Week 3 HW Lab Automation


*You can find the Python Script for Opentrons Artwork part in the Week 3 Lab section!
Post-Lab Questions
1. Find and describe a published paper that utilizes the Opentrons or an automation tool to achieve novel biological applications.
The paper I chose, “An Automated Versatile Diagnostic Workflow for Infectious Disease Detection in Low-Resource Settings” (Urrutia Iturritza et al., 2024), presents a fully automated diagnostic pipeline built using the Opentrons OT-One-S Hood robot. The authors combined open-source modular automation with molecular biology protocols to create a workflow capable of detecting Neisseria meningitidis, a pathogen responsible for meningitis. What I found especially compelling is that the system automates the entire process - from DNA isolation using magnetic beads to isothermal amplification (RPA), enzymatic digestion, and final detection on a paper-based vertical flow microarray. Instead of focusing solely on analytical novelty, the study emphasizes integration: connecting multiple biological modules into a single continuous, robot-executed workflow.
The workflow processes eight samples in parallel and completes the full diagnostic pipeline in about 110 minutes, about 18% faster than manual processing by trained personnel. Even more interesting is the cost analysis: the estimated cost per sample (~USD 16) is significantly lower (~5.8× less per sample) than traditional PCR-based diagnostic tests. The use of recombinase polymerase amplification (RPA), which operates at constant temperature, eliminates the need for expensive thermocycling infrastructure. The detection step uses gold nanoparticles on paper-based microarrays, producing a visible signal, which makes the system potentially adaptable for decentralized or low-resource settings.
It is important to note that the workflow was not completely autonomous. Before running the protocol, researchers manually prepared the samples and placed the required reagents, consumables, microarrays, and supporting equipment on the robot deck. Some user interaction was still needed during the thermal cycling steps, and imaging and quantitative analysis of the microarrays were carried out manually after the run. Overall, while the liquid-handling and main molecular processes were automated, the system still relied on human setup and post-processing - showing both the strengths and current practical limits of laboratory automation.
This work highlights automation not as a luxury but as a strategy for improving accessibility, reproducibility, and safety in molecular diagnostics. By using an open-source robot such as Opentrons, standard laboratory consumables, and modular protocols, the authors demonstrate a strong proof of concept for automating nearly an entire diagnostic workflow. Importantly, this approach minimizes human handling steps, thereby reducing the risk of contamination and operational errors, while allowing scientists to focus on experimental design, interpretation, and optimization. In this sense, the engineering contribution lies not only in the individual modules, but in the successful integration of these modules into a cohesive, largely automated system with clear translational potential.
Resources: Miren, U. I., Mlotshwa, P., Gantelius, J., Alfvén, T., Loh, E., Karlsson, J., . . . Gaudenzi, G. (2024). An automated versatile diagnostic workflow for infectious disease detection in low-resource settings. Micromachines, 15(6), 708. doi:https://doi.org/10.3390/mi15060708
2. Write a description about what you intend to do with automation tools for your final project. You may include example pseudocode, Python scripts, 3D printed holders, a plan for how to use Ginkgo Nebula, and more. You may reference this week’s recitation slide deck for lab automation details.
Automation plan for BioGlue Meniscal Repair project
For my final project, I intend to use lab automation to systematically screen BioGlue formulations that deliver a cartilage-supporting growth factor (e.g., a TGF-β family protein). The goal of automation is to remove pipetting variability and enable parallel testing of many conditions (protein dose, gel composition, crosslinking conditions) in a reproducible way.
What I would automate:
A) Hydrogel formulation + dose matrix (OT-2 / Opentrons)
- Prepare a concentration series of growth factor (e.g., 0×, low, medium, high, very high).
- Dispense hydrogel precursor components into a 96-well plate (or small molds).
- Add growth factor to each well according to a planned matrix (dose × gel composition).
- Mix consistently (pipette mixing program).
- Start a standardized “release study” (how much protein leaves the gel over time) by overlaying buffer and scheduling timed sampling.
B) Automated sampling for release kinetics 6. At defined timepoints (e.g., 1h, 6h, 24h, 48h), the robot removes a small aliquot of supernatant and transfers it to a readout plate (for ELISA / fluorescence / total protein assay).
Final Project Ideas
As requested, I uploaded my ideas to the slide deck, but here they are too:
BioGlue for Meniscal Repair

