Individual Final Project
title: ‘Individual Final Project’ weight: 10 description: ‘Automated Optimization of a DNAzyme–Cas12a Amplified Lead Sensor’
Automated Optimization of a DNAzyme–Cas12a Amplified Lead Sensor
Abstract
Lead contamination in drinking water remains a major public health problem because even low-level chronic exposure can impair neurological development, cardiovascular health, and overall long-term wellbeing. Existing analytical methods such as ICP-MS are highly sensitive, but they usually require centralized laboratory infrastructure, trained personnel, and expensive instrumentation, which limits their accessibility for decentralized or field-based monitoring. The overall goal of this project is to develop a modular environmental biosensing platform that couples a Pb²⁺-responsive DNAzyme with CRISPR-Cas12a signal amplification in order to generate a rapid and amplified fluorescent readout. The central hypothesis is that a DNAzyme-triggered release of a programmable nucleic acid activator can be linked to Cas12a collateral cleavage to improve sensitivity while preserving modularity. To test this idea, the project is structured into three aims: first, computational design and kinetic modeling of the sensing cascade; second, automated experimental optimization using robotic liquid handling; and third, long-term translation into a portable and modular environmental sensing format. The methods include nucleic acid folding analysis, structural plausibility assessment, kinetic simulation, DNA construct design, and future automated wet-lab optimization. Together, this project aims to establish a scalable biosensing framework for environmental monitoring that is adaptable, programmable, and ultimately deployable outside centralized laboratories.
Project Aims
Aim 1: Experimental / Short-term Aim
The first aim of my final project is to computationally design and prioritize a modular DNAzyme–Cas12a lead sensor by optimizing nucleic acid architecture, assessing structural plausibility of the Cas12a activation complex, and building an ODE-based kinetic model to predict signal amplification, leakage, and theoretical sensitivity before wet-lab testing.
Aim 2: Development / Medium-term Aim
The second aim of my final project is to experimentally optimize and validate the sensor using automated liquid handling workflows. Following successful in-silico prioritization, this stage will use an Opentrons OT-2 platform to execute multidimensional parameter sweeps across reaction variables such as pH, Mg²⁺ concentration, reporter concentration, and DNAzyme/Cas12a stoichiometry in order to identify conditions that maximize sensitivity and reproducibility in real water samples.
Aim 3: Visionary / Long-term Aim
The third aim of my final project is to develop the sensing platform into a modular and field-deployable environmental monitoring technology. In the long term, the assay could be adapted into decentralized formats such as lyophilized or paper-based systems and extended to detect additional toxic metals by replacing the upstream recognition module while preserving the downstream CRISPR-based amplification architecture
Recent literature supports the use of DNAzymes as mechanistically well-defined and highly selective recognition elements for lead sensing. Brown et al. described a lead-dependent 8-17 DNAzyme with a two-step catalytic mechanism, providing an important biochemical basis for Pb²⁺-responsive cleavage. Structural work later showed that RNA-cleaving DNAzymes such as 8-17 adopt compact active conformations with a pre-organized Pb²⁺ binding pocket, strengthening the connection between sequence, folding, and catalysis.
Beyond mechanistic studies, DNAzymes have also been successfully translated into sensing platforms. Li et al. reported a single-stranded fluorescent Pb²⁺ DNAzyme sensor with good performance across a broad temperature range, highlighting the practicality of DNAzyme-based environmental sensing. More recently, He et al. developed a DNAzyme-based CRISPR/Cas12a fluorescence sensor for Pb²⁺ detection, directly demonstrating the feasibility of coupling metal-responsive DNAzyme cleavage to CRISPR-mediated signal amplification. Together, these studies support the conceptual basis of my project while also revealing opportunities for improved modularity, automation, and optimization.
- Brown, A. K., Li, J., Pavot, C. M.-B., & Lu, Y. (2003). A lead-dependent DNAzyme with a two-step mechanism. Biochemistry, 42(23), 7152–7161.
