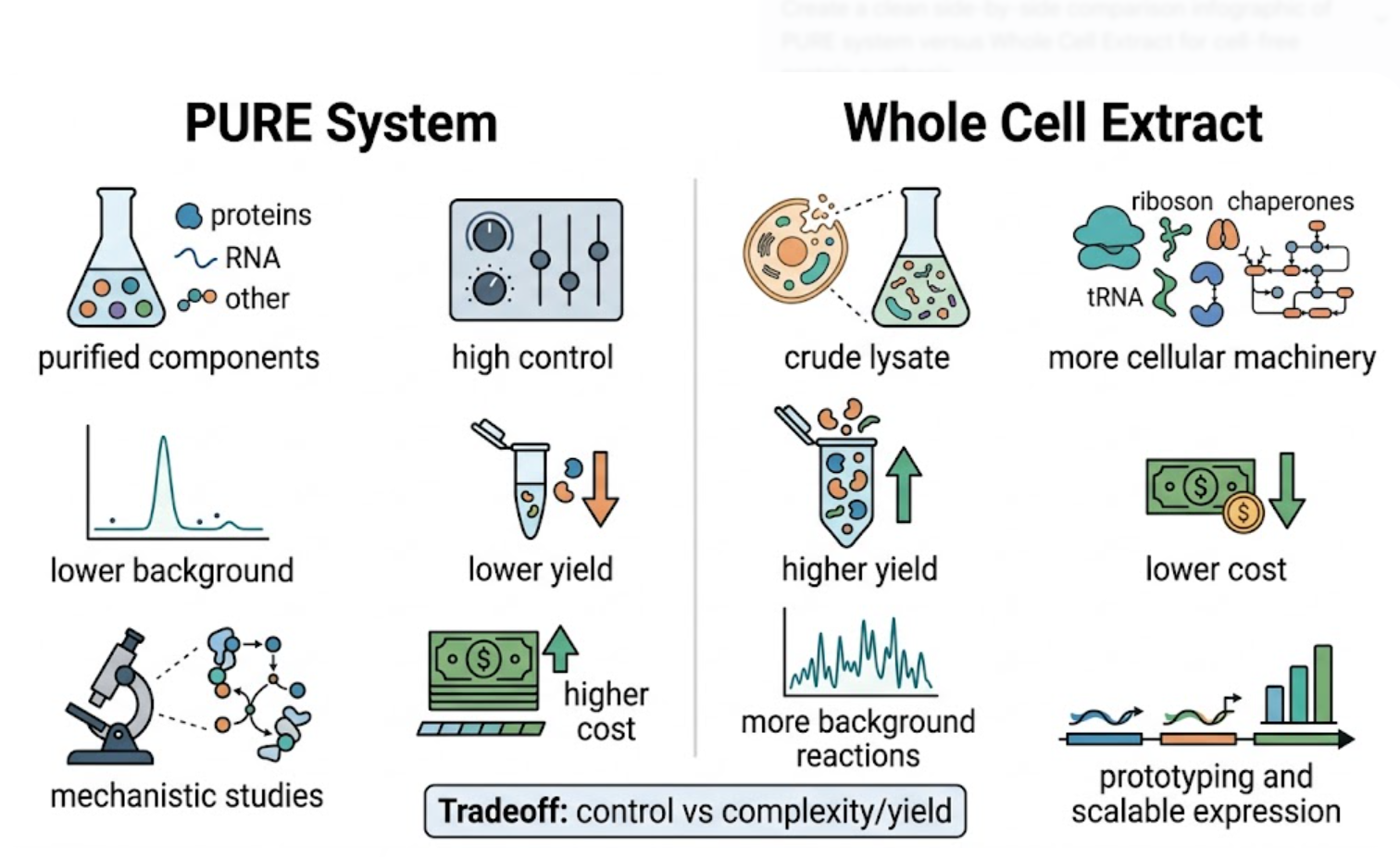
Prokaryotic and eukaryotic cell-free systems differ mainly in complexity, speed, post-translational capability, and the types of proteins they are best suited to express. Prokaryotic systems, especially E. coli-based systems, are typically fast, flexible, and relatively inexpensive. They are ideal for synthetic biology, fluorescent reporters, and proteins that do not require complex post-translational modifications. In contrast, eukaryotic systems such as wheat germ or rabbit reticulocyte extracts are better suited for proteins that require a more eukaryotic folding environment or more complex processing. The HTGAA lab notes directly compare PURE and whole-cell extract systems and note that whole-cell extracts can come from organisms including E. coli, wheat germ, and rabbit reticulocytes.
