Week 1 HW.2: Lecture prep for W2
Answer prep questions from three faculty members:
Homework Questions from Professor Jacobson:
Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
Error rate refers to errors per nucleotide added per replication. An error could be a misincorporation (wrong base expressed for a pair), for example.
Error rate of polymerase synthesis is 1/1e7 (1:10^7).
Human genome has 3.1-3.2 Gbp or 3e9 base pairs.
The rate of errors in polymerase copying the human genome’s DNA is 1/1e7 * 3e9, which is nonzero.
Biology deals with the likely error through multiple levels of mitigation:
- Proofreading during synthesis corrects errors
- Mismatch repair after synthesis repairs errors
- Redundancy and selection at multiple levels - DNA is double-stranded, cells exist in huge populations, misfolded proteins get degraded, defective RNAs are destroyed, faulty cells undergo apoptosis
- Damage repair system
How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
Our assumptions:
- Average Human Protein: 1036 bp.
- ~30,000 proteins observed in mammalian genome.
- A protein of length L = 3L nucleotides (bases) + a stop codon in the genome
Coding is the process by which DNA is transcribed into mRNA (triplets / codons), and mRNA (codons) is translated into a linear chain of amino acids (polypeptides), which folds into 3D protein structures.
How many different ways are there to code for an average human protein, meaning how many different DNA encodings would compile (transcribe and translate) down to the same protein (chain of amino acids) of length 1036 bp?
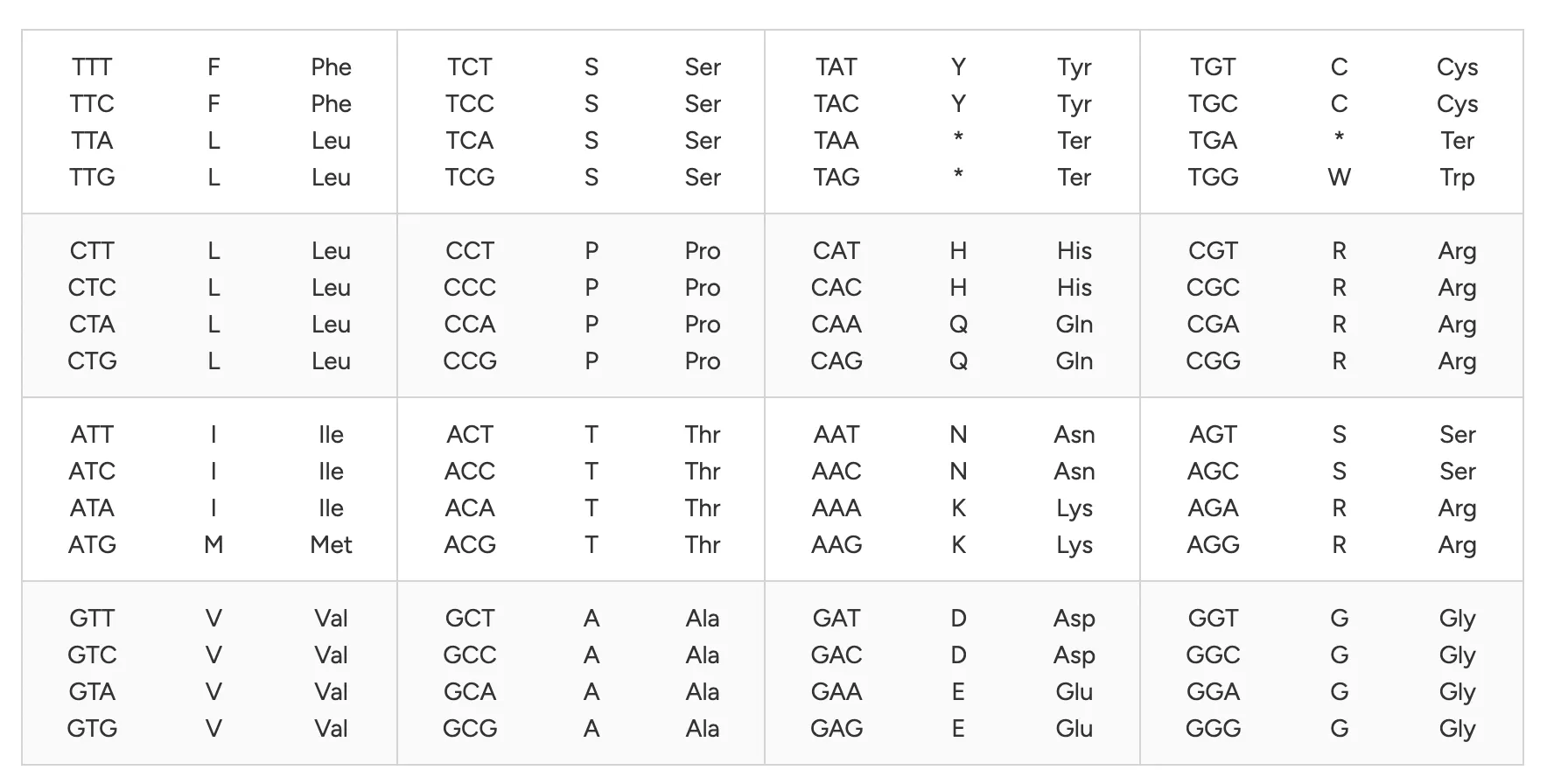

Codons are 3 nucleotides, each which have a base (A,C,G,T). There are 64 possible triplet combinations (codons) using the four bases (A, U, G, C). Each codon encodes one amino acid. An amino acid can be encoded by multiple codons. For instance, codons GAA and GAG both specify glutamic acid and exhibit redundancy. This is referred to as degeneracy.