- Liu, H. et al. (2017). Crystal structure of an RNA-cleaving DNAzyme. Nature Communications.
- Li, H. et al. (2012). Single-stranded DNAzyme-based Pb²⁺ fluorescent sensor that can work well over a wide temperature range. Biosensors and Bioelectronics, 34(1), 159–164.
- He, S. et al. (2025). A DNA concatemer-encoded CRISPR/Cas12a fluorescence sensor for sensitive detection of Pb²⁺ based on DNAzymes. Analyst, 150(9), 1778–1784.
Ethical Implications
This project raises several ethical considerations related to environmental health, public communication, and responsible biosensor development. At its core, the project is motivated by beneficence, because it aims to improve access to lead monitoring tools that could support earlier detection of unsafe water conditions and reduce long-term exposure to a major public health hazard. It also relates to justice, since communities with fewer resources are often the ones most affected by environmental contamination while also having the least access to centralized analytical testing. At the same time, the principle of non-maleficence is especially important here, because an inaccurate sensor could produce false negatives that give users unjustified confidence in contaminated water, or false positives that generate unnecessary alarm. Since the project is based on a modular synthetic biology sensing architecture, it must also be guided by responsibility in how claims are made, how performance is validated, and how limitations are communicated.
To ensure that this project is ethical, several measures should be taken both during research and in any future real-world deployment. First, the sensor should never be presented as a replacement for certified analytical methods unless its performance has been rigorously benchmarked under realistic environmental conditions. Second, all results should be reported transparently, including background leakage, false activation risks, matrix effects, and any uncertainty in the predicted or measured limit of detection. Third, the project should include safe handling and disposal practices for all reagents, especially if future versions use CRISPR components, fluorogenic reporters, lyophilized reaction mixes, or field-deployable formats. A further ethical requirement is to avoid overpromising accessibility if the assay still depends on conditions or materials that are difficult to standardize outside the laboratory. Alternatives such as using the platform only for preliminary screening, while confirming results with certified methods, should remain part of the deployment strategy. In this way, the project can remain aligned with public health goals while minimizing the risk of misuse, misinterpretation, or premature application.
Ethical Implications
This project raises several ethical considerations related to environmental health, public communication, and responsible biosensor development. At its core, the project is motivated by beneficence, because it aims to improve access to lead monitoring tools that could support earlier detection of unsafe water conditions and reduce long-term exposure to a major public health hazard. It also relates to justice, since communities with fewer resources are often the ones most affected by environmental contamination while also having the least access to centralized analytical testing. At the same time, the principle of non-maleficence is especially important here, because an inaccurate sensor could produce false negatives that give users unjustified confidence in contaminated water, or false positives that generate unnecessary alarm. Since the project is based on a modular synthetic biology sensing architecture, it must also be guided by responsibility in how claims are made, how performance is validated, and how limitations are communicated.
To ensure that this project is ethical, several measures should be taken both during research and in any future real-world deployment. First, the sensor should never be presented as a replacement for certified analytical methods unless its performance has been rigorously benchmarked under realistic environmental conditions. Second, all results should be reported transparently, including background leakage, false activation risks, matrix effects, and any uncertainty in the predicted or measured limit of detection. Third, the project should include safe handling and disposal practices for all reagents, especially if future versions use CRISPR components, fluorogenic reporters, lyophilized reaction mixes, or field-deployable formats. A further ethical requirement is to avoid overpromising accessibility if the assay still depends on conditions or materials that are difficult to standardize outside the laboratory. Alternatives such as using the platform only for preliminary screening, while confirming results with certified methods, should remain part of the deployment strategy. In this way, the project can remain aligned with public health goals while minimizing the risk of misuse, misinterpretation, or premature application.