For a prokaryotic cell-free system, I would choose to produce amilGFP or deGFP, because fluorescent proteins are easy to detect, are commonly used as reporters, and generally do not require complex post-translational modifications. They are ideal for fast optimization and proof-of-concept experiments. In fact, the Week 9 lab demonstrates TX-TL functionality using a T7-IPTG-amilGFP plasmid and fluorescence monitoring across IPTG concentrations.
For a eukaryotic cell-free system, I would choose to produce an antibody fragment or a human secreted signaling protein, because these proteins are more likely to benefit from a eukaryotic translation environment, especially if proper folding, disulfide bonding, or more native-like processing is important.

5. How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
To optimize expression of a membrane protein in a cell-free system, I would design a small matrix experiment in which I systematically vary temperature, template concentration, reaction time, salt composition, and especially the presence of membrane-mimicking additives such as detergents, liposomes, or nanodiscs. I would begin with a screening-scale setup to identify conditions that maximize soluble or functional product, not just total expression. This kind of tuning is one of the major strengths of cell-free systems, since the reaction chemistry can be adjusted directly without the constraints of cell viability.
The main challenges with membrane proteins are poor solubility, aggregation, misfolding, and inefficient insertion into membrane-like environments. To address these, I would test a panel of membrane mimics in parallel and compare lower and higher expression temperatures, because slower synthesis often improves folding quality. I would also compare at least two DNA concentrations, because overexpression can worsen aggregation.
To evaluate success, I would not rely only on total protein amount. I would also use a functional readout if possible, such as ligand binding, channel activity, or detergent-stable recovery. In other words, the goal would be to optimize for correctly folded, functional protein, not just maximum yield.
6. Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
One possible reason is poor template quality or incorrect template concentration. If the DNA is degraded, impure, or present at a suboptimal concentration, transcription may be inefficient. A troubleshooting strategy would be to verify DNA quality, confirm concentration accurately, and test a small template titration series.
A second possible reason is suboptimal reaction chemistry, including energy limitation, salt imbalance, or insufficient cofactors. Cell-free systems are highly sensitive to magnesium, potassium, energy substrates, and overall reaction composition. A troubleshooting strategy would be to test several magnesium and energy-support conditions in parallel and compare both kinetics and final yield. The Week 9 lab explicitly emphasizes the importance of salts, nucleotides, cofactors, and energy substrates such as 3-PGA or PEP. :contentReference[oaicite:20]{index=20}
A third possible reason is RNA or protein instability. Transcripts may be degraded by RNases, or the protein itself may misfold, aggregate, or be unstable under the chosen conditions. A troubleshooting strategy would be to include RNase protection, reduce reaction temperature, shorten incubation time, or redesign the construct to improve translation and folding. The lab notes specifically include murine RNase inhibitor as a component used to protect mRNA from degradation. :contentReference[oaicite:21]{index=21}
Homework question from Kate Adamala
Design an example of a useful synthetic minimal cell
Pick a function and describe it.
