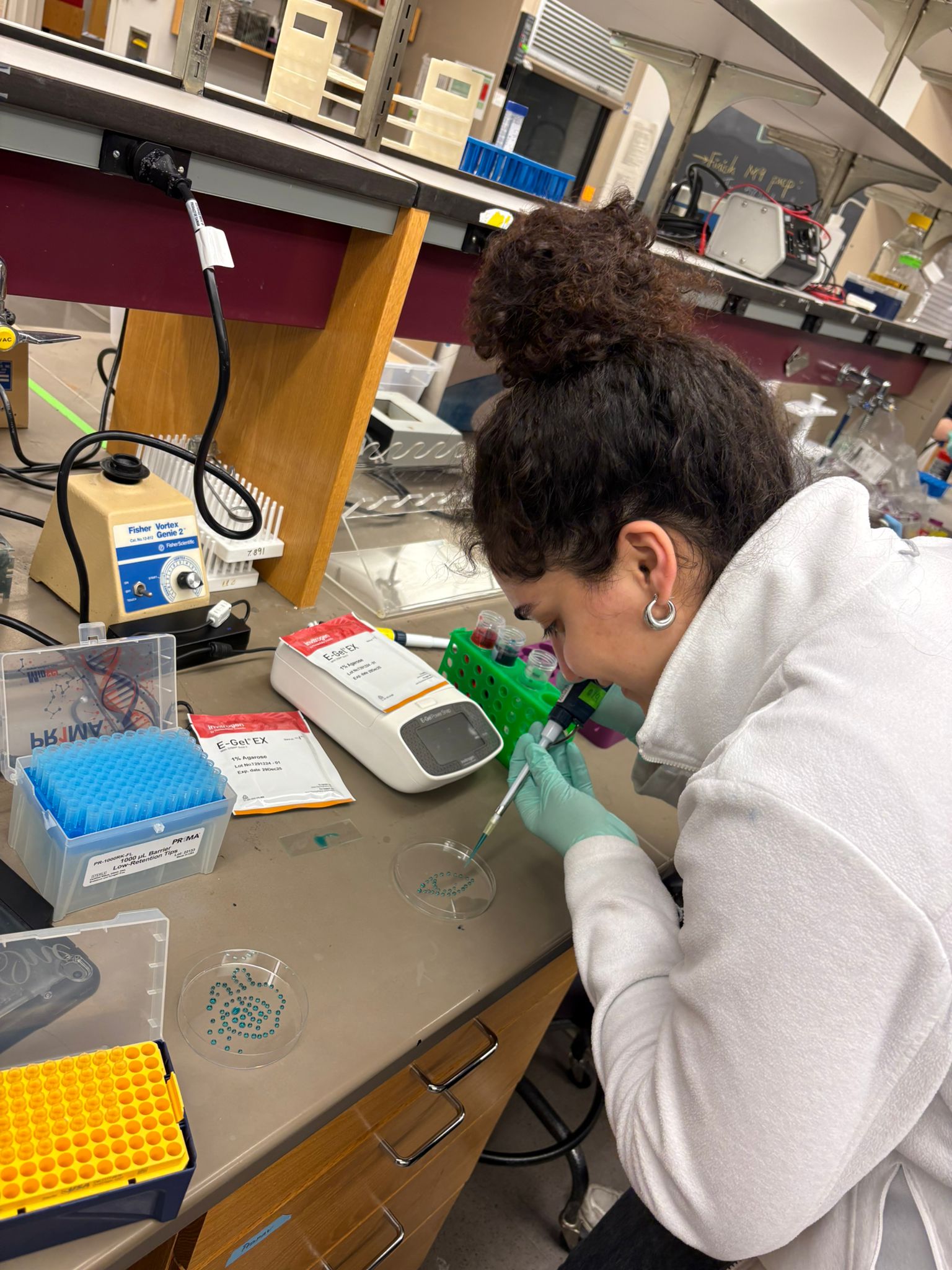
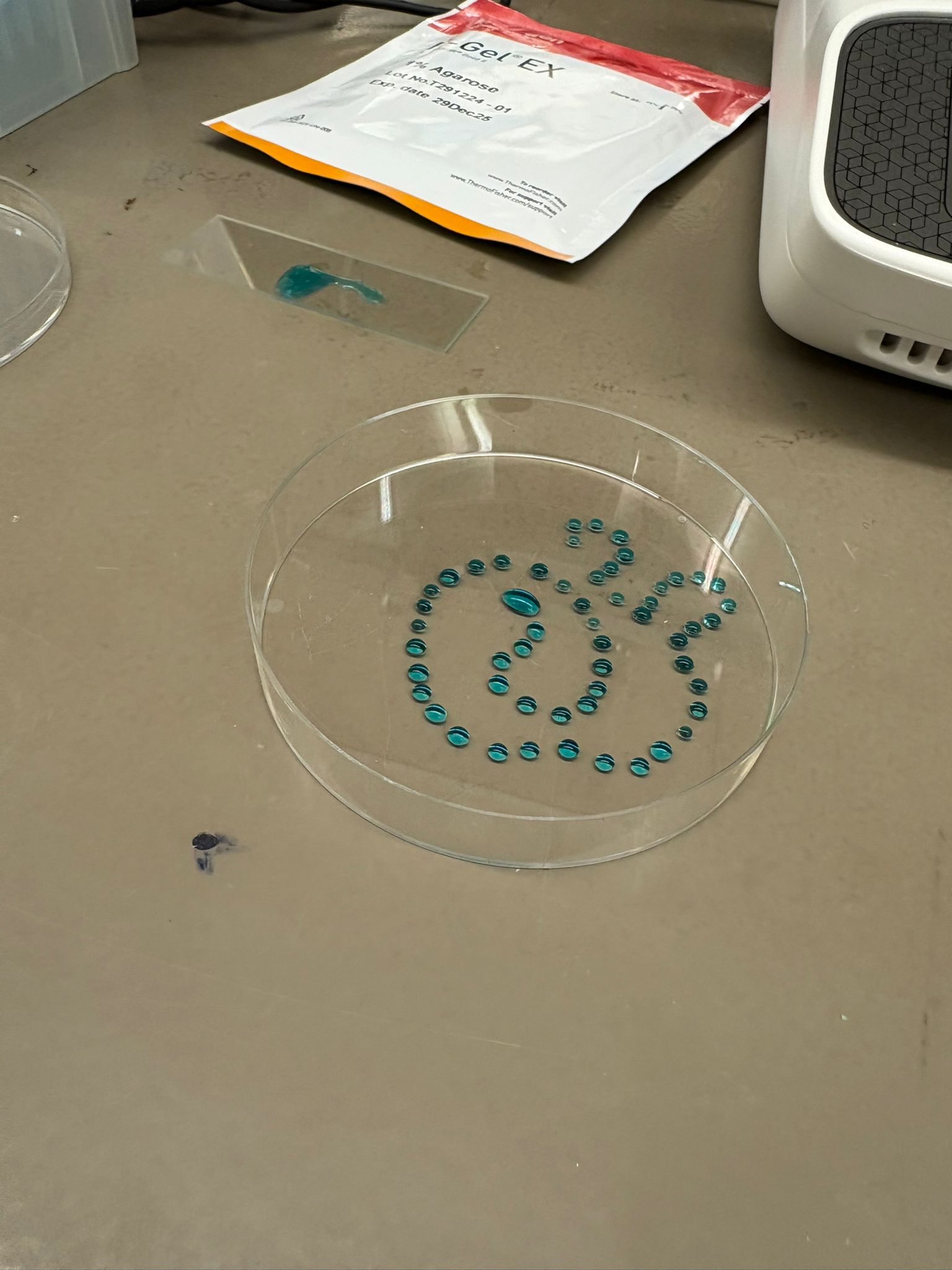
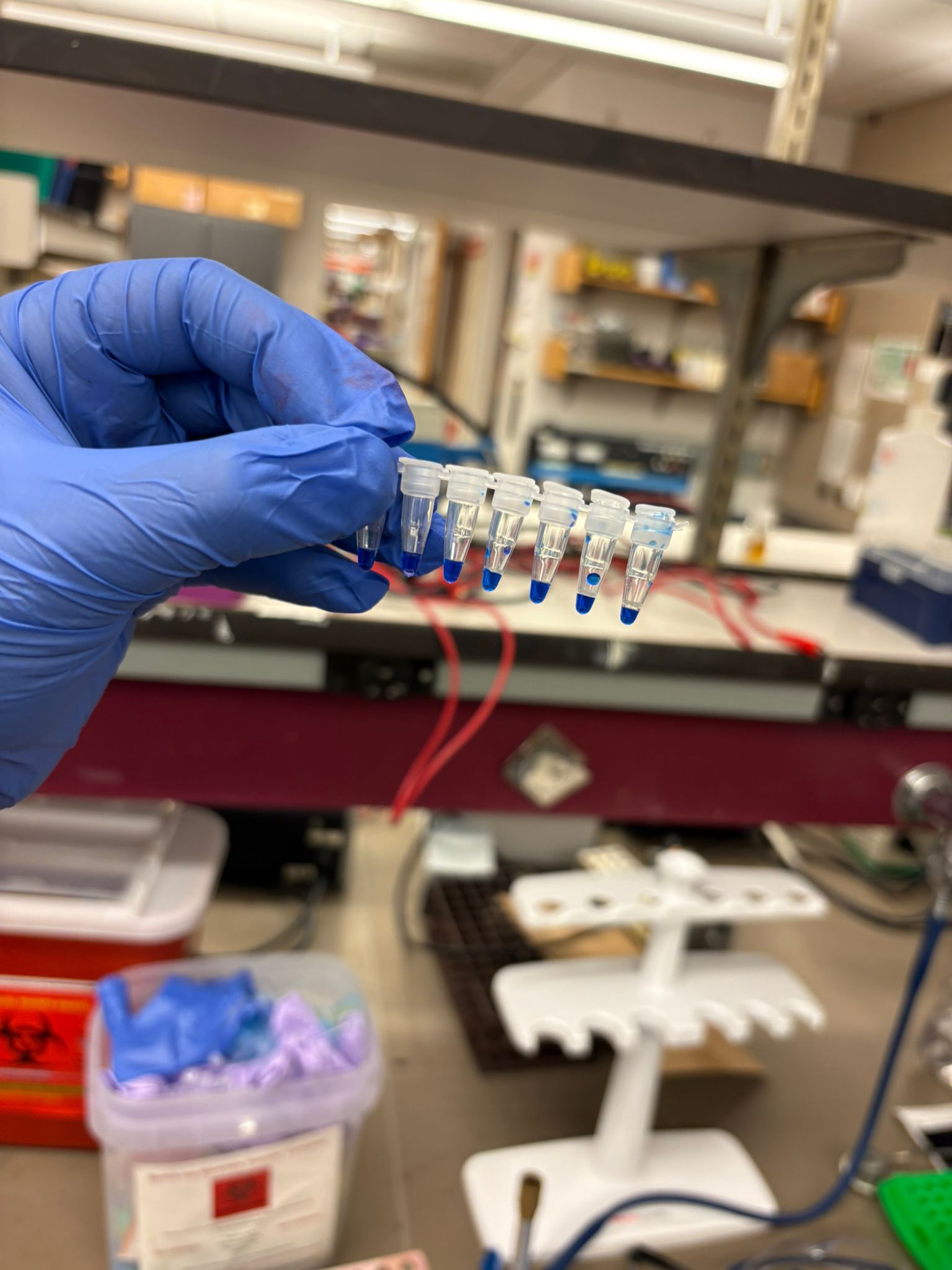
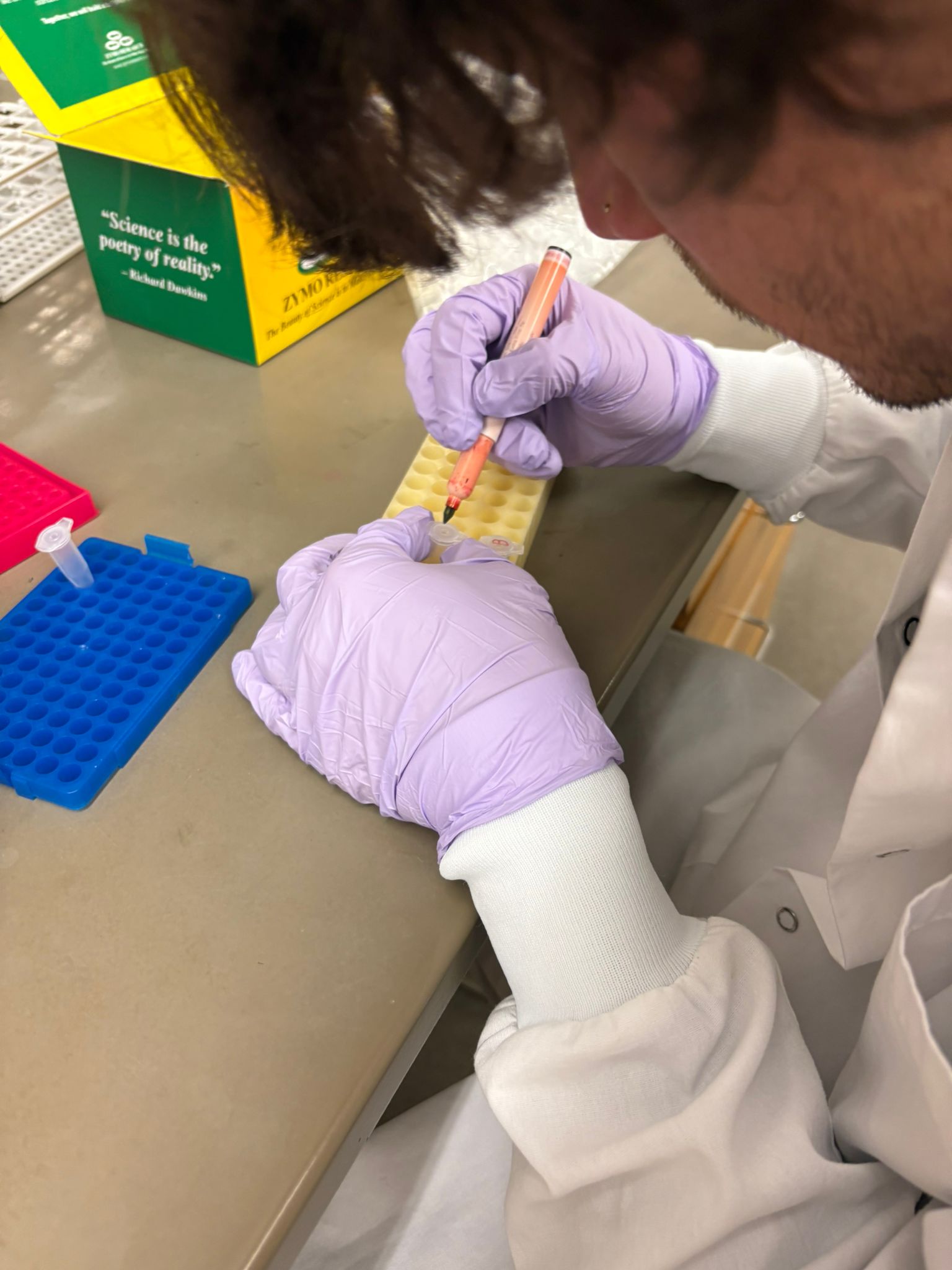
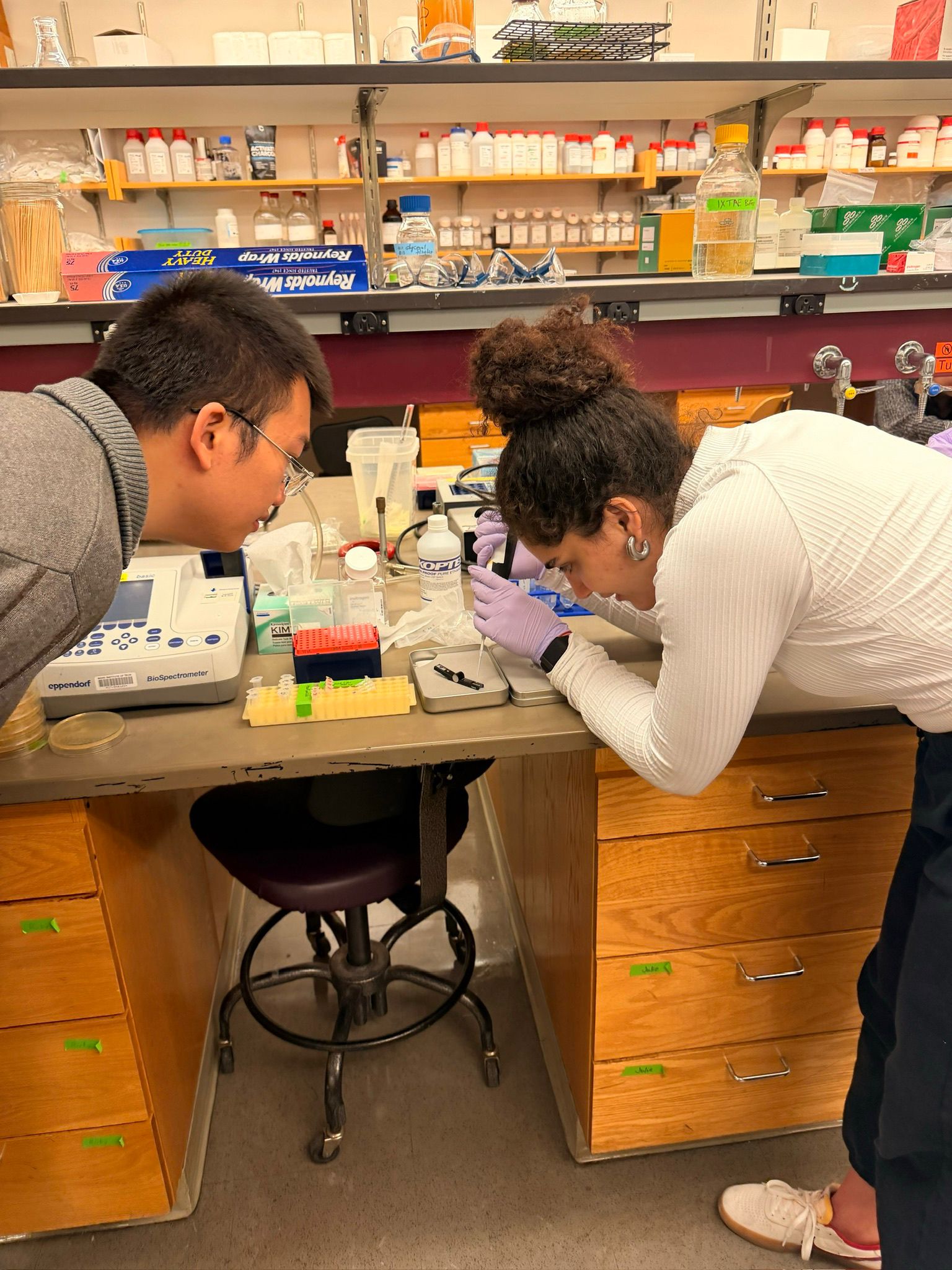
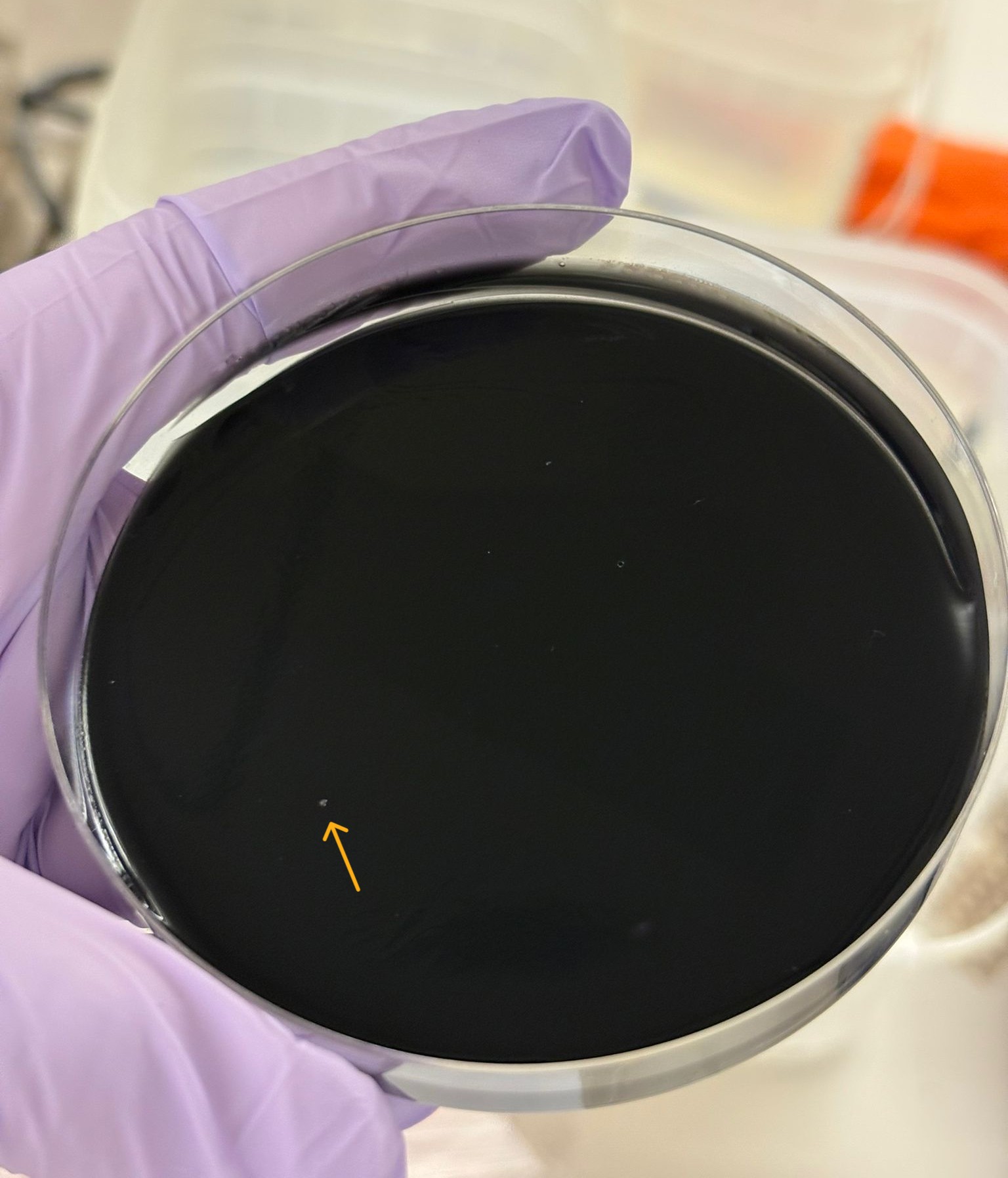
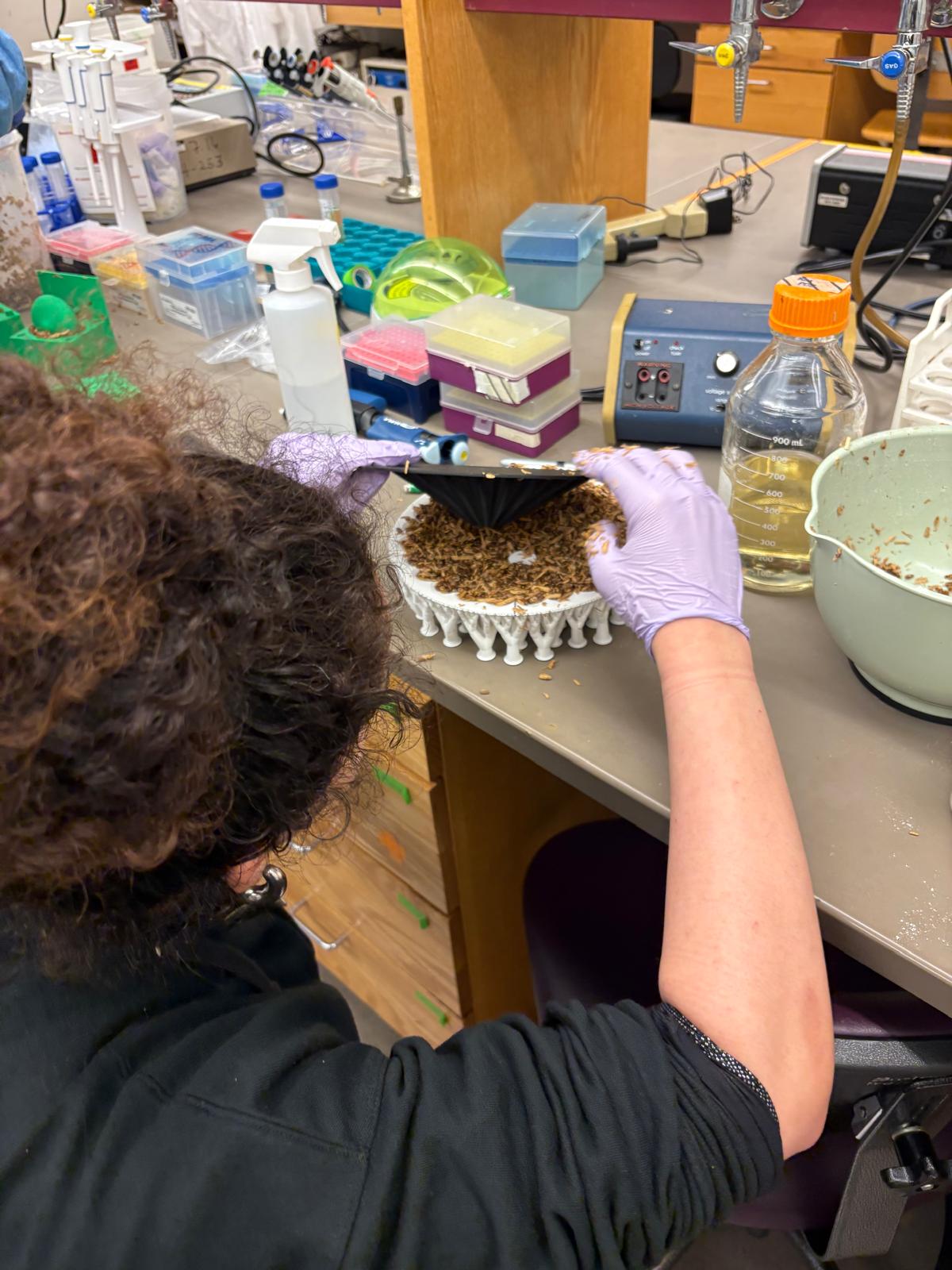
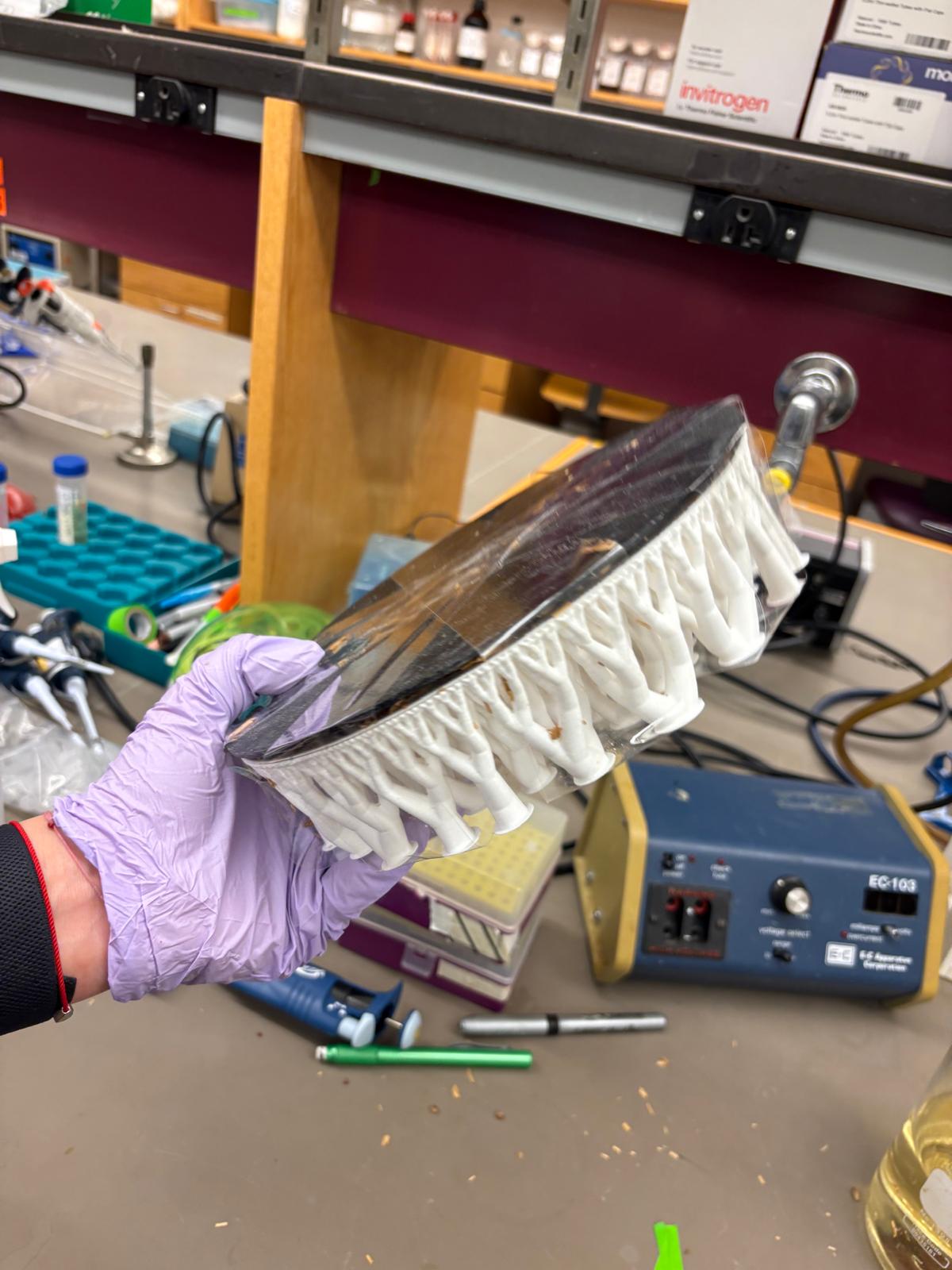
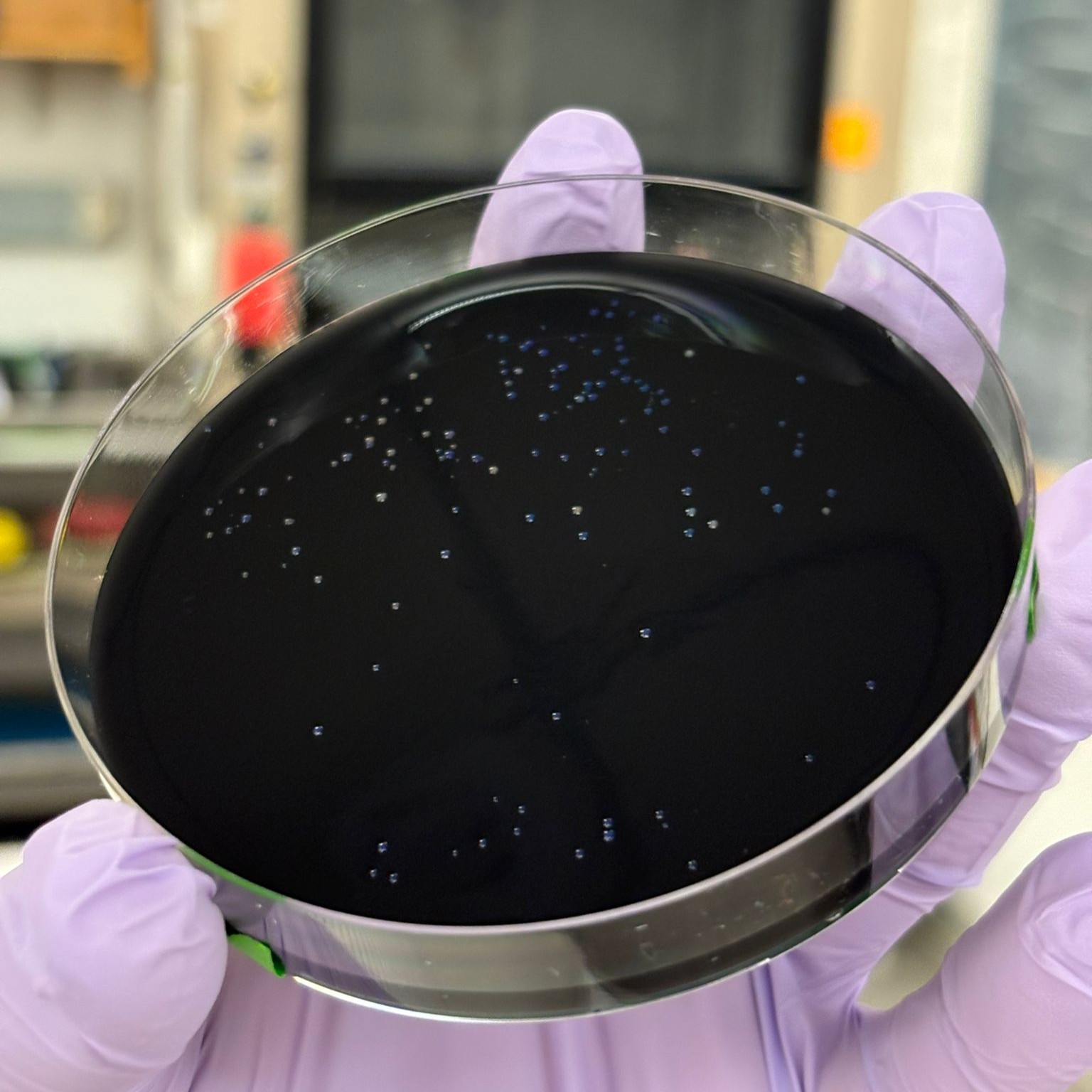
