Week 2 HW: DNA Read, Write, and Edit
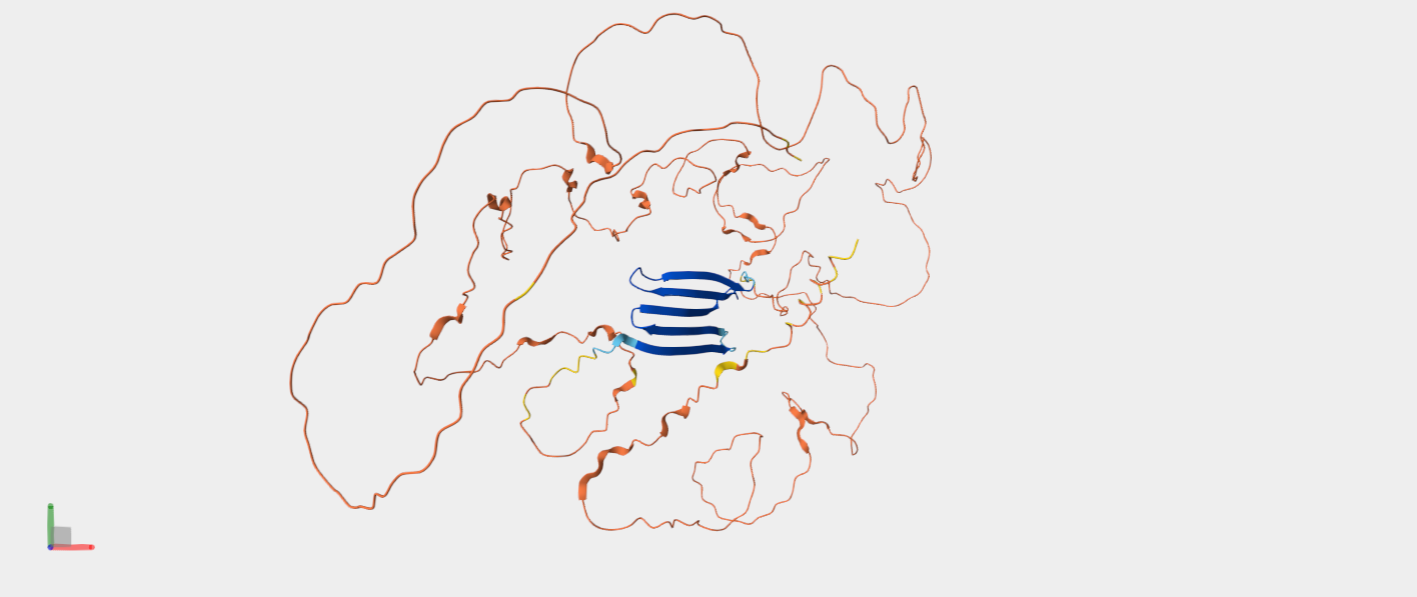

Predicted structure of Pro-resilin (Drosophila melanogaster), generated by AlphaFold and retrieved from UniProt (UniProtKB: Q9V7U0).
*You can find parts 1 and 2 in the week 2 lab section!
Part 3: DNA Design Challenge
My protein choice: Pro-resilin!
Pro-resilin is a highly elastic, rubber-like structural protein found in insects that enables efficient energy storage for jumping, flight, and sound production. About seven years ago, my biotechnology teacher in high school showed us an article about a very cool leading professor in Israel, who was doing things that seemed almost impossible. I vividly remember one of the projects he was working on: scientists had identified a protein that enables a particular spider species to jump remarkably high relative to its body size, and this scientist wanted to give humans the same ability by creating shoes with soles made from some protein similar to Pro-resilin.
That moment has stayed with me until today. I think it would be very cool to look at the structure of this protein, try to understand what gives it its unique characteristics, and potentially think about what could be done with it.
I went to UniProt.org and searched for “Pro-resilin protein Drosophila melanogaster” Here is the AA sequence:
*I chose the Drosophila melanogaster (fruit fly) resilin protein because its resilin is the best-characterized and most widely studied form of the protein.
Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
To determine the nucleotide sequence corresponding to Pro-resilin in Drosophila melanogaster, I followed the link for the canonical transcript (Q9V7U0-1) on UniProt, which directed me to Ensembl Metazoa. There, I downloaded only the coding sequence (CDS) in FASTA format, excluding UTRs and introns, to obtain the exact DNA sequence that encodes the protein. The sequence is shown below:
Codon optimization. Since the Twist Codon Optimization Tool is out of service on the website.

So I went to Google and found the free tool, the Gensmart Codon Optimization Tool. I input my CDS and chose optimization for E.coli:

And after some time, here is what I got:
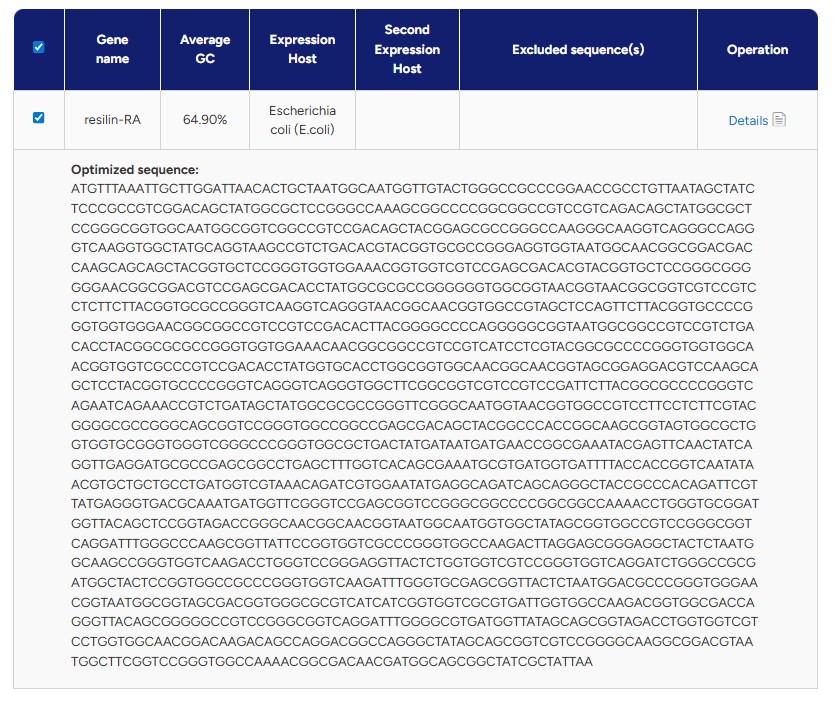

Or in plain text:
There are several reasons why we need to optimize codon usage. Although multiple codons can encode the same amino acid, different organisms have preferences for certain codons over others - a phenomenon known as codon bias. This means that a DNA sequence that works efficiently in one organism may not be translated as efficiently in another. If rare or non-preferred codons are used, the ribosome may stall, leading to lower protein expression or incomplete translation. Therefore, it is important to adapt the codons in the sequence to match the preferred codon usage of the organism in which the protein will be expressed.
I chose to optimize the sequence for Escherichia coli because it is one of the most well-characterized and widely used organisms for recombinant protein expression. E. coli grows quickly, is easy to manipulate genetically, and has a large body of existing protocols and tools available. Since this is my first time going through the full synthetic gene design and expression workflow, I felt it was best to work with a well-established system before moving on to more complex organisms.
What technologies could be used to produce this protein from your DNA?
One technology that can be used to produce this protein from my DNA is recombinant protein expression in bacteria, which is exactly the approach taken here. After obtaining and codon-optimizing the coding sequence (CDS), the gene is inserted into a plasmid vector that contains a promoter, ribosome binding site (RBS), terminator, and a selectable marker such as an antibiotic resistance gene. The plasmid is then transformed into E. coli cells, which are grown under selective conditions to ensure that only bacteria containing the correct vector survive.
Once inside the bacteria, the promoter drives transcription of the inserted DNA into mRNA. The ribosome then binds to the mRNA and translates the coding sequence into the corresponding protein. As the bacteria grow and divide, they produce increasing amounts of the recombinant protein. After sufficient expression, the protein can be purified. This results in the isolation of the desired protein.