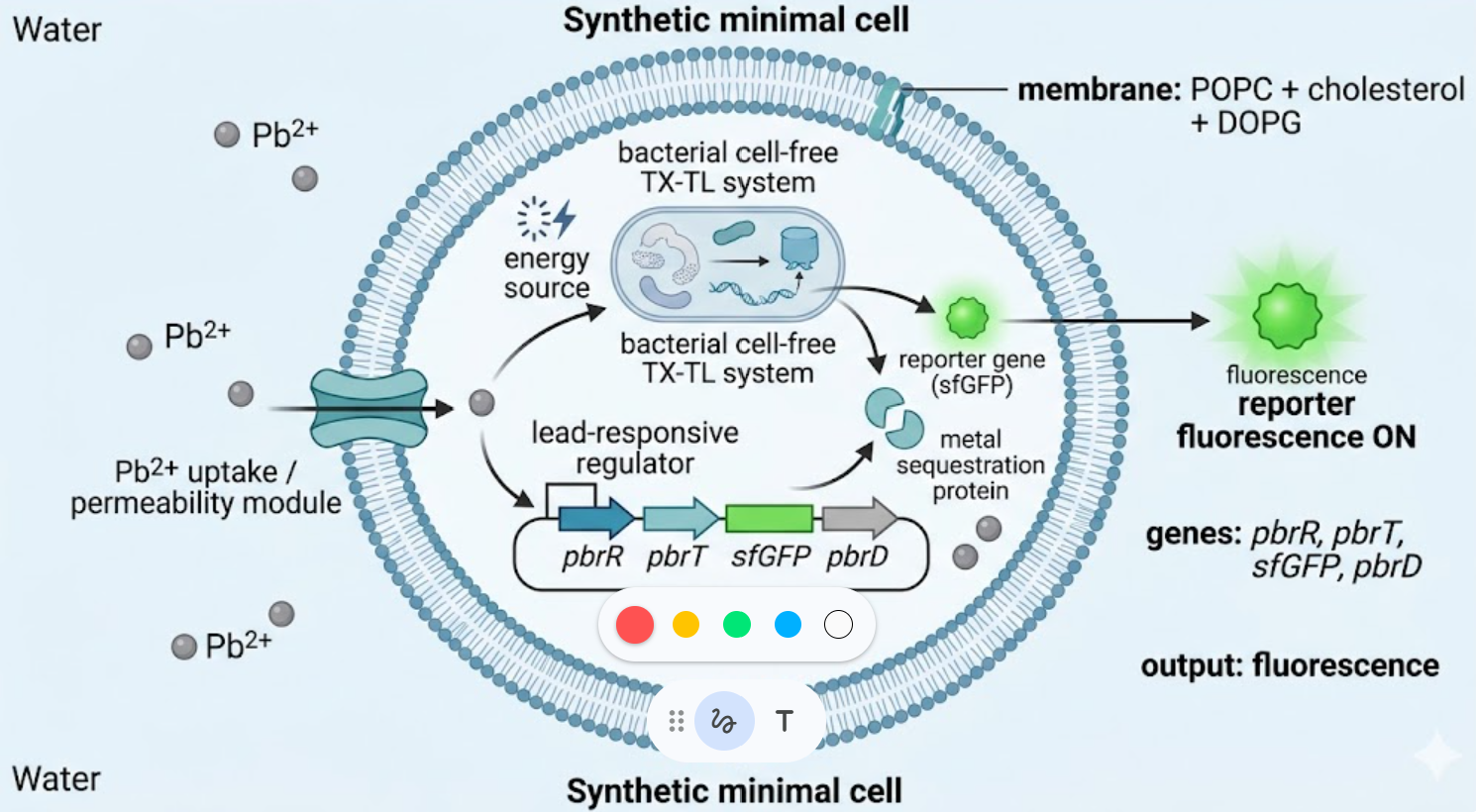
I would design a lead-sensing synthetic minimal cell for environmental monitoring and remediation.
What would your synthetic cell do? What is the input and what is the output?
The synthetic cell would detect Pb²⁺ ions in a water sample and respond by producing a fluorescent readout together with a lead-binding sequestration protein inside the compartment.
Input: Pb²⁺ in the surrounding environment.