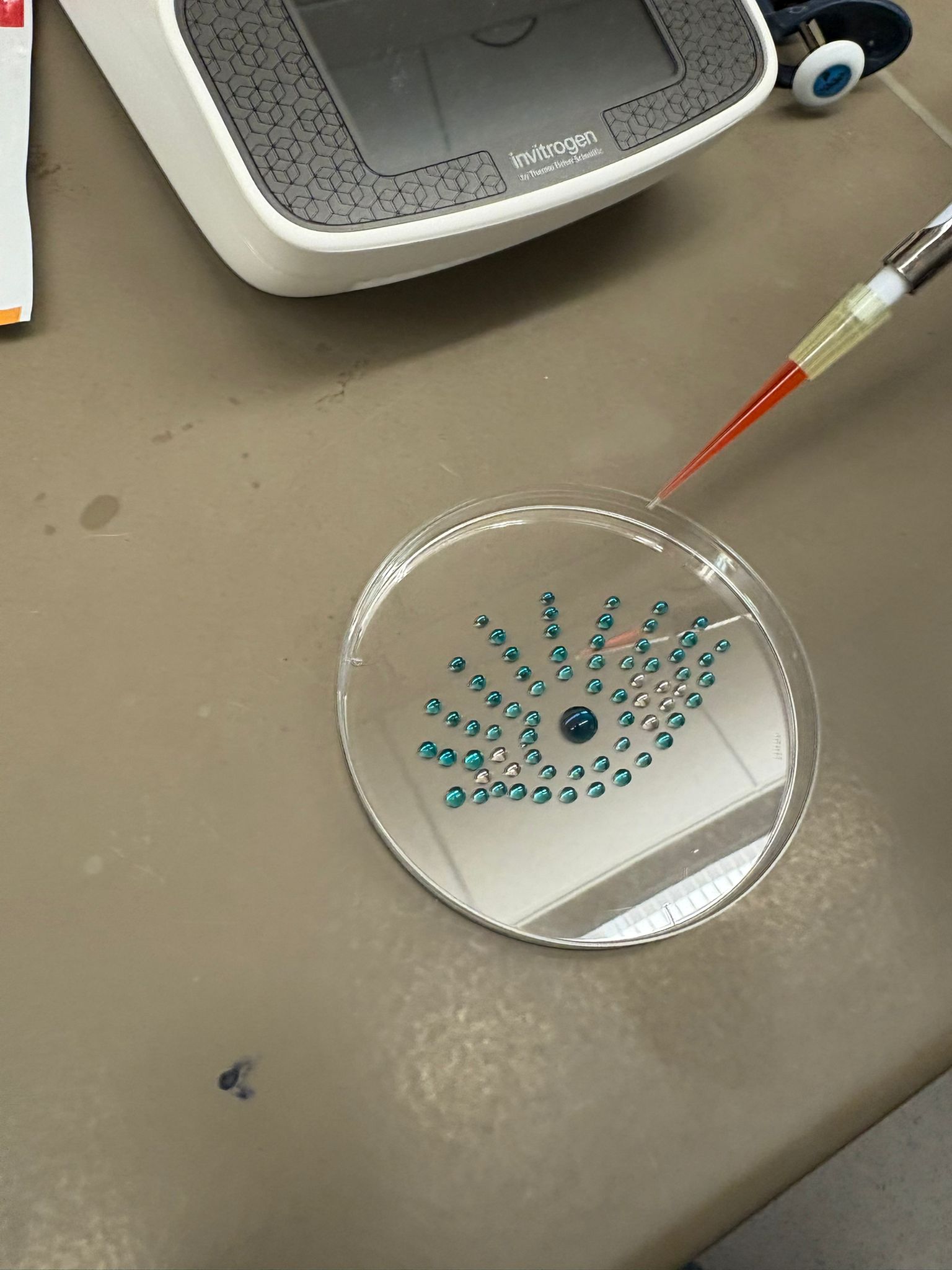
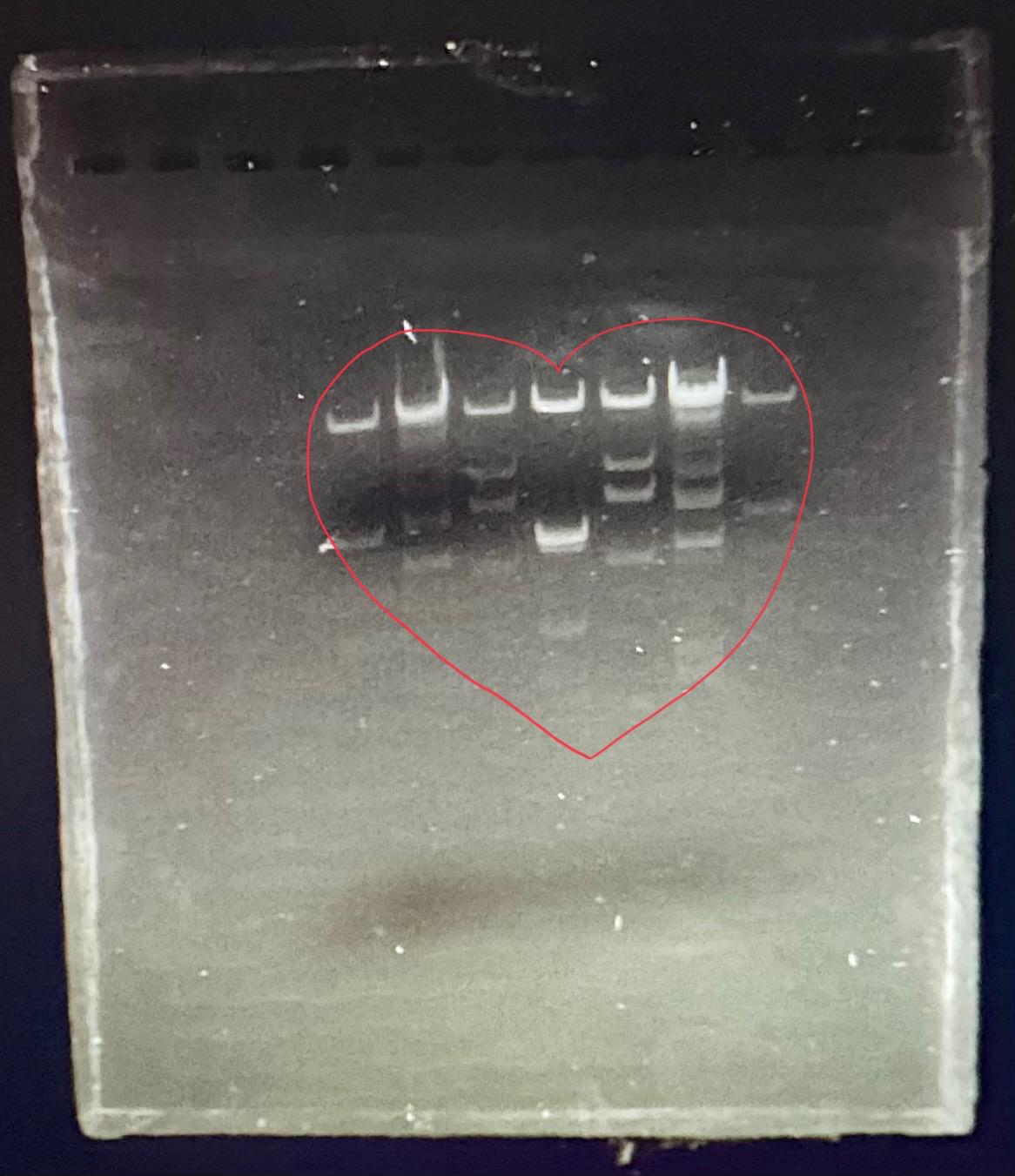
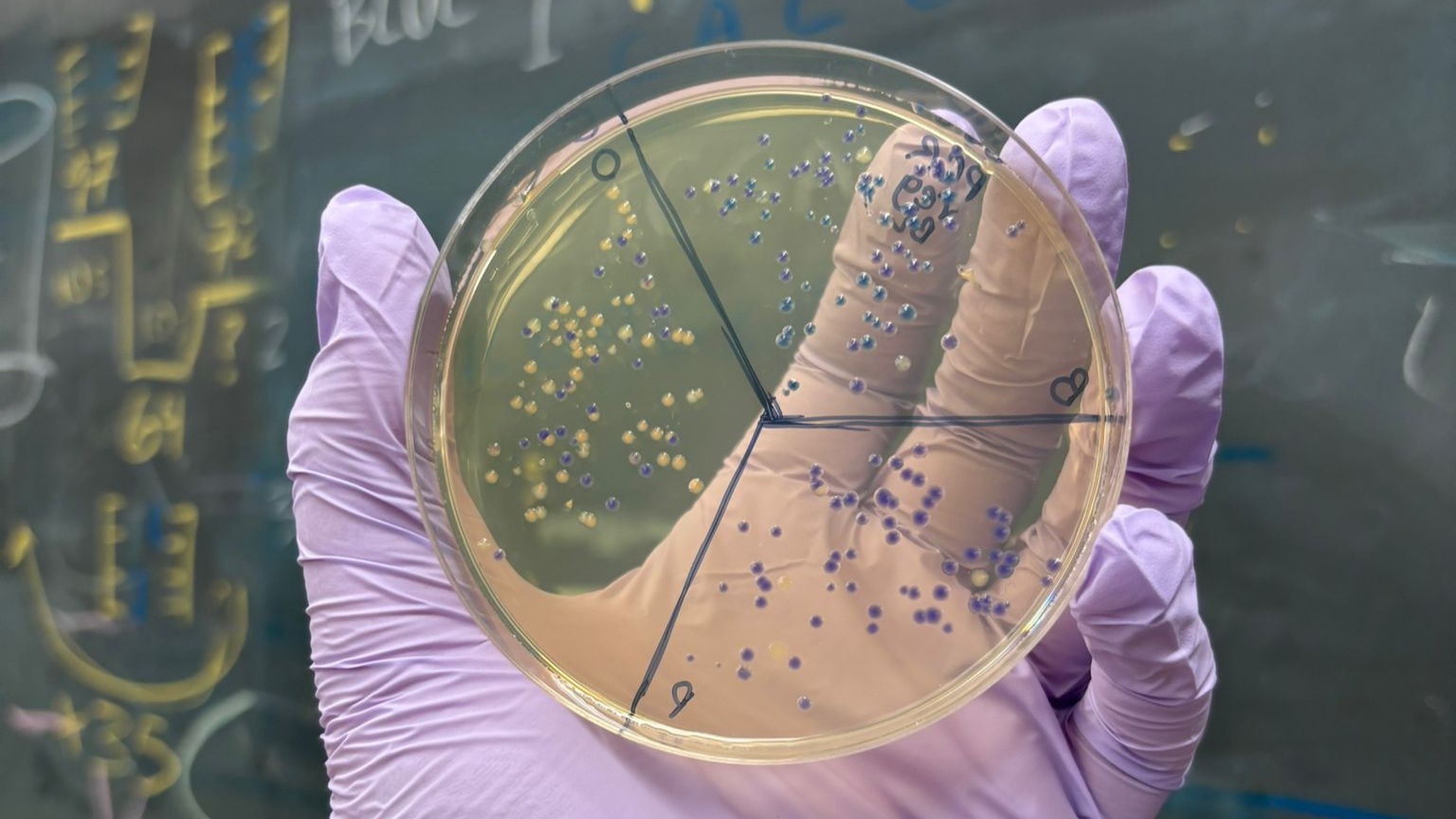
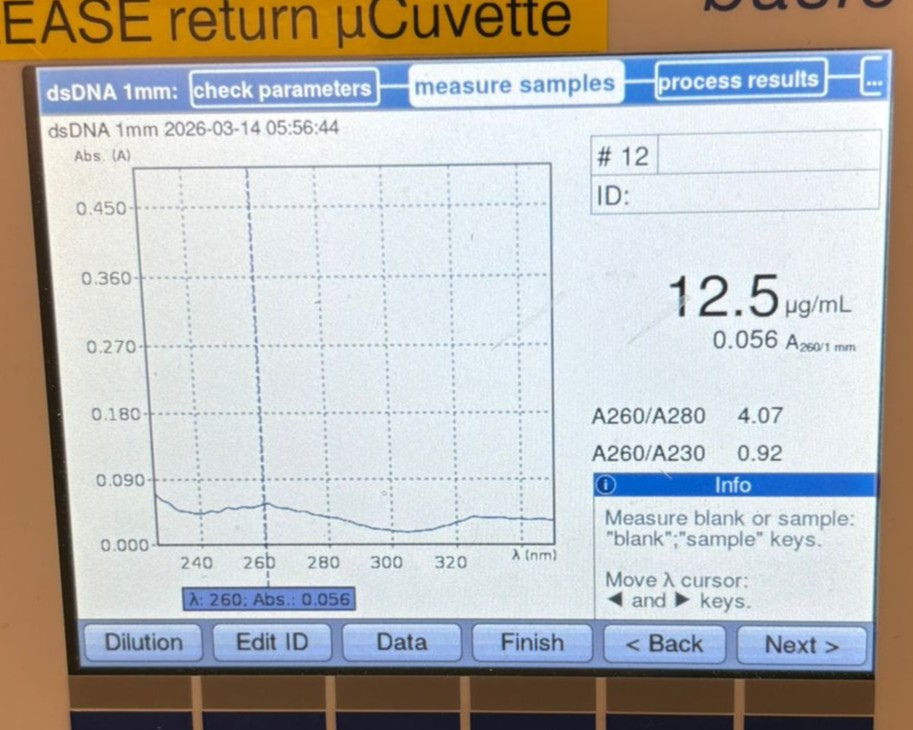
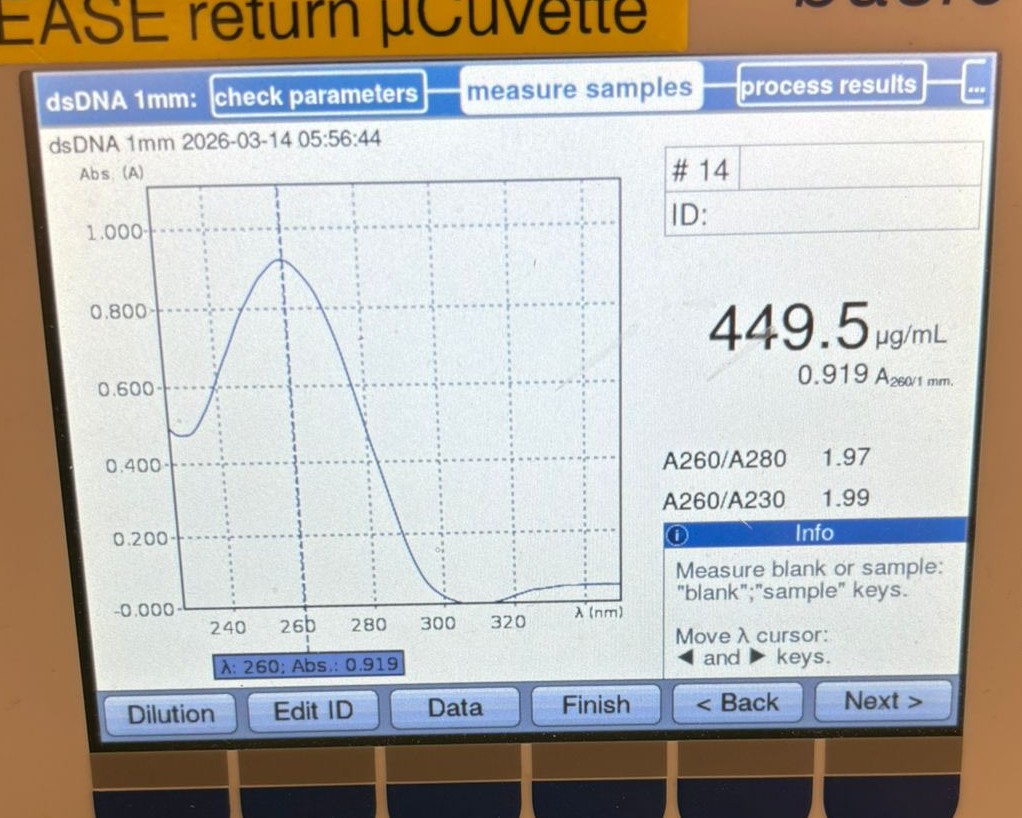
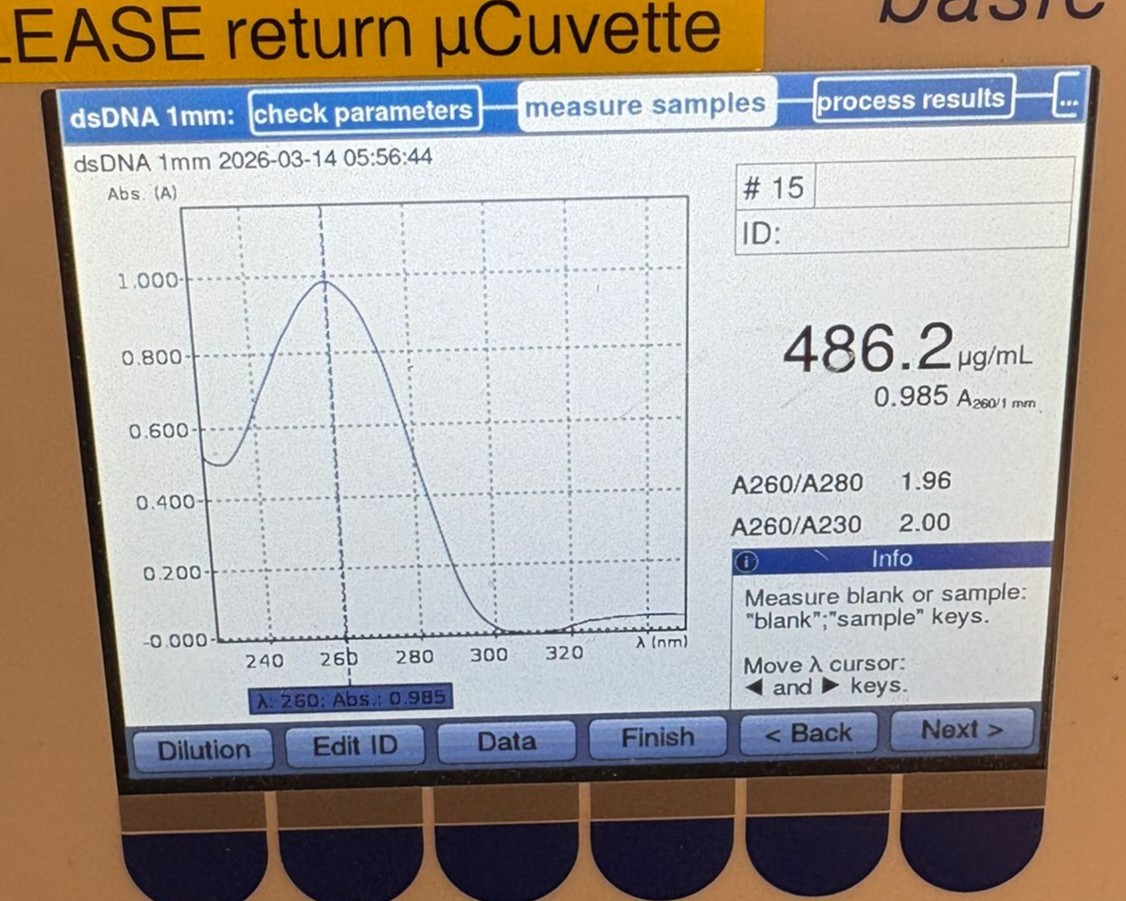
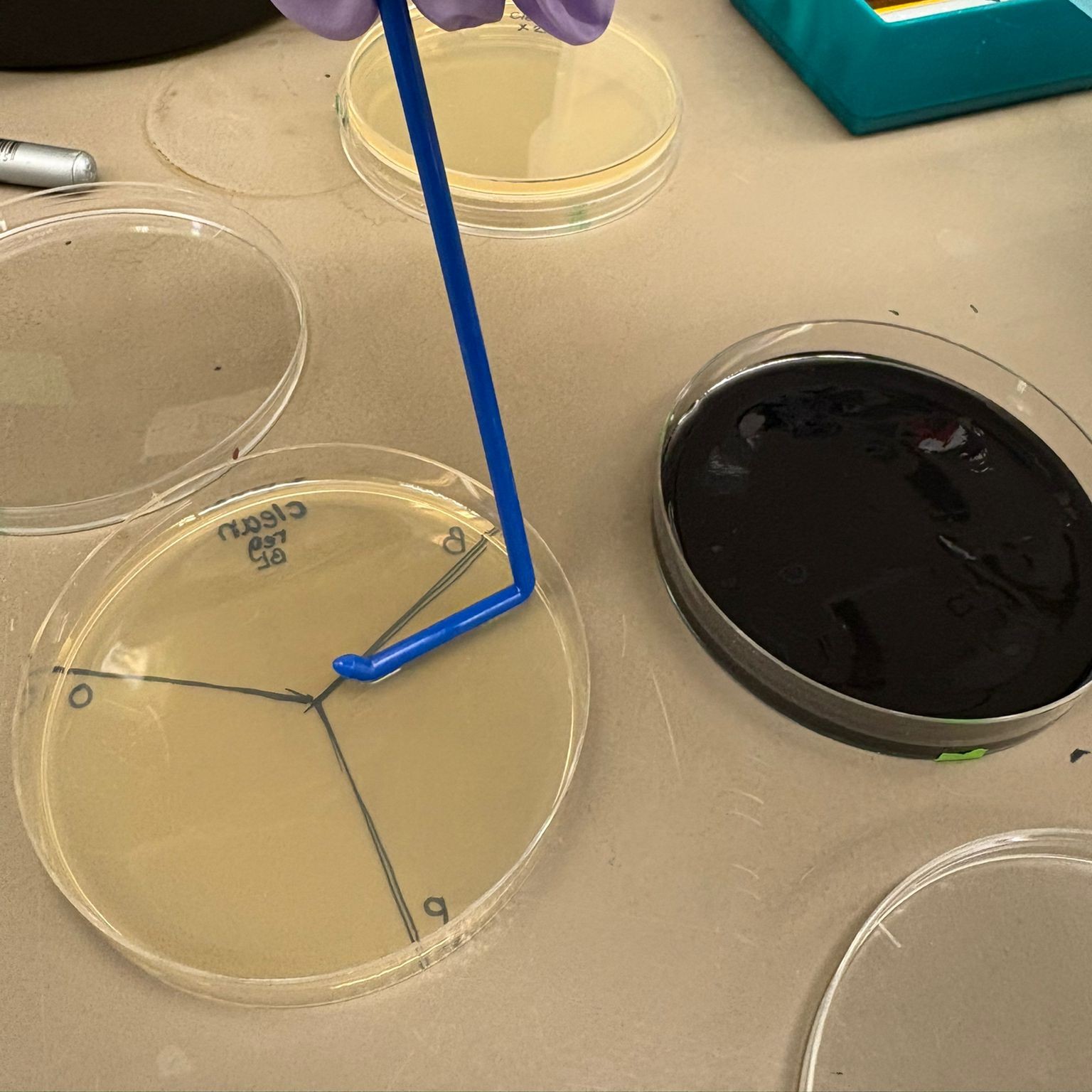
In this week’s lab, we used the Opentrons OT-2 pipetting robot and genetically engineered E. coli bacteria to