Ethical Implications
This project raises several ethical considerations related to environmental health, public communication, and responsible biosensor development. At its core, the project is motivated by beneficence, because it aims to improve access to lead monitoring tools that could support earlier detection of unsafe water conditions and reduce long-term exposure to a major public health hazard. It also relates to justice, since communities with fewer resources are often the ones most affected by environmental contamination while also having the least access to centralized analytical testing. At the same time, the principle of non-maleficence is especially important here, because an inaccurate sensor could produce false negatives that give users unjustified confidence in contaminated water, or false positives that generate unnecessary alarm. Since the project is based on a modular synthetic biology sensing architecture, it must also be guided by responsibility in how claims are made, how performance is validated, and how limitations are communicated.
To ensure that this project is ethical, several measures should be taken both during research and in any future real-world deployment. First, the sensor should never be presented as a replacement for certified analytical methods unless its performance has been rigorously benchmarked under realistic environmental conditions. Second, all results should be reported transparently, including background leakage, false activation risks, matrix effects, and any uncertainty in the predicted or measured limit of detection. Third, the project should include safe handling and disposal practices for all reagents, especially if future versions use CRISPR components, fluorogenic reporters, lyophilized reaction mixes, or field-deployable formats. A further ethical requirement is to avoid overpromising accessibility if the assay still depends on conditions or materials that are difficult to standardize outside the laboratory. Alternatives such as using the platform only for preliminary screening, while confirming results with certified methods, should remain part of the deployment strategy. In this way, the project can remain aligned with public health goals while minimizing the risk of misuse, misinterpretation, or premature application.
Ethical Implications
This project raises several ethical considerations related to environmental health, public communication, and responsible biosensor development. At its core, the project is motivated by beneficence, because it aims to improve access to lead monitoring tools that could support earlier detection of unsafe water conditions and reduce long-term exposure to a major public health hazard. It also relates to justice, since communities with fewer resources are often the ones most affected by environmental contamination while also having the least access to centralized analytical testing. At the same time, the principle of non-maleficence is especially important here, because an inaccurate sensor could produce false negatives that give users unjustified confidence in contaminated water, or false positives that generate unnecessary alarm. Since the project is based on a modular synthetic biology sensing architecture, it must also be guided by responsibility in how claims are made, how performance is validated, and how limitations are communicated.
To ensure that this project is ethical, several measures should be taken both during research and in any future real-world deployment. First, the sensor should never be presented as a replacement for certified analytical methods unless its performance has been rigorously benchmarked under realistic environmental conditions. Second, all results should be reported transparently, including background leakage, false activation risks, matrix effects, and any uncertainty in the predicted or measured limit of detection. Third, the project should include safe handling and disposal practices for all reagents, especially if future versions use CRISPR components, fluorogenic reporters, lyophilized reaction mixes, or field-deployable formats. A further ethical requirement is to avoid overpromising accessibility if the assay still depends on conditions or materials that are difficult to standardize outside the laboratory. Alternatives such as using the platform only for preliminary screening, while confirming results with certified methods, should remain part of the deployment strategy. In this way, the project can remain aligned with public health goals while minimizing the risk of misuse, misinterpretation, or premature application.

Results & Quantitative Expectations
What aspect of the project did I choose to validate?