Output: fluorescence plus intracellular lead-capture activity.
Could this function be realized by cell-free Tx/Tl alone, without encapsulation?
Only partially. A purely open cell-free reaction could detect Pb²⁺ and produce a reporter signal, but it would not behave as a discrete synthetic cell and would have limited control over selective uptake, localization, and containment of the response. Encapsulation adds compartmentalization and makes the design more realistic as a minimal cell.
Could this function be realized by genetically modified natural cell?
Yes, it could be realized in a genetically engineered bacterium. However, using a synthetic minimal cell would reduce concerns related to growth, escape, biocontainment, and environmental release of living engineered organisms.
Describe the desired outcome of your synthetic cell operation.
In the presence of lead, the synthetic minimal cell should generate a clear and measurable fluorescent signal and retain part of the toxic metal within the compartment by expressing a sequestration module.
Design all components that would need to be part of your synthetic cell.
The system would require:
- a membrane compartment
- an internal TX-TL system
- a lead-responsive sensing circuit
- a fluorescent reporter
- a sequestration module
- sufficient salts, cofactors, amino acids, nucleotides, and energy substrate
What would be the membrane made of?
A phospholipid membrane made of POPC + cholesterol, with a small fraction of negatively charged lipid such as DOPG to improve stability and tunability.
What would you encapsulate inside? Enzymes, small molecules.
Inside the vesicle I would encapsulate:
- an E. coli-based cell-free TX-TL system
- nucleotides
- amino acids
- magnesium and potassium salts
- an energy source such as PEP
- a plasmid carrying a lead-responsive regulatory system
- a fluorescent reporter gene such as sfGFP
- a lead-binding protein gene such as smtA or pbrD
Which organism would your Tx/Tl system come from? Is bacterial OK, or do you need a mammalian system for some reason?
A bacterial system is sufficient here. An E. coli-derived TX-TL system is appropriate because the sensing circuit would be based on bacterial regulatory logic, and no mammalian-specific promoter or modification system is required.
How will your synthetic cell communicate with the environment?
Lead ions are not guaranteed to cross the membrane efficiently, so I would include a metal uptake or permeability strategy, such as a membrane transporter or pore. A candidate gene would be pbrT, a lead uptake transporter. The reporter signal would be measured optically from outside the vesicle.
Experimental details
Lipids:
- POPC
- cholesterol
- DOPG
Genes:
- pbrR (lead-responsive transcriptional regulator)
- pbrT (lead uptake transporter)
- sfGFP (fluorescent reporter)
- pbrD or smtA (metal-binding/sequestration protein)
How will you measure the function of your system?
I would measure fluorescence as the primary output and compare signal across a Pb²⁺ concentration gradient. As a secondary assay, I would quantify residual lead in the external solution before and after incubation to assess whether sequestration occurred.