create bio art on black (charcoal) agar plates! The bacteria were engineered to express fluorescent proteins (such as GFP – Green Fluorescent Protein), allowing them to glow in multiple colors under ultraviolet light.
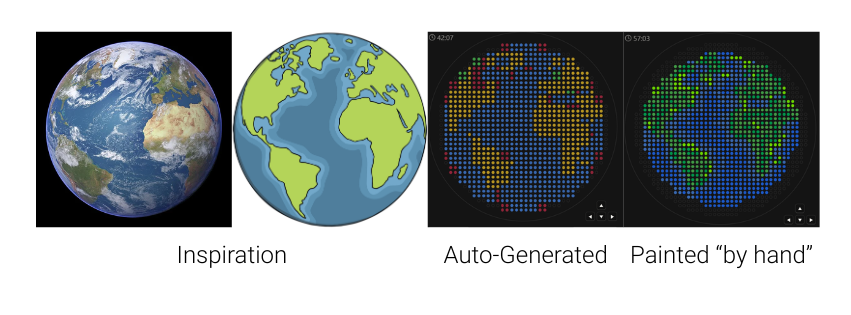
My first idea for this project was to create a picture of Earth that represents both the diverse backgrounds and countries students in the class and at MIT come from, and the global impact I hope to have through my work. I started by finding a reference photo and uploading it to the GUI. The result wasn’t very good, but I used it as a base and painted my final design over it. I would have loved to include some white (which we don’t have) for clouds, but I think it turned out amazing even without it.