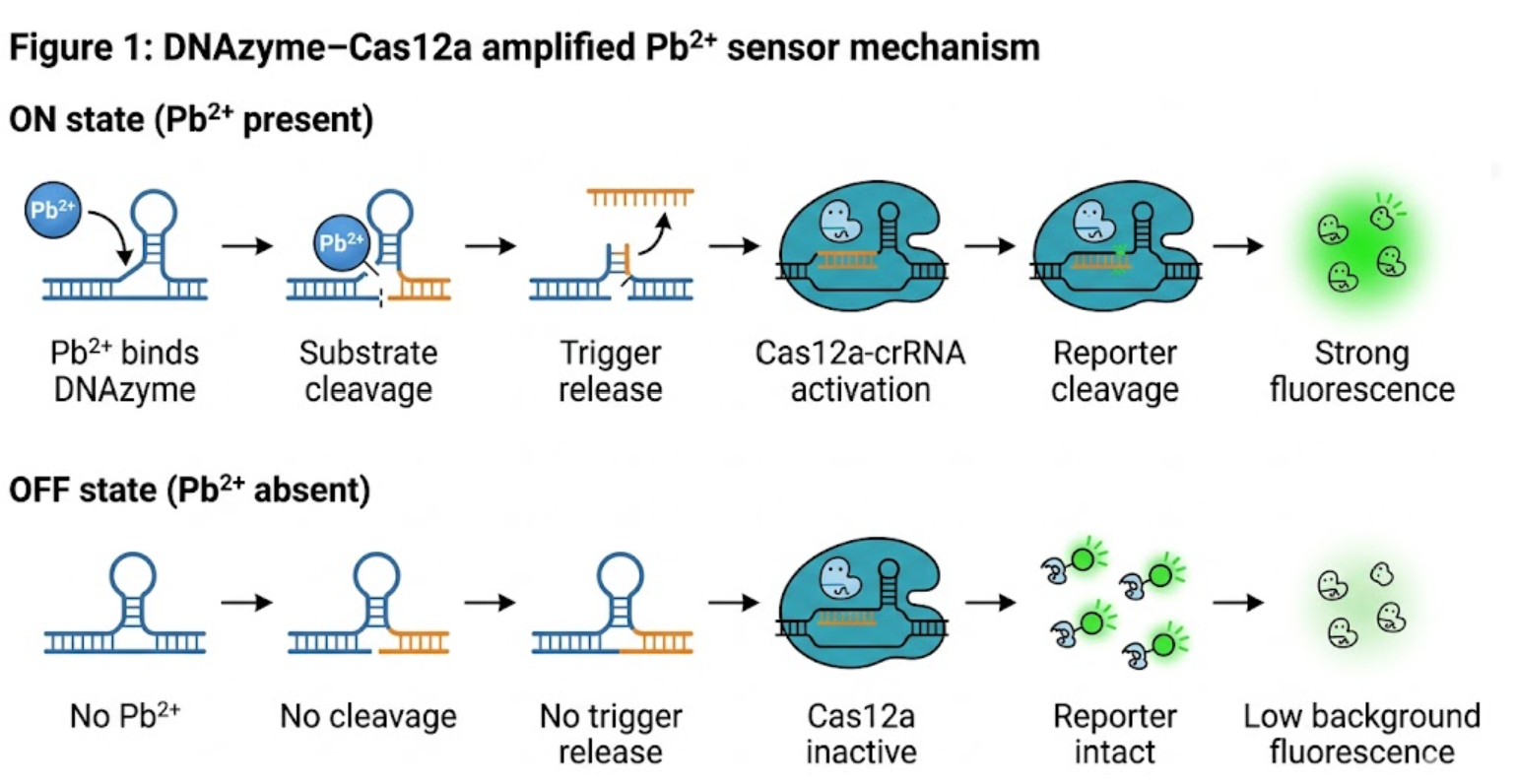
For this stage of the project, I chose to validate the design and computational prioritization workflow of the DNAzyme–Cas12a sensing cascade rather than a fully assembled wet-lab assay. This validation focuses on whether the sensing architecture can be rationally designed in a way that minimizes unwanted folding, preserves trigger accessibility, and supports a plausible downstream Cas12a activation logic. I selected this aspect because it is directly achievable within the current scope of the course and because a poor sequence architecture would undermine all later experimental optimization.