Homework question from Peter Nguyen
Freeze-dried cell-free systems integrated into materials
Application field
Architecture
One-sentence summary pitch
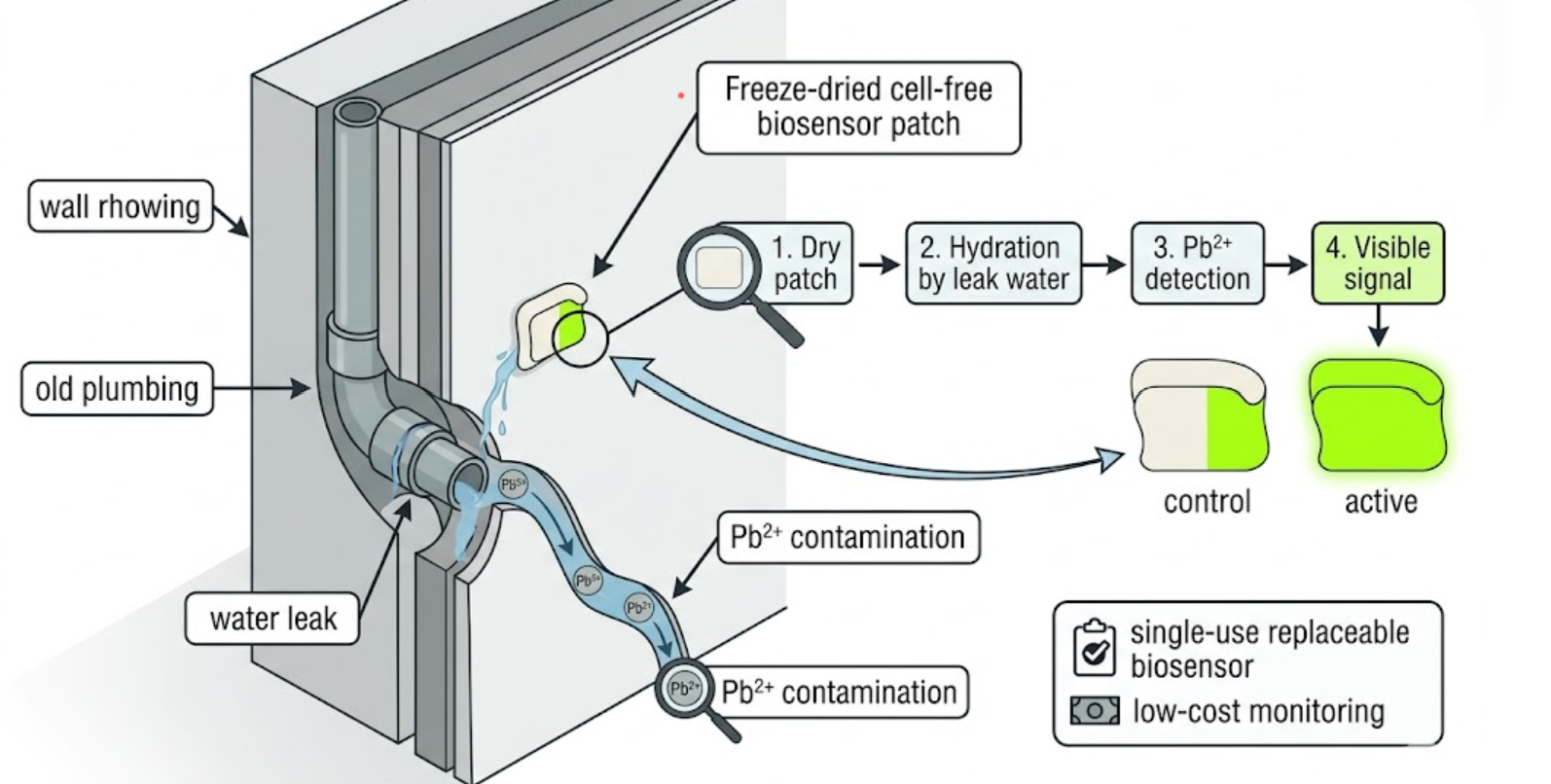
I propose a freeze-dried cell-free wall patch that becomes fluorescent when exposed to lead-contaminated water from leaking pipes.
How will the idea work, in more detail?
The concept is a replaceable patch integrated into high-risk areas of buildings, such as behind sinks, near pipe junctions, or around old plumbing. The patch would contain a freeze-dried cell-free biosensor embedded in a porous material that activates when it becomes wet. If lead-containing water reaches the patch, the biosensor would produce a visible fluorescent or colorimetric signal that indicates contamination. The patch could be read by eye or with a simple handheld fluorescence viewer. Because the reaction is freeze-dried, storage and deployment would be easy, especially in older buildings, schools, or low-resource settings.
What societal challenge or market need will this address?
This addresses the need for fast, low-cost, decentralized detection of water contamination, especially in aging infrastructure where lead exposure remains a major public health problem. It could be especially valuable in schools, public buildings, rental housing, and remote communities.
How do you envision addressing the limitation of cell-free reactions (e.g., activation with water, stability, one-time use)?