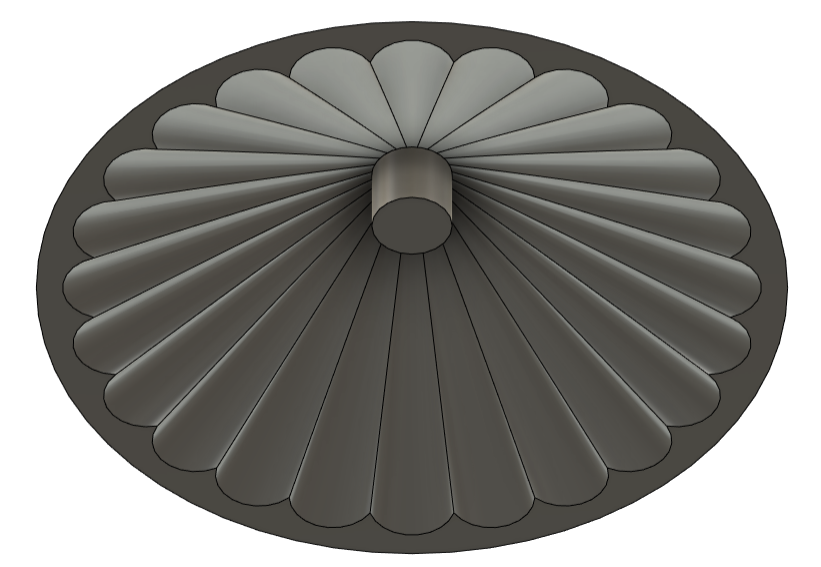
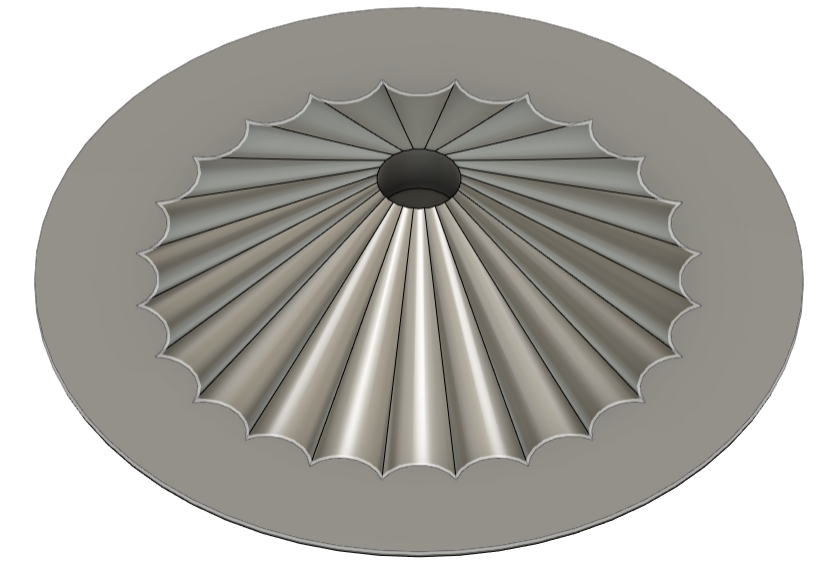
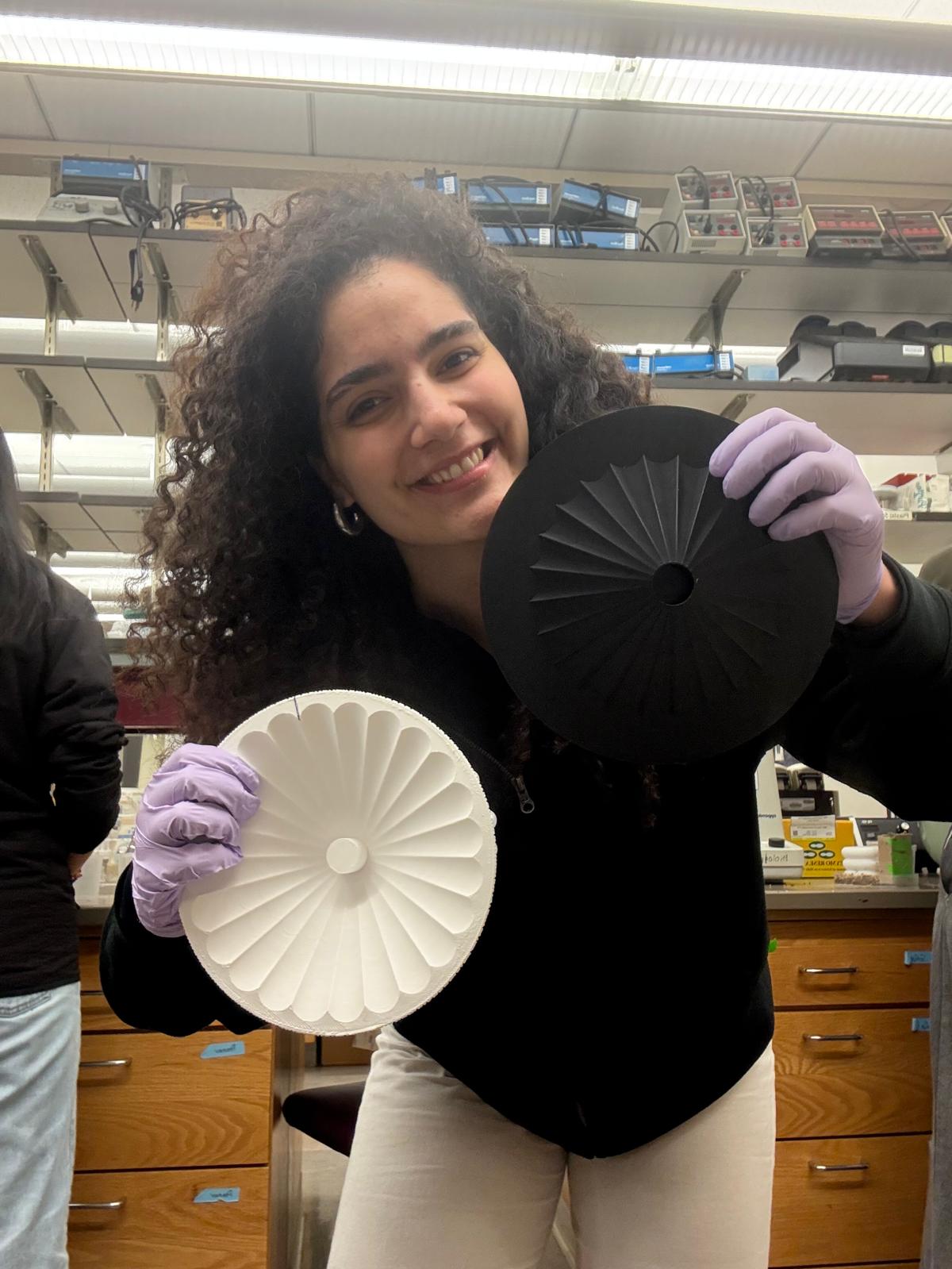
The “pixelated” look of the art examples from previous years reminded both me and my mother, who, among other things, is a textile designer, of cross-stitch embroidery. I wanted to create a design inspired by the classic rose cross patterns - but using this new, cutting-edge technology. A merge between the past and the future.
from opentrons import types
import string
metadata = {
'protocolName': '{YOUR NAME} - Opentrons Art - HTGAA',
'author': 'HTGAA',
'source': 'HTGAA 2026',
'apiLevel': '2.20'
}
Z_VALUE_AGAR = 2.0
POINT_SIZE = 0.75
venus_points = [(-9,33), (5,33), (7,33), (9,33), (11,31), (13,31), (11,29), (-19,25), (-17,25), (1,25), (3,25), (15,25), (-21,23), (-19,23), (-7,23), (-5,23), (-21,21), (-19,21), (-5,21), (-31,19), (-29,19), (-21,19), (-19,19), (-17,19), (-15,19), (-11,19), (29,19), (-31,17), (-29,17), (-13,17), (-11,17), (29,17), (-13,15), (23,15), (-13,13), (11,13), (31,13), (33,13), (11,11), (31,11), (33,11), (-23,9), (-21,9), (11,9), (-33,3), (15,3), (-33,1), (17,1), (29,-3), (27,-5), (-17,-7), (-15,-7), (-13,-7), (23,-7), (25,-7), (27,-7), (-13,-9), (-11,-9), (23,-9), (-9,-11), (-7,-11), (23,-11), (-7,-13), (27,-13), (25,-15), (-5,-17), (-5,-19), (-7,-21)]
mwasabi_points = [(-7,33), (-5,33), (-3,33), (-9,31), (-7,31), (-3,31), (-1,31), (-17,29), (-15,29), (-5,29), (-1,29), (1,29), (21,29), (-19,27), (-17,27), (-15,27), (-13,27), (-5,27), (-3,27), (-1,27), (13,27), (19,27), (-21,25), (11,25), (17,25), (23,25), (17,23), (19,23), (25,23), (27,23), (27,21), (-27,19), (-13,19), (7,19), (13,19), (15,19), (19,19), (25,19), (7,17), (11,17), (-27,15), (-25,15), (-17,15), (13,15), (15,15), (17,15), (29,15), (-27,13), (-23,13), (23,13), (25,13), (-35,11), (-31,11), (-21,11), (-15,11), (-13,11), (23,11), (-35,9), (-31,9), (-27,9), (-25,9), (-19,9), (7,9), (25,9), (27,9), (29,9), (-35,7), (-33,7), (-29,7), (-27,7), (5,7), (11,7), (13,7), (15,7), (17,7), (19,7), (21,7), (23,7), (27,7), (29,7), (31,7), (-33,5), (-29,5), (-23,5), (5,5), (11,5), (13,5), (23,5), (27,5), (33,5), (-29,3), (5,3), (19,3), (23,3), (29,3), (-31,1), (-29,1), (5,1), (11,1), (13,1), (25,1), (-31,-1), (5,-1), (15,-1), (17,-1), (25,-1), (-23,-3), (7,-3), (9,-3), (11,-3), (17,-3), (21,-3), (25,-3), (-19,-5), (19,-5), (23,-5), (25,-5), (19,-7), (-17,-9), (-15,-9), (19,-9), (-23,-11), (-21,-11), (-15,-11), (-13,-11), (-11,-11), (19,-11), (-21,-13), (19,-13), (21,-13), (23,-13), (-17,-15), (19,-15), (27,-15), (-17,-17), (-7,-17), (19,-17), (-17,-19), (-15,-19), (-11,-19), (-7,-19), (19,-19), (-13,-21), (-11,-21), (-13,-23), (-9,-23), (-7,-23), (-15,-25), (-11,-25), (-9,-25)]
azurite_points = [(-1,33), (3,33), (-15,31), (-13,31), (1,31), (7,31), (9,31), (15,31), (-11,29), (-9,29), (-7,29), (3,29), (7,29), (9,29), (-11,27), (-7,27), (1,27), (3,27), (5,27), (9,27), (11,27), (-15,25), (-3,25), (-1,25), (5,25), (7,25), (9,25), (13,25), (-17,23), (-3,23), (-1,23), (1,23), (5,23), (7,23), (11,23), (-17,21), (-15,21), (-13,21), (-11,21), (-9,21), (-7,21), (-3,21), (-1,21), (1,21), (7,21), (9,21), (11,21), (-9,19), (-7,19), (-5,19), (-1,19), (5,19), (-9,17), (-7,17), (-3,17), (-1,17), (5,17), (-11,15), (-9,15), (-1,15), (1,15), (3,15), (5,15), (7,15), (25,15), (27,15), (-11,13), (-9,13), (1,13), (5,13), (7,13), (13,13), (17,13), (19,13), (21,13), (-11,11), (-9,11), (-1,11), (3,11), (5,11), (7,11), (9,11), (13,11), (15,11), (19,11), (21,11), (-17,9), (-15,9), (-13,9), (-11,9), (-9,9), (5,9), (-21,7), (-19,7), (-17,7), (-15,7), (-3,7), (1,7), (3,7), (-21,5), (-19,5), (-17,5), (1,5), (3,5), (-25,3), (-23,3), (-21,3), (-15,