Part 4: Prepare a Twist DNA Synthesis Order
Linear Map
After following all of the instructions on the website, I have obtained the following Linear Map:
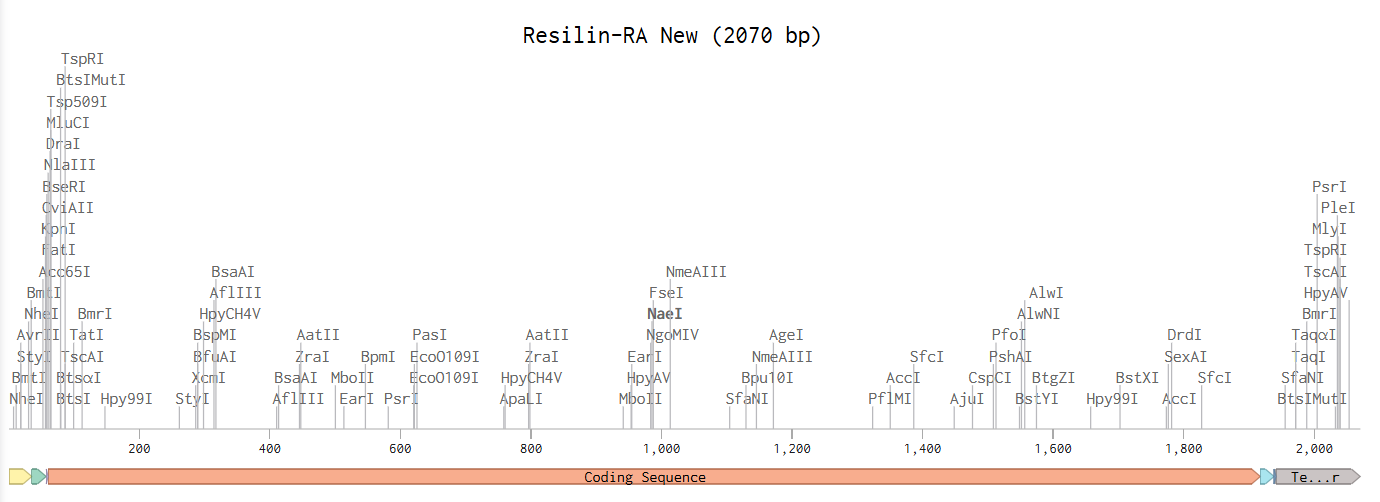

And here is the link to view it in Benchling: Resilin-RA expression cassette
When I proceeded to Twist, the platform flagged issues with my sequence and recommended re-optimization. After running their internal optimization, the system still classified the sequence as too complex, however, it did allow me to download the plasmid design.
Here it is:
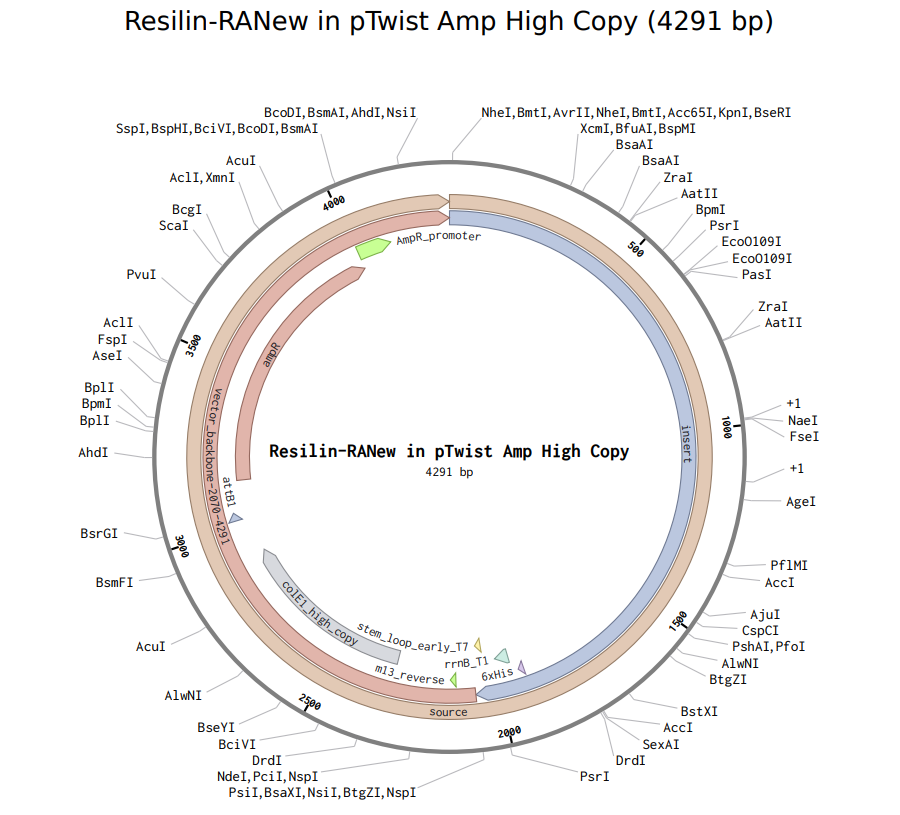


Yayyyyy :)
Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
I would like to sequence the gut microbiome of athletes before and after intense training, or before and after a traumatic injury (such as ligament tears or broken bones), to investigate whether specific bacterial compositions or functional genes correlate with recovery speed, inflammation levels, or injury resilience. I think it would be fascinating to explore whether the microbiome, often referred to as “the second brain”, plays an even greater role in recovery and overall health than we currently understand.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why
To obtain a microbiome sample, I would probably rely on a non-invasive stool sample collection method, since from what I have learned, this is typically how researchers study the gut microbiome. For sequencing, I would probably choose next-generation sequencing (NGS), specifically shotgun metagenomic sequencing. NGS allows for a lot of DNA fragments to be sequenced in parallel, enabling comprehensive analysis of complex microbial communities. This approach would allow me not only to identify the bacterial species present, but also to analyze functional genes that may be associated with inflammation regulation, tissue repair, and recovery capacity.
- Shotgun metagenomic sequencing using Illumina technology is considered a second-generation sequencing method because it enables massively parallel sequencing of millions of short DNA fragments simultaneously. Unlike first-generation Sanger sequencing, which reads one fragment at a time, second-generation platforms generate high-throughput short reads using sequencing-by-synthesis chemistry.
- The input for this method is the total DNA extracted from a stool sample. The extracted DNA is fragmented into smaller pieces, typically a few hundred base pairs long. Synthetic adapter sequences are ligated to both ends of each fragment, allowing them to bind to the sequencing flow cell. The fragments are then amplified by PCR to generate sufficient material, creating a sequencing library ready for analysis.