The patch would be packaged in a moisture-protective housing until installation and would be designed as a single-use replaceable sensor. Stability would be improved by lyophilization and sealed storage. Since accidental hydration is the main activation trigger, the patch would only be exposed at the desired monitoring location. One-time use is acceptable here because the material is intended as a cheap diagnostic indicator rather than a reusable electronic sensor.

Homework question from Ally Huang
Mock Genes in Space proposal
Background information (maximum 100 words)
Long-duration space missions depend on safe recycled water and fast biological monitoring, but current detection workflows can be slow, equipment-intensive, or dependent on return-to-Earth analysis. A freeze-dried cell-free biosensor could provide a lightweight, low-maintenance method for detecting microbial contamination on orbit. This is significant for astronaut health, highly relevant for future missions with limited resupply, and scientifically interesting because it combines molecular detection, low-resource biotechnology, and space-compatible synthetic biology.
Molecular or genetic target (maximum 30 words)
A bacterial 16S rRNA-derived sequence amplified from recycled spacecraft water samples.
How your target relates to the space biology question (maximum 100 words)
If bacterial nucleic acids are detected in recycled spacecraft water, that indicates possible contamination or biofilm-related risk within the life-support system. Monitoring a bacterial nucleic acid target is therefore directly relevant to astronaut health and to the reliability of long-duration water recycling infrastructure. A sequence-based target is also practical because it can be amplified and then linked to a cell-free biosensor readout.
Hypothesis or research goal (maximum 150 words)
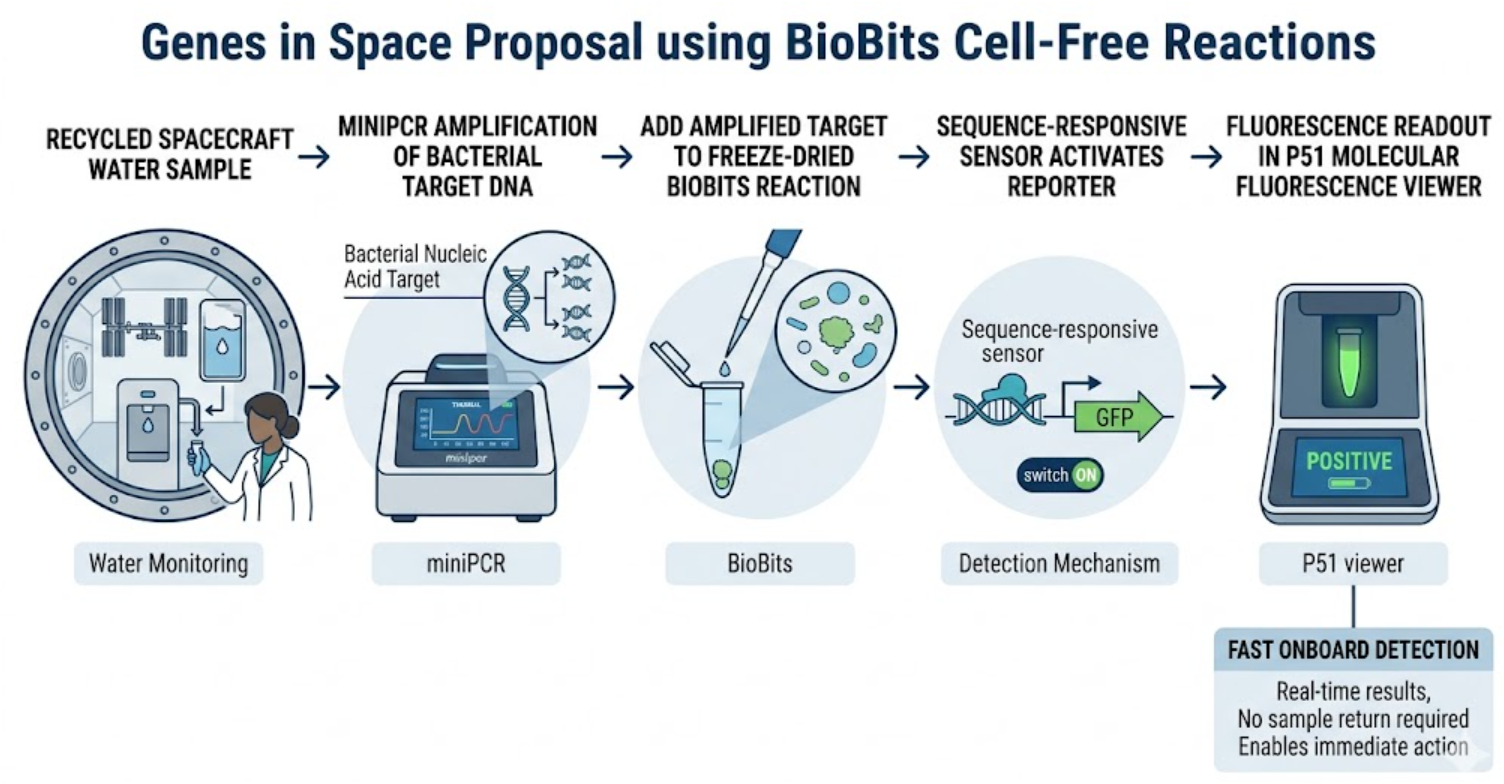
My hypothesis is that a freeze-dried BioBits® cell-free reaction coupled to a sequence-specific RNA sensing module can provide a simple and space-compatible readout for bacterial contamination in recycled water. I expect that if a bacterial target sequence is first enriched using the miniPCR® thermal cycler, then the amplified product can trigger a cell-free sensor and generate a visible fluorescence output in the P51 Molecular Fluorescence Viewer. The reasoning is that cell-free systems are lightweight, low-maintenance, and compatible with freeze-dried deployment, which makes them attractive for spaceflight where mass, storage, and user complexity are constrained.
