Week 11 HW: Bioproduction and Cloud Labs
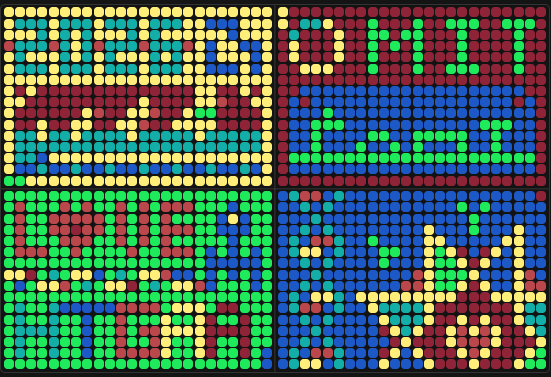


Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
My Contribution
My involvement in the community bioart project started right at the beginning. I found myself in a “digital battle” with other users over whether the top row on the left plate would be blue or yellow - I was on Team Yellow! No particular reason, just a gut feeling:) After a few minutes, others joined me:

It did indeed end up yellow in the final design!


What I liked
I really enjoyed the global aspect of the project - the fact that hundreds of people from all around the world contributed to a single piece of art is remarkable. I also loved the “evolutionary” dimension of it: being able to trace how the design changed and developed over time gave it a unique, living quality.
My favourite snapshot from the evolution:

What could be improved
For next year, I’d suggest two things:
- More colors - a broader palette would give participants more creative flexibility and make the process even more fun.
- More surface area - a larger canvas would allow for higher-resolution details and give more people the space to leave their mark on the final piece.
Overall, it was a wonderful idea and a genuinely fun experience to be part of!
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
1. Cell-Free Protein Synthesis Reaction Components
E. coli Lysate
| Component | Role |
|---|---|
| BL21 (DE3) Star Lysate (includes T7 RNA Polymerase) | Provides the core cellular machinery needed for transcription and translation, including ribosomes, enzymes, and co-factors. The T7 RNA Polymerase specifically transcribes DNA into mRNA using the T7 promoter system. |
Salts / Buffer
| Component | Role |
|---|---|
| Potassium Glutamate | Maintains ionic strength and mimics the intracellular environment, supporting enzyme activity and ribosome stability. |
| HEPES-KOH pH 7.5 | A buffering agent that maintains a stable, physiological pH (7.5), which is optimal for transcription and translation enzymes. |
| Magnesium Glutamate | Provides Mg²⁺ ions, which are essential cofactors for ribosomes, RNA polymerase, and many enzymes involved in protein synthesis. |
| Potassium Phosphate Monobasic | Works together with the dibasic form to maintain pH and provide phosphate ions, which are important for energy metabolism and nucleotide synthesis. |
| Potassium Phosphate Dibasic | Works together with the monobasic form to maintain pH and provide phosphate ions, which are important for energy metabolism and nucleotide synthesis. |
Energy / Nucleotide System
| Component | Role |
|---|---|
| Ribose | A simple sugar that serves as a carbon and energy source; can be metabolized to regenerate nucleotides and sustain the reaction over longer incubation periods. |
| Glucose | Another energy source that feeds into central metabolic pathways to regenerate ATP and other nucleotides needed for sustained transcription and translation. |
| AMP | A nucleotide monophosphate precursor that is phosphorylated to ATP, the primary energy currency that drives protein synthesis. |
| CMP | Precursor to CTP, which is required for RNA synthesis during transcription. |
| GMP | Precursor to GTP, which is required for both transcription and translation (e.g., ribosome translocation). |
| UMP | Precursor to UTP, which is required for RNA synthesis during transcription. |
| Guanine | A nucleobase that can be salvaged and converted into GMP/GTP, supplementing the guanosine nucleotide pool for transcription and translation. |
Translation Mix (Amino Acids)
| Component | Role |
|---|---|
| 17 Amino Acid Mix | Provides the bulk of the amino acid building blocks needed for the ribosome to synthesize polypeptide chains. |
| Tyrosine | Supplied separately because of its low solubility at neutral pH; it is an essential amino acid for building complete proteins. |
| Cysteine | Supplied separately due to its reactivity; it is essential for forming disulfide bonds and maintaining the structural integrity of many proteins. |
Additives
| Component | Role |
|---|---|
| Nicotinamide | A precursor to NAD⁺, which is a critical cofactor in metabolic redox reactions that regenerate energy to sustain the cell-free reaction. |
Backfill
| Component | Role |
|---|---|
| Nuclease-Free Water | Used to bring the reaction to its final volume without introducing RNases (ribonucleases) and DNases (deoxyribonucleases), enzymes that degrade RNA and DNA respectively, that could degrade the nucleic acids in the reaction. |
2. 1-Hour PEP/NTP vs. 20-Hour NMP-Ribose Master Mix: Key Differences
The most fundamental difference between the two master mixes is their energy and nucleotide system: the 1-hour PEP/NTP mix supplies energy directly via pre-made NTPs (ATP, GTP, CTP, UTP) and PEP-Mono (phosphoenolpyruvate monopotassium salt; regenerates NTPs by donating phosphate groups to NDPs produced during transcription) as an immediate energy source, while the 20-hour NMP-Ribose mix supplies energy indirectly via NMPs and simple sugars (ribose and glucose) that the lysate enzymes must first convert into usable NTPs. This makes the PEP/NTP system fast but short-lived, while the NMP-Ribose system is slower to get going but more sustainable over longer incubation periods. The 1-hour mix also contains more additives (Spermidine, DMSO, cAMP, NAD, Folinic Acid) to boost immediate transcription/translation activity, whereas the 20-hour mix keeps additives minimal (only Nicotinamide) since the reaction has more time to proceed on its own.
3. Bonus question
Although GMP is listed as 0.00 uM in the 20-hour mix, Guanine (the nucleobase) is still provided and can be converted into GMP through a process called nucleotide salvage, where enzymes in the lysate attach a ribose-phosphate group to the free Guanine base. The resulting GMP is then phosphorylated up to GTP, which can be used for transcription. This is a more cost-effective way to supply guanosine nucleotides compared to adding pre-made GMP directly.