Week 6 HW: Genetic Circuits Part I

DNA Assembly Questions
- What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The Phusion High-Fidelity PCR Master Mix contains several key components that enable accurate and efficient DNA amplification. First, it includes the Phusion DNA polymerase, which is responsible for synthesizing new DNA strands. This enzyme has both 5’→3’ polymerase activity and 3’→5’ exonuclease (proofreading) activity, which allows it to correct mistakes during DNA replication and results in very high fidelity compared to standard polymerases like Taq. The mix also contains deoxynucleotides (dNTPs), which are the building blocks used by the polymerase to construct the new DNA strands. In addition, it includes an optimized reaction buffer with MgCl₂, which provides the proper chemical environment for the enzyme to function. Because all of these components are already included in the master mix, only the template DNA, primers, and water need to be added to perform the PCR.
- What are some factors that determine primer annealing temperature during PCR?
The primer annealing temperature is mainly determined by the primer’s melting temperature (Tm), which depends on:
- Primer length - longer primers have higher Tm
- GC content - more G–C pairs increase stability and raise Tm
- Salt concentration - higher ionic strength stabilizes primer–template binding and increases Tm
- There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
Both PCR and restriction enzyme digests can be used to generate linear DNA fragments, but they differ in how they work and when they are best used.
PCR:
- Uses primers and a DNA polymerase to amplify a specific region of DNA
- Can introduce mutations or modifications (e.g., through primer design)
- Does not require pre-existing restriction sites
- Useful when you want flexibility, to create new sequences, or to amplify a certain region
Restriction enzyme digest:
- Uses enzymes to cut DNA at specific recognition sites
- Produces predictable ends (sticky or blunt)
- Requires that the desired cut sites already exist in the DNA
- Useful when you want precise, reproducible cuts without introducing mutations