- In Illumina sequencing, DNA fragments bind to a flow cell and undergo cluster amplification. During sequencing-by-synthesis, fluorescently labeled nucleotides are incorporated one base at a time to synthesize the complementary strand. After each incorporation, a camera detects the emitted fluorescence signal, and the color corresponds to a specific base (A, T, C, or G). This process repeats in cycles, allowing the machine to determine the DNA sequence through base calling.
- The output consists of millions of short DNA reads, typically stored in FASTQ format. Each read includes both the nucleotide sequence and a quality score (a measure of confidence in each base call) for each base. These reads can then be computationally analyzed to determine microbial composition and identify functional genes within the microbiome.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?
I would like to synthesize DNA that can be transcribed and translated into a therapeutic protein “paste” (or hydrogel-like material) to support healing of joint cartilage, and specifically the meniscus. Since there isn’t a sufficient solution for meniscal tears today, and the effects of surgeries can be lifelong, I think that creating something like this could be game-changing. One of the main challenges with meniscus healing is that blood supply is not consistent across the tissue (especially toward the inner region), which makes delivery of therapeutics through the bloodstream difficult. Because of that, an externally applied material that can consistently deliver needed proteins or signaling molecules directly to the injury site might help the meniscus heal better and faster. More specifically, I would start by synthesizing a construct encoding a cartilage-supporting growth factor, such as members of the TGF-β superfamily (Fortier et al., 2011), or a small engineered protein designed to promote extracellular matrix production. This construct could eventually be incorporated into a biomaterial scaffold for localized and sustained protein delivery.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
The DNA would be synthesized using solid-phase phosphoramidite synthesis, which builds short oligonucleotides one nucleotide at a time. These short oligonucleotides are then assembled into a full-length gene using enzymatic methods such as ligation or Gibson Assembly, and cloned into a plasmid for verification and use. I would use this method because it is the current standard for accurate, high-fidelity DNA synthesis and allows precise control over the exact sequence being produced. It also enables incorporation of specific design elements such as codon optimization, regulatory sequences, or tags, making it great for constructing engineered therapeutic genes.