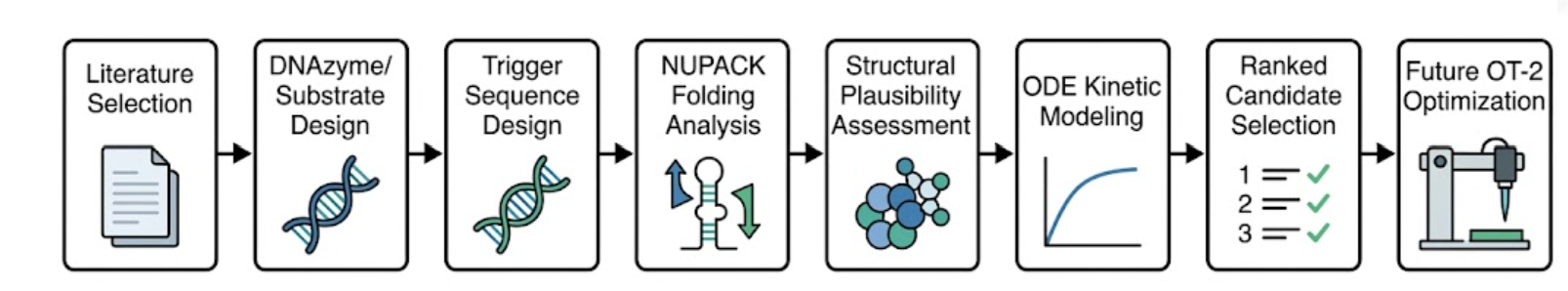
Validation protocol
- I defined the overall sensing architecture as a modular cascade composed of a Pb²⁺-responsive DNAzyme, a cleavable substrate, a released trigger strand, a Cas12a-crRNA activation module, and a fluorescent reporter output.
- I selected literature-supported DNAzyme designs relevant to Pb²⁺ sensing and used them as the mechanistic basis for the upstream recognition module.
- I drafted candidate trigger-release strategies in which cleavage of the substrate would expose or release a DNA sequence capable of activating the downstream CRISPR module.
- I annotated project-relevant sequence elements and organized the design logic in Benchling.
- I evaluated sequence-level folding behavior using NUPACK to identify unwanted secondary structures that could interfere with cleavage, trigger release, or Cas12a activation.
- I compared candidate designs by qualitatively prioritizing those with better trigger accessibility and lower predicted risk of OFF-state leakage.
- I translated the sensing cascade into a reaction-level kinetic framework suitable for ODE-based simulation.
- I defined the major kinetic steps as DNAzyme cleavage, trigger release, Cas12a activation, reporter cleavage, and background leakage.
- I used the model structure to define which variables would most strongly affect sensitivity, including cleavage efficiency, trigger concentration, activation kinetics, reporter concentration, and background activity.
- I documented a DNA design workflow compatible with future synthesis and screening steps, including Benchling annotation and plasmid-level documentation.
What synthetic biology techniques did I use in this validation?
This validation used several synthetic biology techniques even though it did not yet include a full wet-lab assay. The first was DNA construct design, because the project depends on clearly defined sequence modules and activation logic rather than on a vague conceptual pathway. The second was computational nucleic acid analysis, especially folding-based evaluation, because secondary structure directly affects accessibility and leakage in sequence-programmed sensing systems. The third was model-based analysis, since the reaction cascade must be understood not only structurally but also dynamically. Finally, the validation included Benchling-based design documentation and a Twist-compatible DNA workflow, which are essential for translating the concept into experimentally testable constructs later in the project.
What data will I present?
The main data for this stage of the project will be computational and design-derived data rather than experimental fluorescence measurements. These data may include NUPACK structure predictions, ranked candidate architectures, annotated sequence maps, and simulated kinetic trajectories from the ODE model. Together, these outputs will serve as an evidence-based justification for selecting one or more sensing architectures for future experimental optimization.
Quantitative expectations
At this stage, my quantitative expectations are focused on relative performance trends rather than final environmental performance claims. I expect that useful candidate designs will show lower predicted OFF-state leakage, improved trigger accessibility, and stronger separation between simulated ON and OFF trajectories compared with less optimized alternatives. In the next phase of the project, these computational outputs would be used to prioritize experimental conditions for automated screening and to narrow the design space before wet-lab validation.