In general, PCR is preferable when you need to modify or design DNA fragments and amplify them, while restriction digests are preferable when you want to cut DNA in a precise and controlled way using existing sequences.
- How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
To be appropriate for Gibson cloning, the DNA fragments need to have overlapping homologous ends. This means that the ends of the fragments to be assembled should share matching sequences, usually around 20–40 bp, so they can anneal to each other during the Gibson reaction.
You can ensure this by:
- Designing PCR primers that add the correct overlap sequences to the ends of the amplified fragment
- Choosing restriction sites carefully so that the digested fragments have ends compatible with each other
So overall, the main idea is to make sure the fragments have the right matching overlaps for Gibson assembly to join them correctly.
- How does the plasmid DNA enter the E. coli cells during transformation?
Plasmid DNA enters E. coli during transformation when competent cells are heat shocked. The sudden temperature change causes the cell membrane to temporarily “open up,” allowing the DNA to enter the cell.
- Describe another assembly method in detail (such as Golden Gate Assembly)
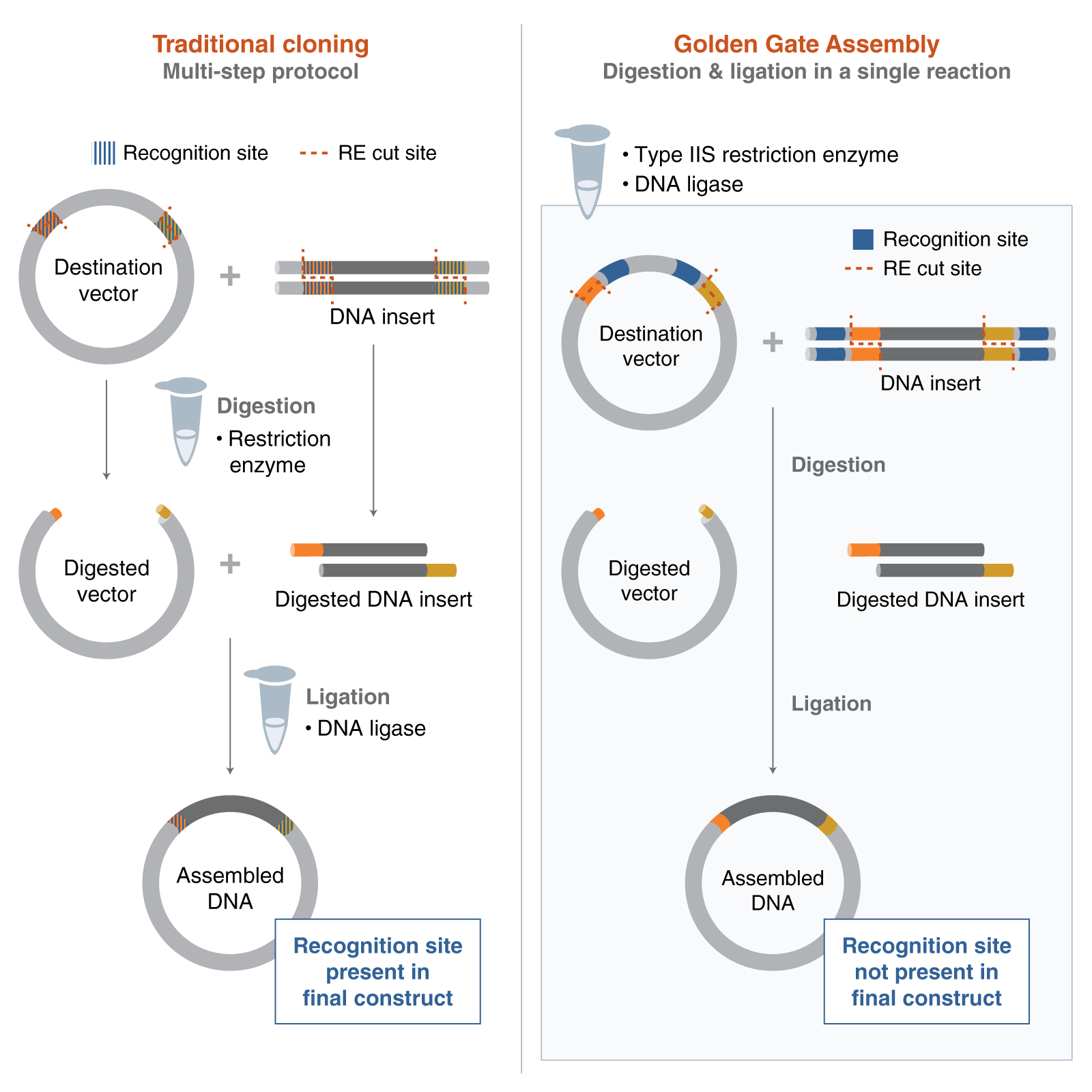
- Golden Gate Assembly is a DNA assembly method that uses Type IIS restriction enzymes and DNA ligase to join multiple fragments in a single reaction. Unlike standard restriction enzymes, Type IIS enzymes cut outside of their recognition site, which allows the creation of custom overhangs. These overhangs can be designed so that different DNA fragments assemble in a specific order and orientation.
In the reaction, the DNA is repeatedly cut and ligated in cycles, which increases the efficiency of correct assembly. Because the recognition sites are removed during the process, the final construct does not contain unwanted extra sequences. This makes Golden Gate especially useful for assembling multiple fragments seamlessly.
Here is a diagram taken from New England Biolabs that describes the method:

- Model this assembly method with Benchling or Asimov Kernel!
Asimov Kernel
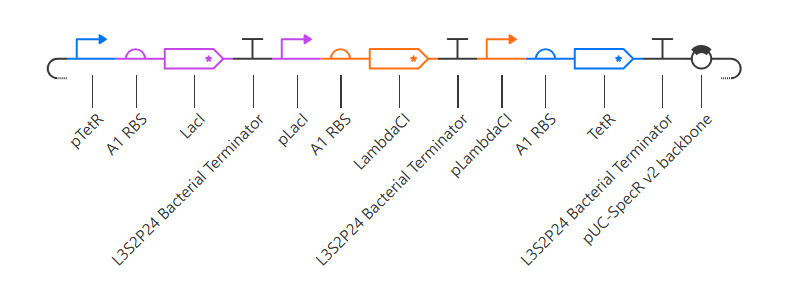
Here is my recreated repressilator:


Note that I used pUC-SpecR v2 backbone instead of pUC-SpecR v1 backbone, since I couldn’t find it.