3), (-3,3), (-1,3), (1,3), (3,3), (31,3), (33,3), (-35,1), (-25,1), (-23,1), (-21,1), (1,1), (3,1), (29,1), (31,1), (-35,-1), (-33,-1), (-21,-1), (-19,-1), (-17,-1), (-5,-1), (3,-1), (29,-1), (31,-1), (-35,-3), (-33,-3), (-31,-3), (-27,-3), (-21,-3), (-19,-3), (-17,-3), (-15,-3), (-11,-3), (-3,-3), (3,-3), (5,-3), (31,-3), (-35,-5), (-33,-5), (-31,-5), (-29,-5), (-27,-5), (-17,-5), (-15,-5), (-13,-5), (3,-5), (5,-5), (7,-5), (9,-5), (11,-5), (29,-5), (31,-5), (-35,-7), (-33,-7), (-25,-7), (-11,-7), (-9,-7), (-7,-7), (5,-7), (7,-7), (9,-7), (11,-7), (13,-7), (29,-7), (-35,-9), (-33,-9), (-31,-9), (-25,-9), (-23,-9), (-9,-9), (-7,-9), (-5,-9), (-3,-9), (7,-9), (11,-9), (13,-9), (25,-9), (27,-9), (-25,-11), (-5,-11), (13,-11), (15,-11), (25,-11), (27,-11), (-25,-13), (-5,-13), (-3,-13), (1,-13), (13,-13), (15,-13), (25,-13), (29,-13), (-23,-15), (-3,-15), (1,-15), (13,-15), (15,-15), (23,-15), (29,-15), (-23,-17), (-21,-17), (-19,-17), (-3,-17), (1,-17), (13,-17), (15,-17), (23,-17), (25,-17), (-19,-19), (-3,-19), (13,-19), (15,-19), (23,-19), (-19,-21), (-17,-21), (-5,-21), (-3,-21), (-1,-21), (9,-21), (13,-21), (15,-21), (17,-21), (21,-21), (23,-21), (-27,-23), (-25,-23), (-17,-23), (15,-23), (17,-23), (-23,-25), (-17,-25), (-7,-25), (-5,-25), (-21,-27), (-19,-27), (-15,-27), (-9,-27), (-7,-27), (-21,-29), (-19,-29), (-15,-29), (-13,-29), (9,-29), (-15,-31), (-13,-31), (5,-31), (9,-31)]
electra2_points = [(-7,35), (-1,35), (1,35), (3,35), (5,35), (1,33), (3,31), (5,31), (5,29), (-9,27), (7,27), (-13,25), (-11,25), (-15,23), (-13,23), (-11,23), (3,23), (9,23), (3,21), (5,21), (-3,19), (1,19), (3,19), (-5,17), (1,17), (3,17), (-7,15), (-5,15), (-3,15), (-7,13), (-5,13), (-3,13), (-1,13), (3,13), (15,13), (-7,11), (-5,11), (-3,11), (1,11), (17,11), (-7,9), (-5,9), (-3,9), (-1,9), (1,9), (3,9), (-13,7), (-11,7), (-9,7), (-7,7), (-5,7), (-1,7), (-15,5), (-13,5), (-11,5), (-9,5), (-7,5), (-5,5), (-3,5), (-1,5), (-19,3), (-17,3), (-13,3), (-11,3), (-9,3), (-7,3), (-5,3), (35,3), (-19,1), (-17,1), (-15,1), (-13,1), (-11,1), (-9,1), (-7,1), (-5,1), (-3,1), (-1,1), (33,1), (35,1), (-15,-1), (-13,-1), (-11,-1), (-9,-1), (-7,-1), (-3,-1), (-1,-1), (1,-1), (33,-1), (35,-1), (-13,-3), (-9,-3), (-7,-3), (-5,-3), (-1,-3), (1,-3), (33,-3), (35,-3), (-11,-5), (-9,-5), (-7,-5), (-5,-5), (-3,-5), (-1,-5), (1,-5), (33,-5), (35,-5), (-31,-7), (-29,-7), (-27,-7), (-5,-7), (-3,-7), (-1,-7), (1,-7), (3,-7), (31,-7), (33,-7), (35,-7), (-29,-9), (-27,-9), (-1,-9), (1,-9), (3,-9), (5,-9), (9,-9), (29,-9), (31,-9), (33,-9), (-33,-11), (-31,-11), (-29,-11), (-27,-11), (-3,-11), (-1,-11), (1,-11), (3,-11), (5,-11), (7,-11), (9,-11), (11,-11), (29,-11), (31,-11), (-33,-13), (-31,-13), (-29,-13), (-27,-13), (-1,-13), (3,-13), (5,-13), (7,-13), (9,-13), (11,-13), (31,-13), (-31,-15), (-29,-15), (-27,-15), (-25,-15), (-1,-15), (3,-15), (5,-15), (7,-15), (9,-15), (11,-15), (-31,-17), (-29,-17), (-27,-17), (-25,-17), (-1,-17), (3,-17), (5,-17), (7,-17), (9,-17), (11,-17), (27,-17), (29,-17), (-29,-19), (-27,-19), (-25,-19), (-23,-19), (-21,-19), (-1,-19), (1,-19), (3,-19), (5,-19), (7,-19), (9,-19), (11,-19), (25,-19), (27,-19), (29,-19), (-27,-21), (-25,-21), (-23,-21), (-21,-21), (1,-21), (3,-21), (5,-21), (7,-21), (11,-21), (25,-21), (27,-21), (-23,-23), (-21,-23), (-19,-23), (-5,-23), (-3,-23), (-1,-23), (1,-23), (3,-23), (5,-23), (7,-23), (9,-23), (11,-23), (13,-23), (19,-23), (21,-23), (23,-23), (25,-23), (27,-23), (-21,-25), (-19,-25), (-3,-25), (-1,-25), (1,-25), (3,-25), (5,-25), (7,-25), (9,-25), (11,-25), (13,-25), (15,-25), (17,-25), (19,-25), (21,-25), (23,-25), (-17,-27), (-11,-27), (-5,-27), (-3,-27), (-1,-27), (1,-27), (3