Experimental plan (maximum 100 words)
I would test mock water samples containing either bacterial target DNA, non-target DNA, or no DNA. The target region would first be amplified using miniPCR. Amplified material would then be added to a BioBits® reaction containing a sequence-responsive sensing construct and reporter output. Controls would include a positive target control, a negative no-template control, and a non-target sequence control. The main measurements would be fluorescence intensity over time and endpoint signal discrimination between positive and negative samples.

Homework Part B: Individual Final Project
For this week, I focused on defining Aim 1 of my final project.
Final project title
Automated Optimization of a DNAzyme–CRISPR Amplified Lead Sensor
Aim 1
Design and computationally optimize a lead-responsive DNAzyme-to-Cas12a signal transduction architecture before wet-lab screening.
Aim 1 rationale
The first objective is to establish a robust in silico framework for the biosensor before experimental optimization. This includes designing the DNAzyme substrate and release trigger, tuning the coupling between DNAzyme cleavage and Cas12a activation, minimizing unintended secondary structures, and selecting reporter architectures that maximize signal gain while minimizing background. By defining these design constraints early, the wet-lab phase can focus on a smaller and more rational set of candidate constructs.
Initial experimental and design focus
Aim 1 will include:
- sequence design and secondary structure analysis
- trigger and reporter architecture comparison
- specificity considerations for Pb²⁺-dependent activation
- initial planning for automated parameter screening in later stages
Note
The slide deck submission, final project form, and ordering spreadsheet tasks will be completed through the required external course materials separately.


References
- HTGAA 2026 Cell-free Systems Lab. :contentReference[oaicite:22]{index=22}
- DNAdots: Cell-free protein synthesis. :contentReference[oaicite:23]{index=23}
- Kocalar et al., 2024. Validation of Cell-Free Protein Synthesis Aboard the International Space Station. :contentReference[oaicite:24]{index=24}
- Week 9 assignment notes. :contentReference[oaicite:25]{index=25}