My Repressilator’s simulation:
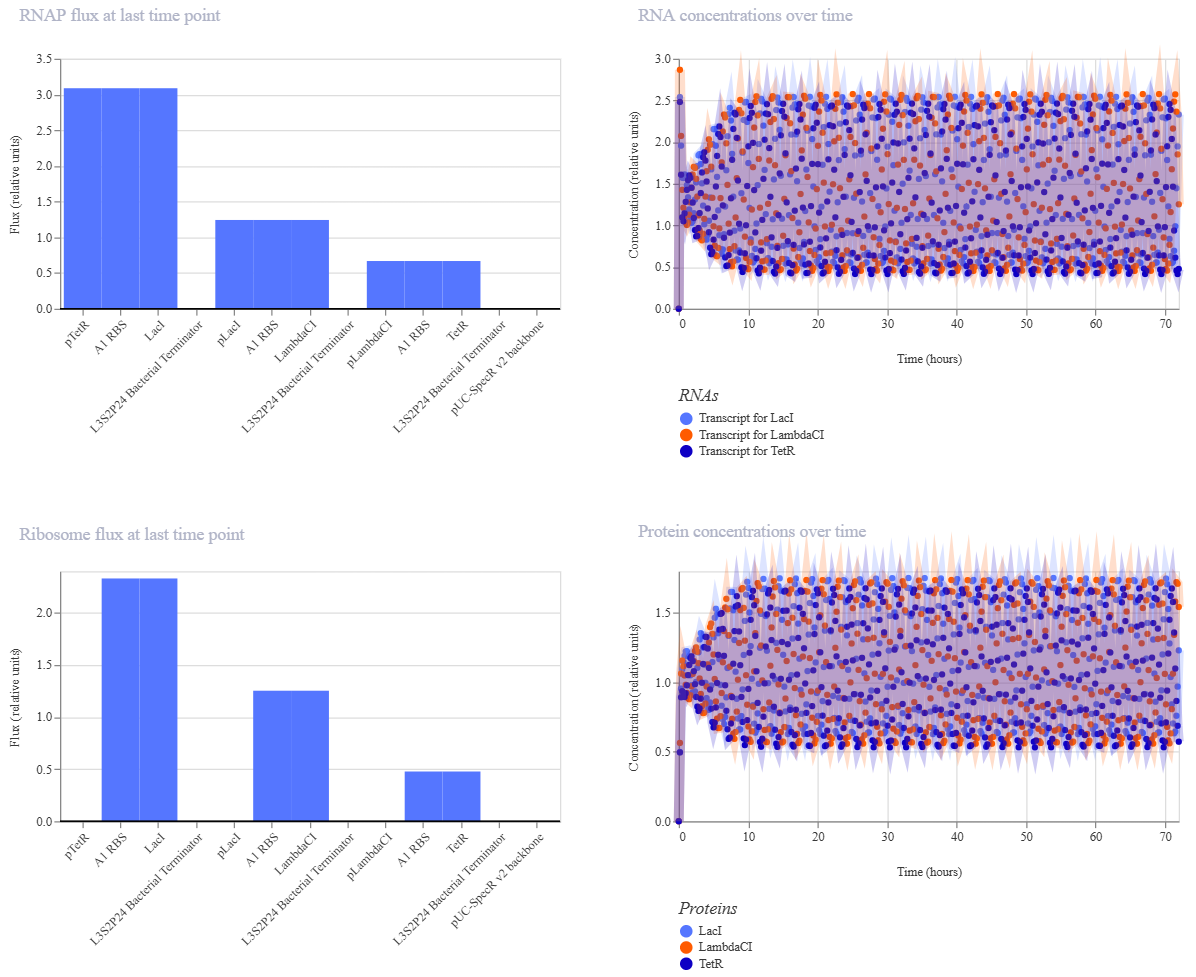

The Repressilator Construct from the Bacterial