,-27), (5,-27), (7,-27), (9,-27), (11,-27), (13,-27), (15,-27), (17,-27), (19,-27), (21,-27), (-17,-29), (-11,-29), (-9,-29), (-7,-29), (-5,-29), (-3,-29), (-1,-29), (1,-29), (3,-29), (5,-29), (7,-29), (11,-29), (13,-29), (15,-29), (17,-29), (19,-29), (21,-29), (-11,-31), (-9,-31), (-7,-31), (-5,-31), (-3,-31), (-1,-31), (1,-31), (3,-31), (7,-31), (11,-31), (13,-31), (15,-31), (-9,-33), (-7,-33), (-5,-33), (-3,-33), (-1,-33), (1,-33), (3,-33), (5,-33), (7,-33), (9,-33), (-7,-35), (-5,-35), (-3,-35), (-1,-35), (1,-35), (3,-35), (5,-35), (7,-35)]
mclover3_points = [(-5,35), (-3,35), (-11,31), (-5,31), (-21,29), (-19,29), (-13,29), (-3,29), (13,29), (15,29), (17,29), (19,29), (-21,27), (15,27), (17,27), (21,27), (-23,25), (-9,25), (-7,25), (-5,25), (19,25), (21,25), (-27,23), (-25,23), (-23,23), (-9,23), (13,23), (15,23), (21,23), (23,23), (-27,21), (-25,21), (-23,21), (13,21), (15,21), (17,21), (19,21), (21,21), (23,21), (25,21), (-25,19), (-23,19), (9,19), (11,19), (17,19), (21,19), (23,19), (27,19), (-27,17), (-25,17), (-23,17), (-21,17), (-19,17), (-17,17), (-15,17), (9,17), (13,17), (15,17), (17,17), (19,17), (21,17), (23,17), (25,17), (27,17), (-31,15), (-29,15), (-23,15), (-21,15), (-19,15), (-15,15), (9,15), (11,15), (19,15), (21,15), (-33,13), (-31,13), (-29,13), (-25,13), (-21,13), (-19,13), (-17,13), (-15,13), (9,13), (27,13), (29,13), (-33,11), (-29,11), (-27,11), (-25,11), (-23,11), (-19,11), (-17,11), (25,11), (27,11), (29,11), (-33,9), (-29,9), (9,9), (13,9), (15,9), (17,9), (19,9), (21,9), (23,9), (31,9), (33,9), (35,9), (-31,7), (-25,7), (-23,7), (7,7), (9,7), (25,7), (33,7), (35,7), (-35,5), (-31,5), (-27,5), (-25,5), (7,5), (9,5), (15,5), (17,5), (19,5), (21,5), (25,5), (29,5), (31,5), (35,5), (-35,3), (-31,3), (-27,3), (7,3), (9,3), (11,3), (13,3), (17,3), (21,3), (25,3), (27,3), (-27,1), (7,1), (9,1), (15,1), (19,1), (21,1), (23,1), (27,1), (-29,-1), (-27,-1), (-25,-1), (-23,-1), (7,-1), (9,-1), (11,-1), (13,-1), (19,-1), (21,-1), (23,-1), (27,-1), (-29,-3), (-25,-3), (13,-3), (15,-3), (19,-3), (23,-3), (27,-3), (-25,-5), (-23,-5), (-21,-5), (13,-5), (15,-5), (17,-5), (21,-5), (-23,-7), (-21,-7), (-19,-7), (15,-7), (17,-7), (21,-7), (-21,-9), (-19,-9), (15,-9), (17,-9), (21,-9), (-19,-11), (-17,-11), (17,-11), (21,-11), (-19,-13), (-17,-13), (-15,-13), (-13,-13), (-11,-13), (-9,-13), (17,-13), (-21,-15), (-19,-15), (-15,-15), (-13,-15), (-11,-15), (-9,-15), (-7,-15), (-5,-15), (17,-15), (21,-15), (-15,-17), (-13,-17), (-11,-17), (-9,-17), (17,-17), (21,-17), (-13,-19), (-9,-19), (17,-19), (21,-19), (-15,-21), (-9,-21), (19,-21), (-15,-23), (-11,-23), (-13,-25), (-13,-27)]
mjuniper_points = [(-23,-13)]
point_name_pairing = [("venus", venus_points),("mwasabi", mwasabi_points),("azurite", azurite_points),("electra2", electra2_points),("mclover3", mclover3_points),("mjuniper", mjuniper_points)]
# Robot deck setup constants
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
# Place the PCR tubes in this order
well_colors = {
'A1': 'sfGFP',
'A2': 'mRFP1',
'A3': 'mKO2',
'A4': 'Venus',
'A5': 'mKate2_TF',
'A6': 'Azurite',
'A7': 'mCerulean3',
'A8': 'mClover3',
'A9': 'mJuniper',
'A10': 'mTurquoise2',
'A11': 'mBanana',
'A12': 'mPlum',
'B1': 'Electra2',
'B2': 'mWasabi',
'B3': 'mScarlet_I',
'B4': 'mPapaya',
'B5': 'eqFP578',
'B6': 'tdTomato',
'B7': 'DsRed',
'B8': 'mKate2',
'B9': 'EGFP',
'B10': 'mRuby2',
'B11': 'TagBFP',
'B12': 'mChartreuse_TF',
'C1': 'mLychee_TF',
'C2': 'mTagBFP2',
'C3': 'mEGFP',
'C4': 'mNeonGreen',
'C5': 'mAzamiGreen',
'C6': 'mWatermelon',
'C7': 'avGFP',
'C8': 'mCitrine',
'C9': 'mVenus',
'C10': 'mCherry',
'C11': 'mHoneydew',
'C12': 'TagRFP',
'D1': 'mTFP1',
'D2': 'Ultramarine',