At-Home Androgen Biosensing for PCOS (Wearable / Microfluidic Kit)


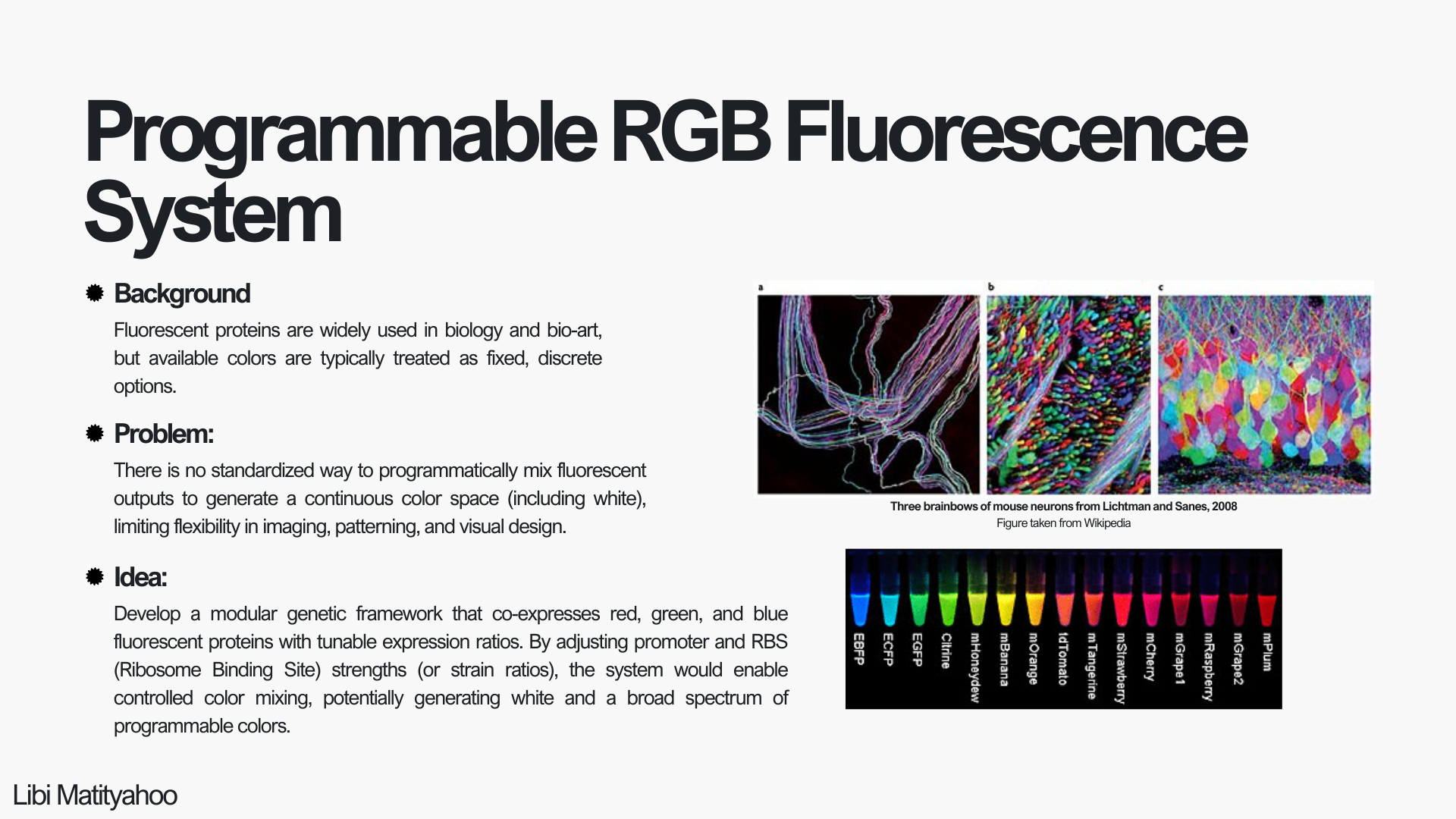
Programmable RGB Fluorescence System