Demos repository:
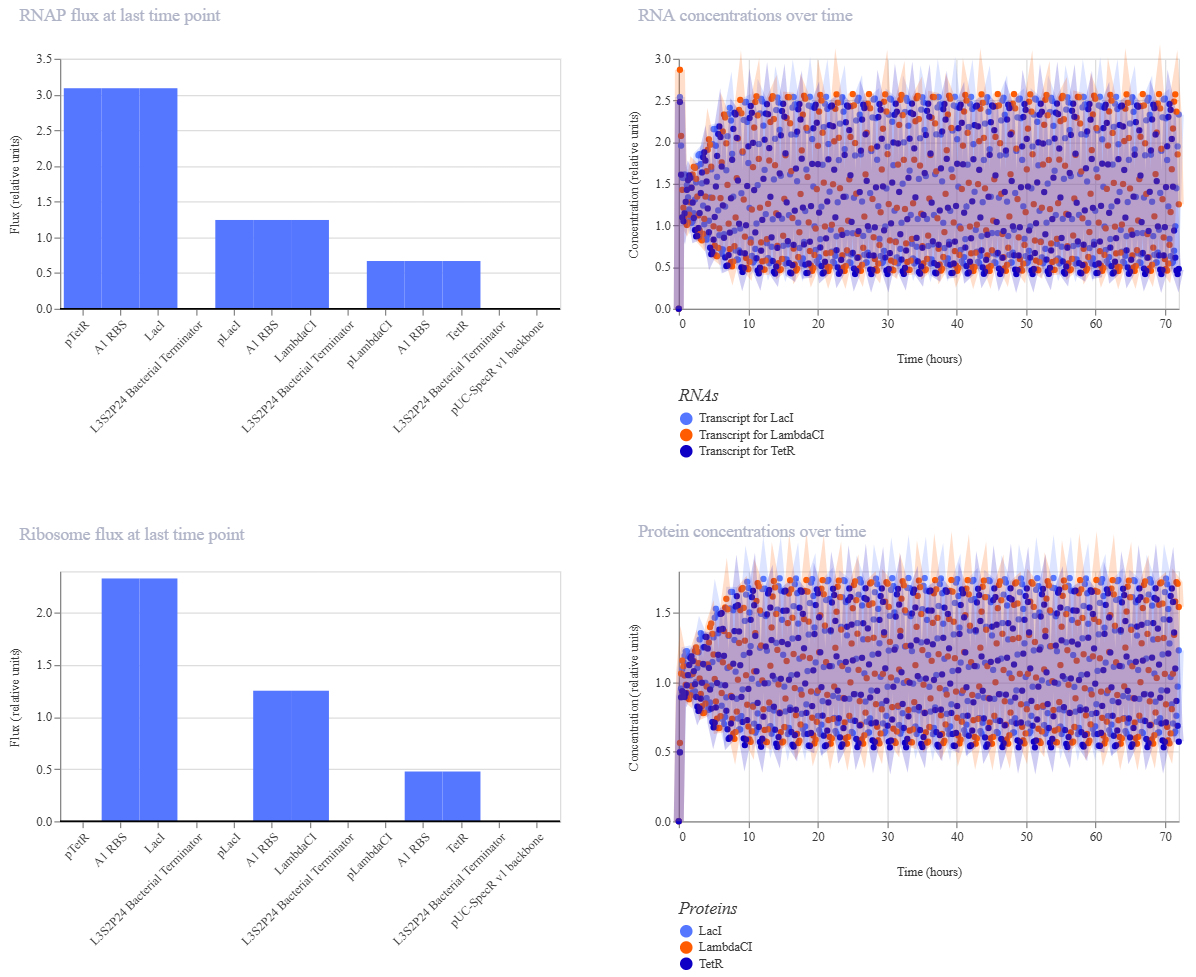

The two simulations look identical to the naked eye.