The degeneracy of an amino acid refers to the number of codons which encode it. ie. d(Leu)=6, meaning Leucine has 6 codons which encode it.
Average codon degeneracy across amino acids is roughly 3.
So to calculate the number of possible encodings for a protein of length L=5 amino acids, we compute the degeneracy of each amino acid, and compute their product to find the maximum number of permutations. ie. for a protein of L=5, average degeneracy d(*)=3, num_permutations=d(*) * d(*) * d(*) * d(*) * d(*) = d(*)^L = 3^L
So for an average human protein of L=1036 bp, the number of possible encodings could be 3^L = 3^1036.
There is an intractable number of possible encodings. However, functional “good” encodings are a tiny subset constrained by expression, folding, RNA processing, regulation, and host biology.
Homework Questions from Dr. LeProust:
What’s the most commonly used method for oligo synthesis currently?
solid-phase chemical synthesis with phosphoramidite chemistry
Why is it difficult to make oligos longer than 200nt via direct synthesis?
Because direct phosphoramidite synthesis has a per-step yield <1.0, errors compound exponentially with length. P(success)=(1-e)^200 is improbable (e ~= 0.01)
Why can’t you make a 2000bp gene via direct oligo synthesis?
(1-e)^2000 is near impossible, due to errors accumulating from each synthetic cycle/step.
- expected number of cleavage events scales ~linearly with cycle count and purine content
- Misincorporations accumulate (wrong base addition)
Homework Question from George Church:
Choose ONE of the following three questions to answer; and please cite AI prompts or paper citations used, if any.
[Using Google & Prof. Church’s slide #4] What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
[Given slides #2 & 4 (AA:NA and NA:NA codes)] What code would you suggest for AA:AA interactions?
[(Advanced students)] Given the one paragraph abstracts for these real 2026 grant programs sketch a response to one of them or devise one of your own:
https://arpa-h.gov/explore-funding/programs/boss
What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
Histidine, isoleucine, leucine, lysine, methionine, phenylalanine, threonine, tryptophan, valine (+ arginine conditional).
Out of the 20 amino acids needed, the body synthesizes 11-12, while the remaining 8-9, known as essential amino acids, must be obtained through diet.
This is not accurate to all animals, it seems? Counterexample: cats. Cats require taurine.
The Lysine Contingency was a genetic alteration Henry Wu performed in the dinosaur genome. The modification knocked out the ability of the dinosaurs to produce the amino acid Lysine.
This forced the dinosaurs to depend on lysine supplements provided by the park’s veterinary staff. In this way, dinosaurs could never escape from the park because they would never survive long without the food supplements.
Haha, I have to rewatch this film.
The way I would hack around this would be to introduce a substance containing the microbes that cows digest and feed it to the dinosaurs. These microbes synthesise the essential amino acids from nitrogen, thus mitigating the need for the dinosaurs to produce Lysine themselves, instead forming a symbiotic relationship with the microbes in their gut.
I don’t know what this question means, but it reminds me also of Liebig’s law - would the restriction of one amino acid necessarily debilitate the dinosaurs so they can’t escape, or is nature more nonlinear and complex than that?
LLM prompts used:
- 10 essential amino acids in all animals?
- across all animals?
- cows can synthesise most of their needed amino acids? how many which ones
- how long can you survive without just one of the amnio acids ?