- Solid-phase phosphoramidite synthesis builds DNA one nucleotide at a time on a solid support. The process occurs in repeated chemical cycles. First, the growing DNA strand is attached to a solid surface. A protected nucleotide (phosphoramidite) is then added to the chain in a coupling reaction. After coupling, a capping step inactivates any strands that did not successfully react, which helps reduce errors. The protecting group on the newly added nucleotide is then removed (deprotection) to expose a reactive site for the next cycle. Finally, an oxidation step stabilizes the newly formed bond. This cycle repeats until the full oligonucleotide sequence is synthesized. After synthesis is complete, the DNA is cleaved from the solid support, fully deprotected, and purified. Since this chemical method efficiently produces short oligonucleotides, multiple synthesized fragments are then assembled using enzymes into a longer gene construct and sequence-verified..
- In solid-phase phosphoramidite synthesis, each chemical coupling step is slightly less than 100% efficient, meaning that errors accumulate as the sequence length increases. For this reason, individual oligonucleotides are typically limited to around 150–200 base pairs, and longer genes must be assembled from shorter fragments. Additionally, sequences that are highly repetitive or have extreme GC content can be more difficult to synthesize accurately.
5.3 DNA Edit
(i) What DNA would you want to edit and why?
I would like to explore editing the DNA of cancerous cells to selectively induce apoptosis (programmed cell death) and prevent uncontrolled cell division as a therapeutic strategy for cancer treatment. Many cancers arise from mutations in genes that regulate cell growth and survival. By using targeted gene editing tools to either restore normal tumor suppressor function or disrupt oncogenes specifically in cancer cells, it may be possible to stop tumor progression while minimizing damage to healthy tissue. A major challenge in this approach would be ensuring precise delivery to cancer cells while avoiding unintended edits in healthy cells.
(ii) What technology or technologies would you use to perform these DNA edits and why?
To perform these DNA edits, I would use a CRISPR-based editing platform, specifically base editing or prime editing, depending on the type of mutation involved. These technologies allow precise genetic modifications without introducing full double-strand breaks, which reduces the risk of unintended insertions or deletions. However, significant challenges remain, particularly achieving efficient delivery to cancer cells while minimizing off-target effects in healthy tissue.
- I would use prime editing, a CRISPR-based technology that uses a Cas9 nickase fused to a reverse transcriptase and a specialized guide RNA, pegRNA (a specially designed guide RNA that identifies the target site and contains the template for the desired genetic edit). The guide RNA directs the enzyme to a specific mutation in the cancer cell DNA. Instead of creating a full double-strand break, the system makes a single-strand nick and directly writes the corrected DNA sequence into the genome. The cell’s repair machinery then stabilizes the edit, making the change permanent.
- First, I would identify a mutation specific to the cancer type and design a pegRNA targeting that sequence. The editing components (Cas9 nickase–reverse transcriptase and pegRNA) would need to be packaged into a delivery system, such as targeted lipid nanoparticles that bind receptors overexpressed on cancer cells. The key inputs include the editing enzyme, the guide RNA, and a selective delivery vehicle to minimize effects on healthy cells.
- Although prime editing is more precise than traditional CRISPR-Cas9, challenges remain. Editing efficiency may be incomplete, and off-target effects are still possible. The biggest limitation is safe and selective delivery to cancer cells, since unintended edits in healthy tissues could cause harm.
References
Fortier, L. A., Barker, J. U., Strauss, E. J., McCarrel, T. M., & Cole, B. J. (2011).
The role of growth factors in cartilage repair. Clinical Orthopaedics and Related Research, 469(10), 2706–2715.