Challenges, limitations, and alternative strategies
A major limitation of the current stage is that computational prioritization cannot prove that the full sensing cascade will behave as expected in real reaction conditions. Nucleic acid folding predictions and structural plausibility assessments are helpful, but they do not fully capture the complexity of reaction kinetics, matrix effects, incomplete cleavage, or unintended interactions between components. Another challenge is that the released trigger may be theoretically accessible in silico while still performing poorly in practice due to concentration effects, sequence context, or competing structures.
A second limitation is that the project currently depends on simplified assumptions about Cas12a activation and background behavior. These assumptions are useful for building an initial model, but they may underestimate leakage or overestimate amplification efficiency. To address this, future versions of the project should compare multiple trigger architectures and explicitly include background-cleavage scenarios in the modeling framework.
An additional challenge is that real environmental water samples may contain salts, competing ions, inhibitors, or contaminants that reduce the performance of both the DNAzyme and the CRISPR module. A promising alternative strategy would be to first optimize the system in buffered model solutions and only then move into increasingly complex matrices. Another useful alternative would be to compare several Pb²⁺-responsive DNAzyme configurations rather than relying on a single canonical design from the beginning.

References
- Brown, A. K., Li, J., Pavot, C. M.-B., & Lu, Y. (2003). A lead-dependent DNAzyme with a two-step mechanism. Biochemistry, 42(23), 7152–7161.
- Liu, H., Yu, X., Chen, Y., et al. (2017). Crystal structure of an RNA-cleaving DNAzyme. Nature Communications, 8, 2006.
- Li, H., Zhang, Q., Cai, Y., Kong, D.-M., & Shen, H.-X. (2012). Single-stranded DNAzyme-based Pb²⁺ fluorescent sensor that can work well over a wide temperature range. Biosensors and Bioelectronics, 34(1), 159–164.
- He, S., Lin, W., Liu, X., et al. (2025). A DNA concatemer-encoded CRISPR/Cas12a fluorescence sensor for sensitive detection of Pb²⁺ based on DNAzymes. Analyst, 150(9), 1778–1784.
- HTGAA 2026 Genetic Circuits II Lab Protocol.
- HTGAA Spring 2026 — Week 2: DNA Read, Write, & Edit.
- HTGAA 2026: Final Project Selection.
- HTGAA 2026: Individual Final Project Documentation.
Supply List and Budget
Core reagents and supplies
- Pb²⁺-responsive DNAzyme oligonucleotides
- Cleavable substrate oligonucleotides
- Trigger strand oligonucleotides
- crRNA for Cas12a activation
- Cas12a enzyme
- Fluorogenic ssDNA reporter
- Reaction buffers
- MgCl₂ and other salts for optimization
- Nuclease-free water
- Microcentrifuge tubes
- PCR tubes or 96-well plates
- Plate seals
- Pipette tips
- Benchling/Twist-compatible DNA design materials
- Optional lyophilization consumables for future deployment studies
Equipment
- Micropipettes
- Mini centrifuge
- Fluorescence plate reader or qPCR-style fluorescence instrument
- Thermal block or incubator
- Computer for design, simulation, and sequence analysis
- Optional Opentrons OT-2 liquid handler for automated optimization
Estimated budget categories
- Oligonucleotides: medium
- Cas12a enzyme and reporter reagents: medium to high
- Buffers and consumables: low to medium
- Plate-based fluorescence readout: depends on local instrumentation access
- Automation cost: low if institutional OT-2 access is available, high if new acquisition is required
Practical note
The most cost-sensitive components of this project are likely to be the CRISPR reagents, custom oligonucleotide sets, and any repeated optimization screens. Costs can be reduced by beginning with a computationally prioritized shortlist of designs before expanding into multidimensional wet-lab screening.