'D3': 'ZsGreen1',
'D4': 'mMiCy',
'D5': 'mStayGold2',
'D6': 'PA_GFP'
}
volume_used = {
'venus': 0,
'mwasabi': 0,
'azurite': 0,
'electra2': 0,
'mclover3': 0,
'mjuniper': 0
}
def update_volume_remaining(current_color, quantity_to_aspirate):
rows = string.ascii_uppercase
for well, color in list(well_colors.items()):
if color == current_color:
if (volume_used[current_color] + quantity_to_aspirate) > 250:
# Move to next well horizontally by advancing row letter, keeping column number
row = well[0]
col = well[1:]
# Find next row letter
next_row = rows[rows.index(row) + 1]
next_well = f"{next_row}{col}"
del well_colors[well]
well_colors[next_well] = current_color
volume_used[current_color] = quantity_to_aspirate
else:
volume_used[current_color] += quantity_to_aspirate
break
def run(protocol):
# Load labware, modules and pipettes
protocol.home()
# Tips
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
# Pipettes
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
# Deep Well Plate
temperature_plate = protocol.load_labware('nest_96_wellplate_2ml_deep', 6)
# Agar Plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate')
agar_plate.set_offset(x=0.00, y=0.00, z=Z_VALUE_AGAR)
# Get the top-center of the plate, make sure the plate was calibrated before running this
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
# Helper function (dispensing)
def dispense_and_jog(pipette, volume, location):
assert(isinstance(volume, (int, float)))
# Go above the location
above_location = location.move(types.Point(z=location.point.z + 2))
pipette.move_to(above_location)
# Go downwards and dispense
pipette.dispense(volume, location)
# Go upwards to avoid smearing
pipette.move_to(above_location)
# Helper function (color location)
def location_of_color(color_string):
for well,color in well_colors.items():
if color.lower() == color_string.lower():
return temperature_plate[well]
raise ValueError(f"No well found with color {color_string}")
# Print pattern by iterating over lists
for i, (current_color, point_list) in enumerate(point_name_pairing):
# Skip the rest of the loop if the list is empty
if not point_list:
continue
# Get the tip for this run, set the bacteria color, and the aspirate bacteria of choice
pipette_20ul.pick_up_tip()
max_aspirate = int(18 // POINT_SIZE) * POINT_SIZE
quantity_to_aspirate = min(len(point_list)*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Iterate over the current points list and dispense them, refilling along the way
for i in range(len(point_list)):
x, y = point_list[i]
adjusted_location = center_location.move(types.Point(x, y))
dispense_and_jog(pipette_20ul, POINT_SIZE, adjusted_location)
if pipette_20ul.current_volume == 0 and len(point_list[i+1:]) > 0:
quantity_to_aspirate = min(len(point_list[i:])*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Drop tip between each color
pipette_20ul.drop_tip()
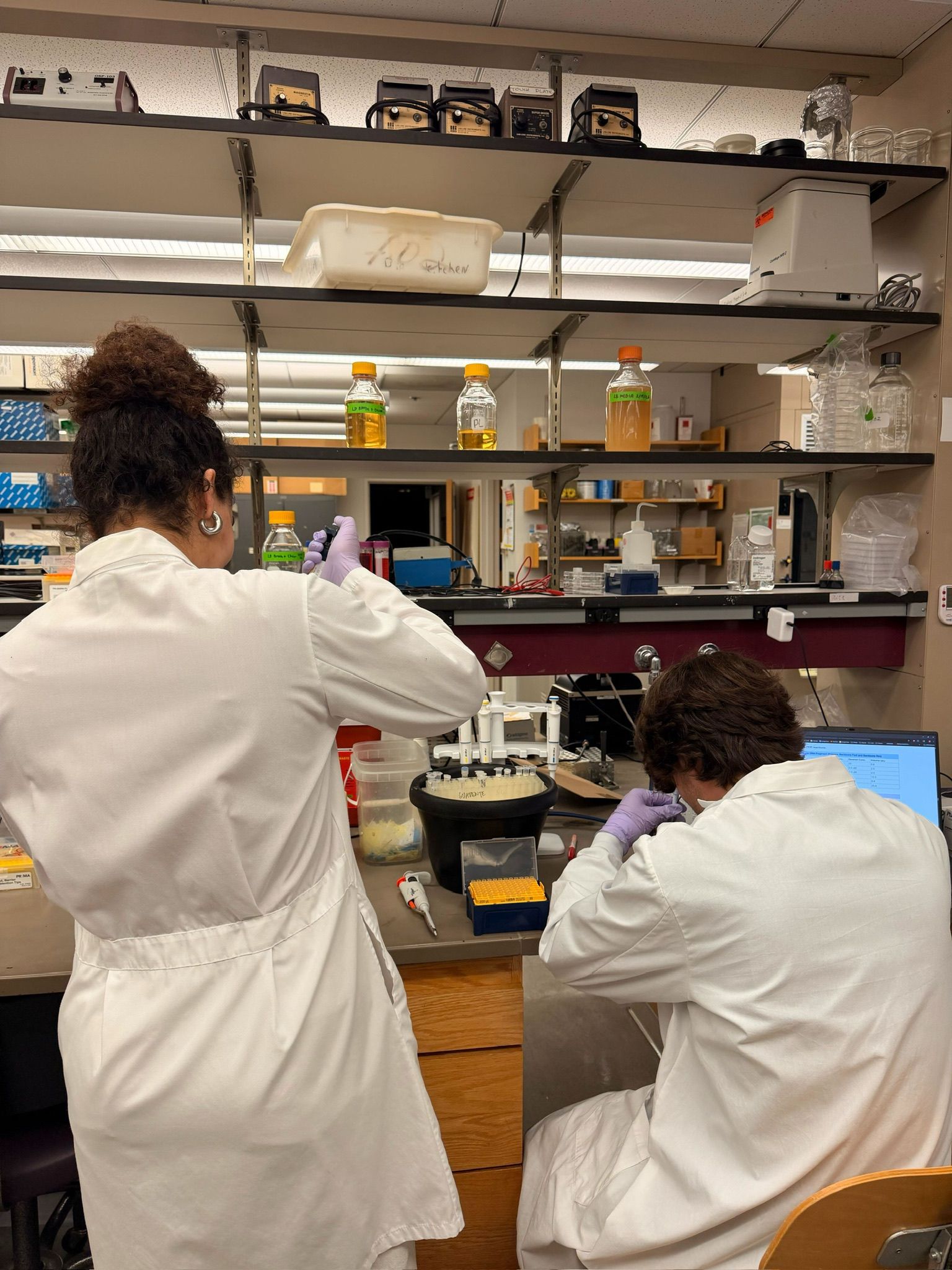
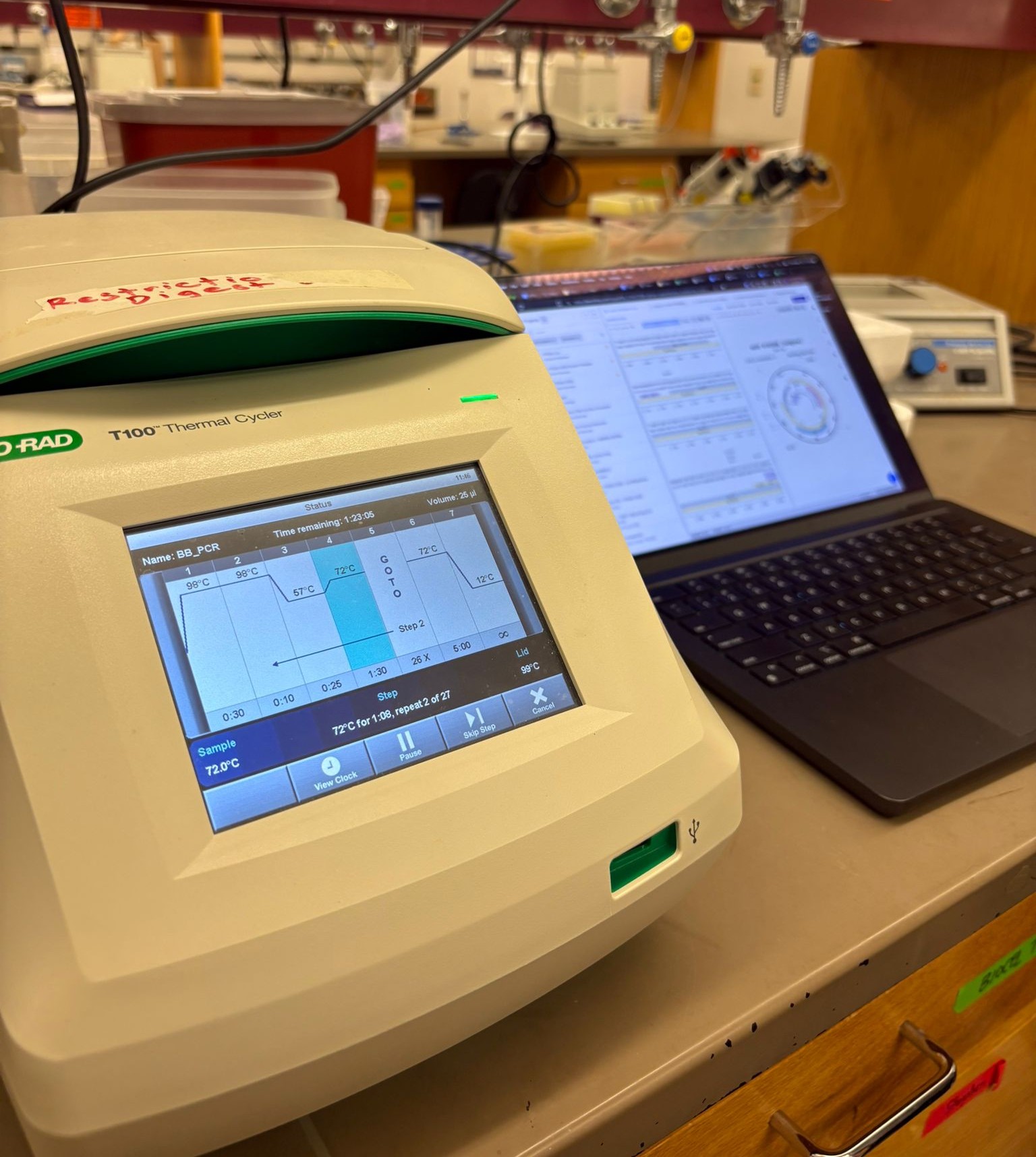
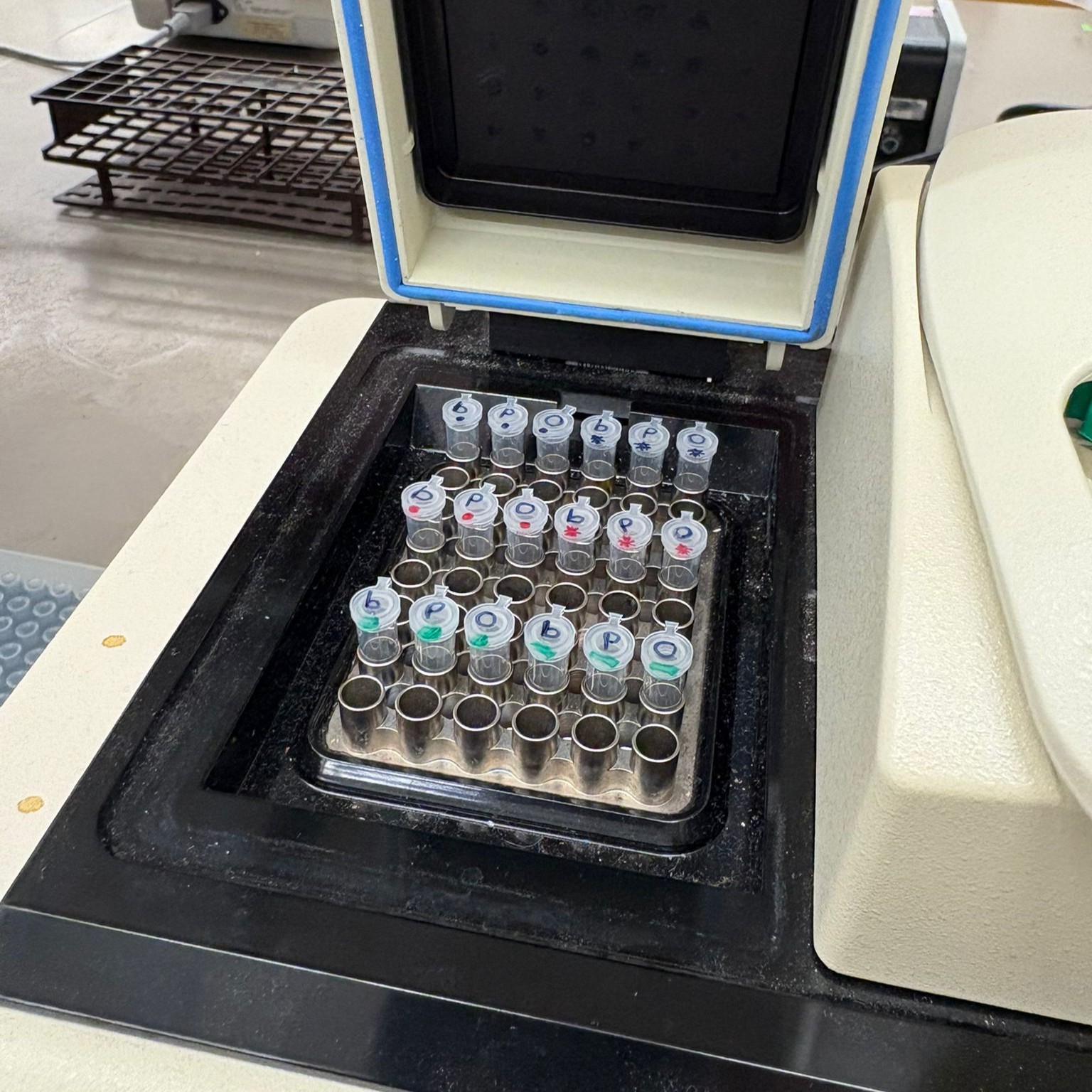
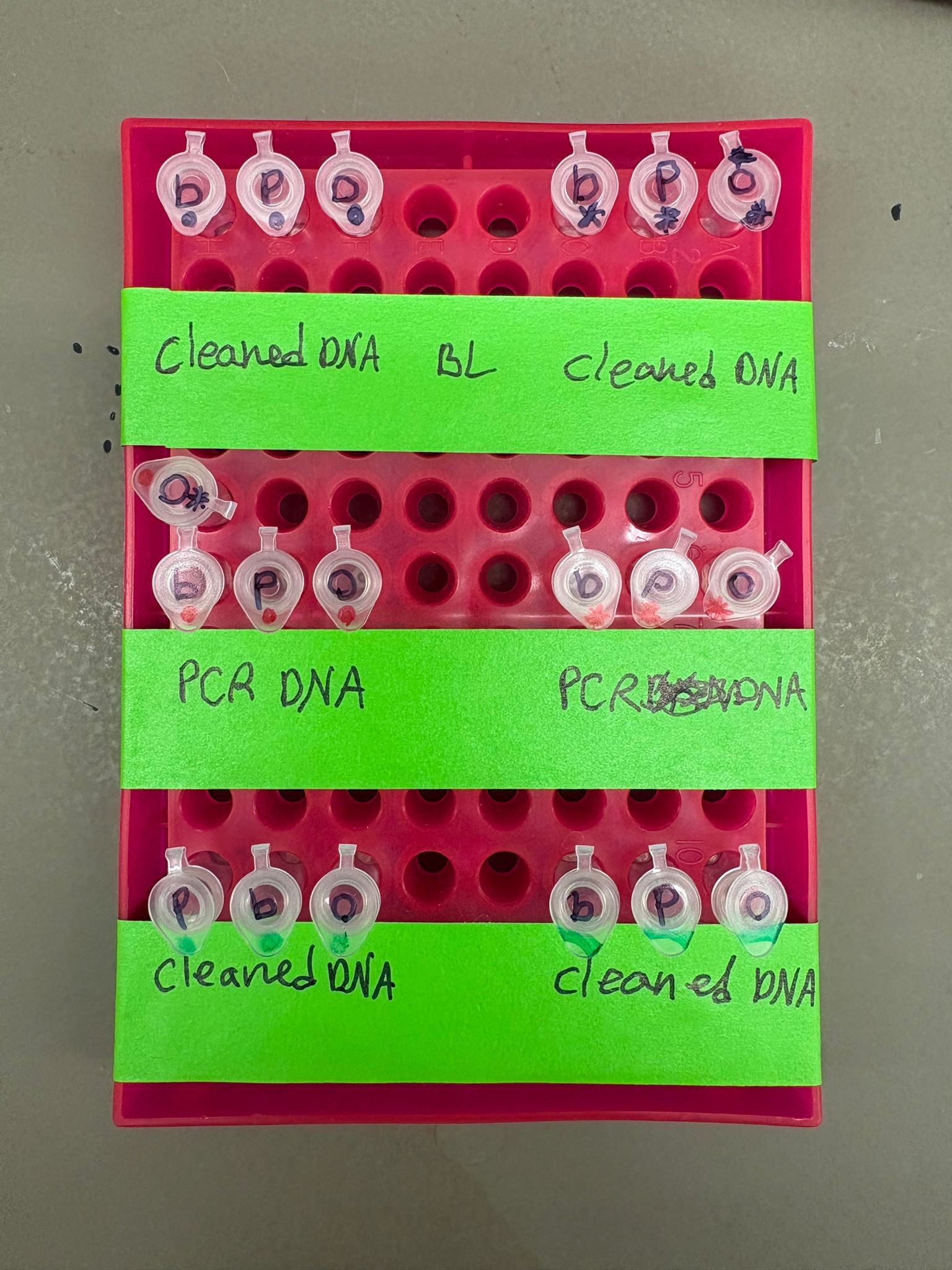
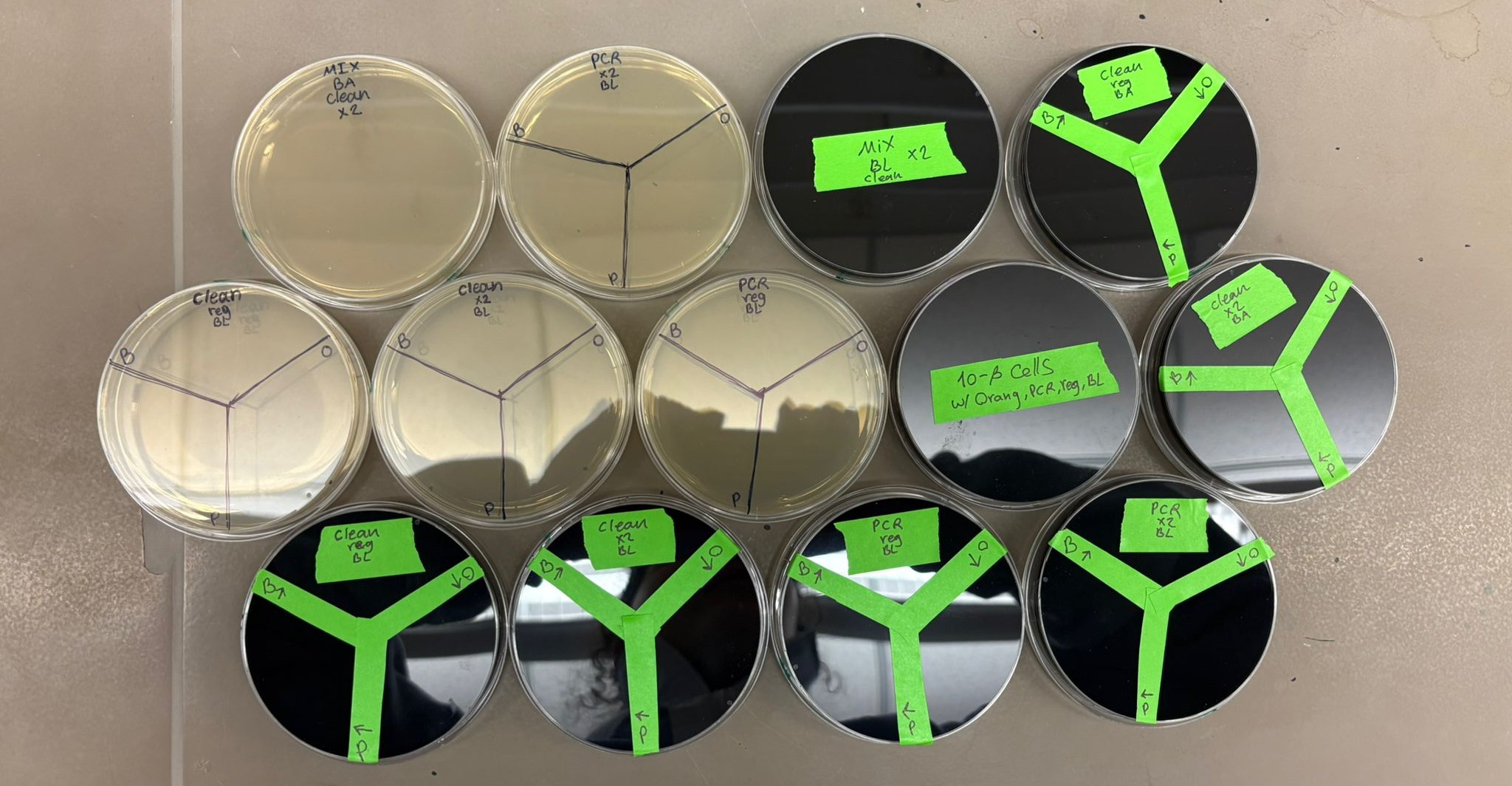
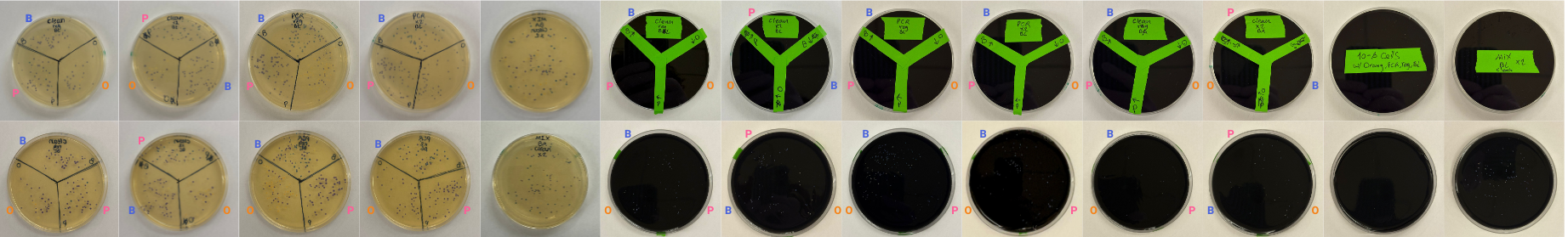
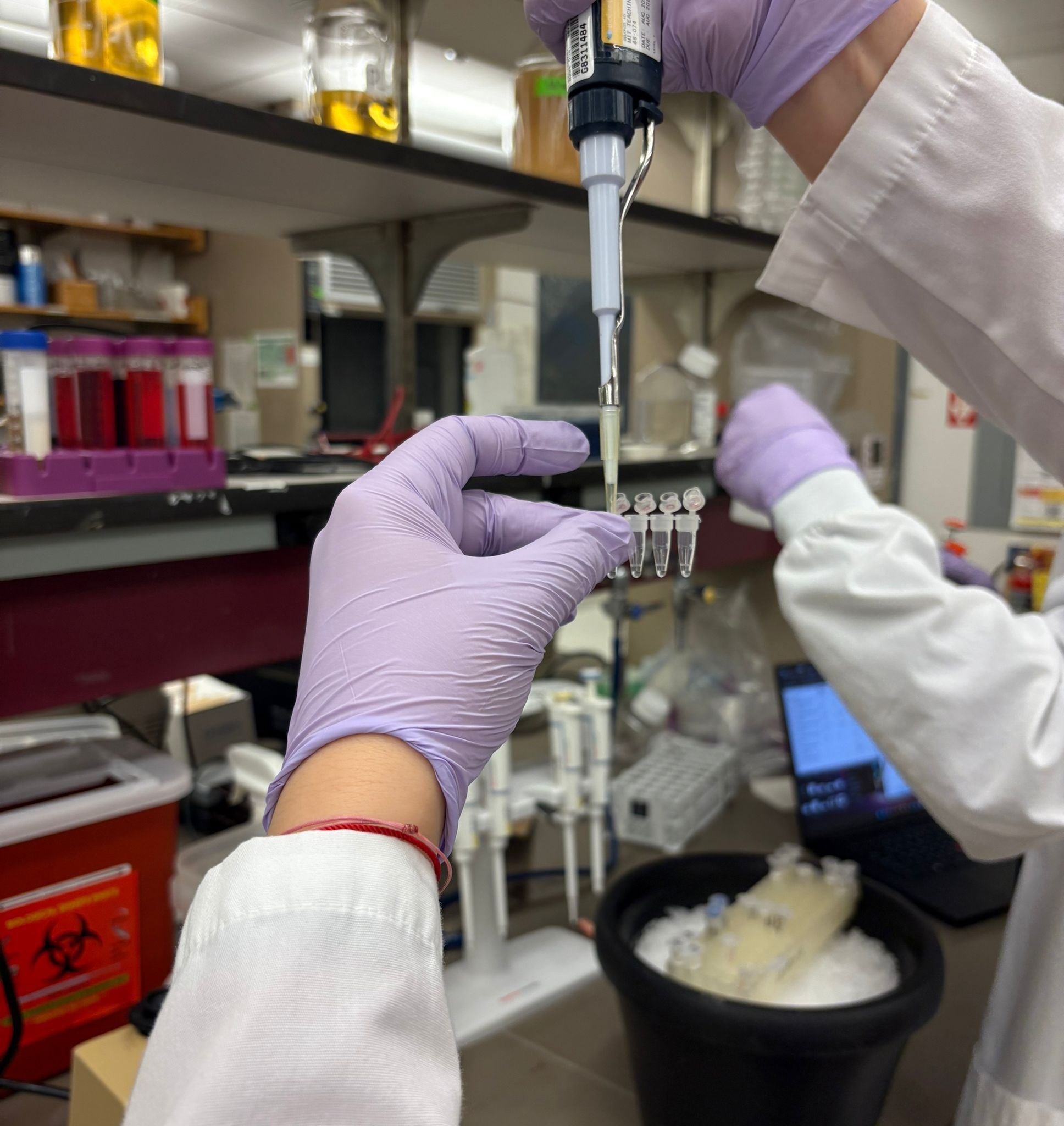
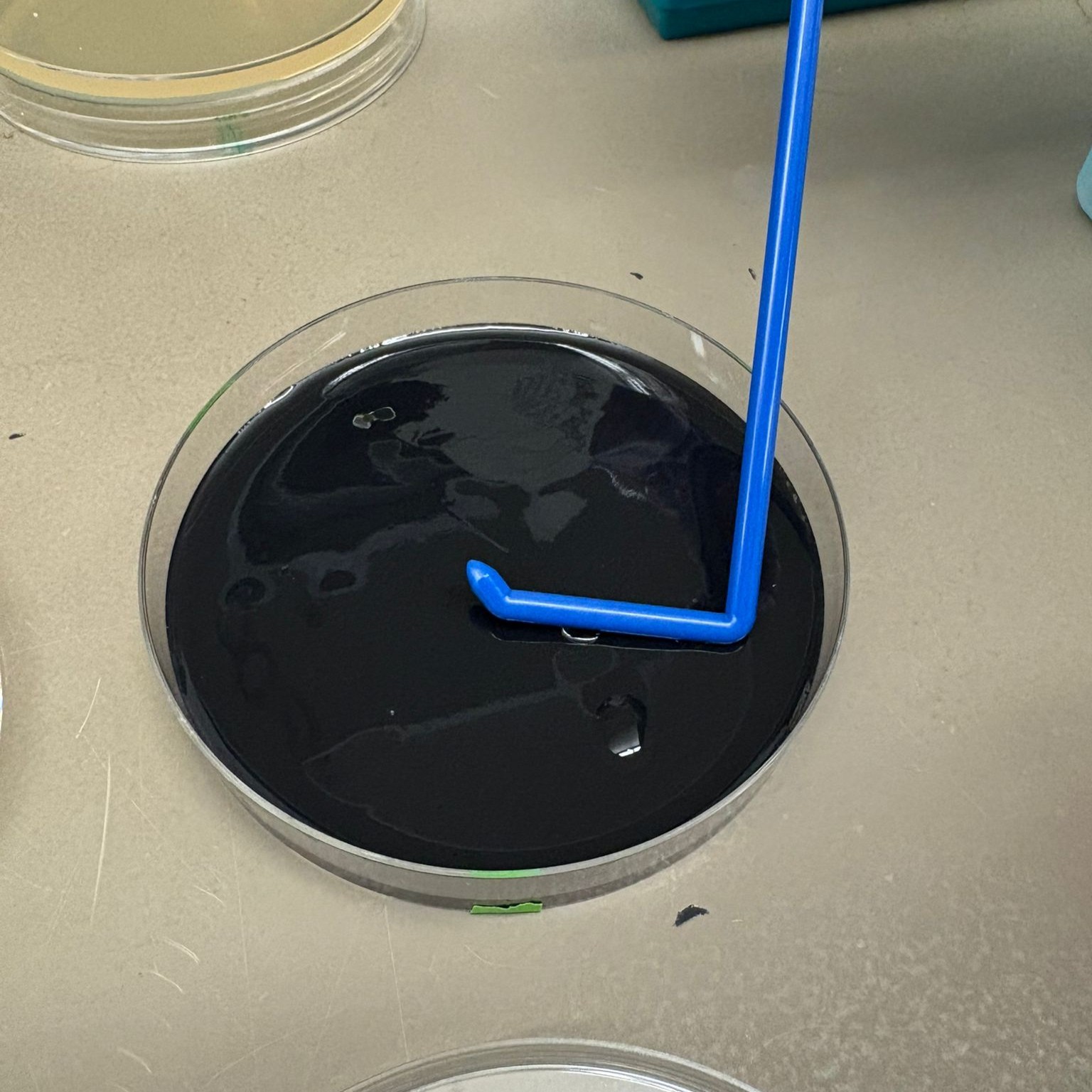
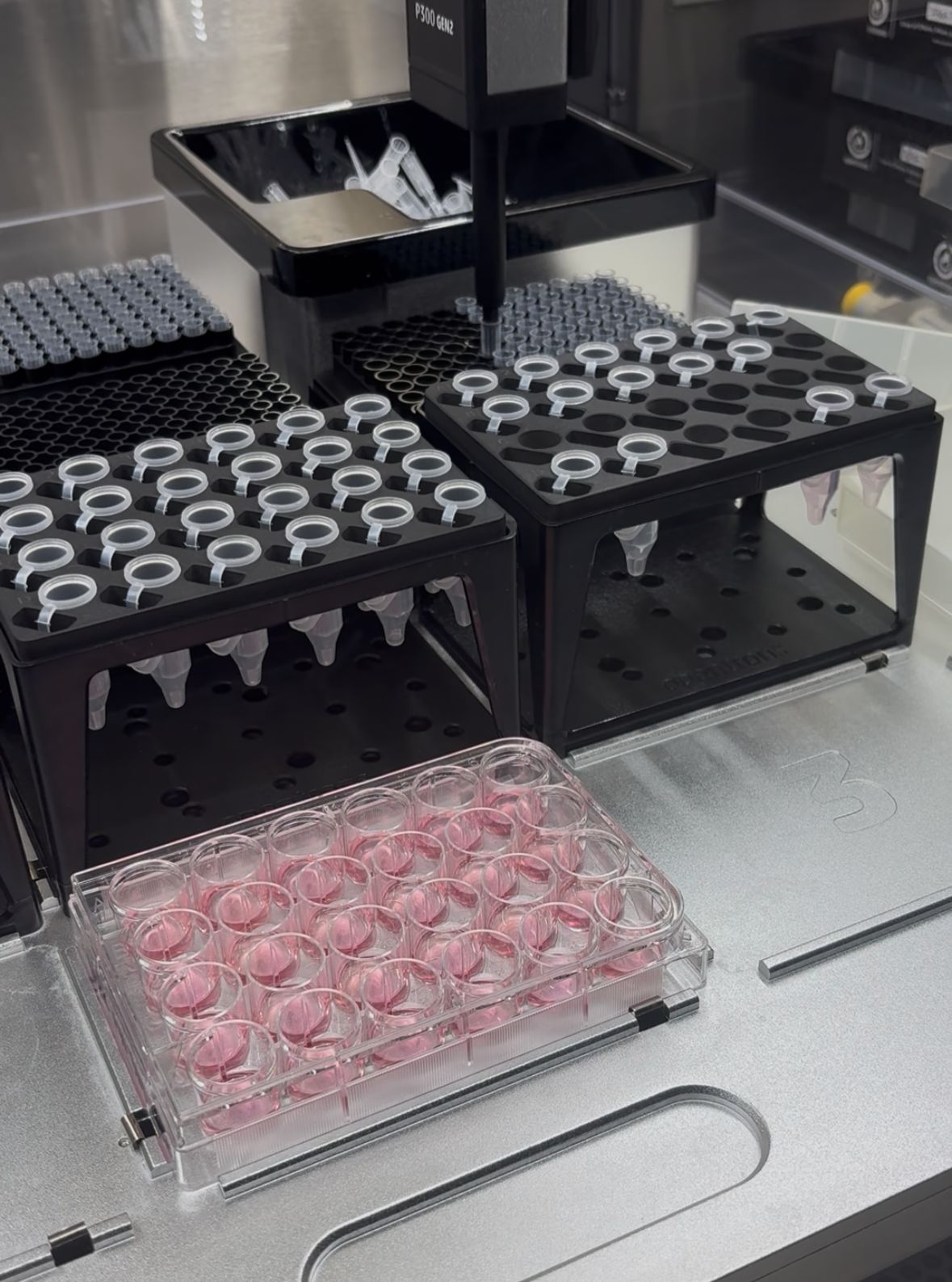
In the lab, we uploaded the design protocol to the Opentrons, loaded the trays with the engineered E. coli, the black (charcoal) agar plate, pipette tips, and a “trash” bin. It was fascinating to watch the machine work. I was especially happy to discover that the Opentrons OT-2 operates using a 3D gantry system - very similar to 3D printers and CNC cutters - which made it very intuitive for me to use (and fun to watch).
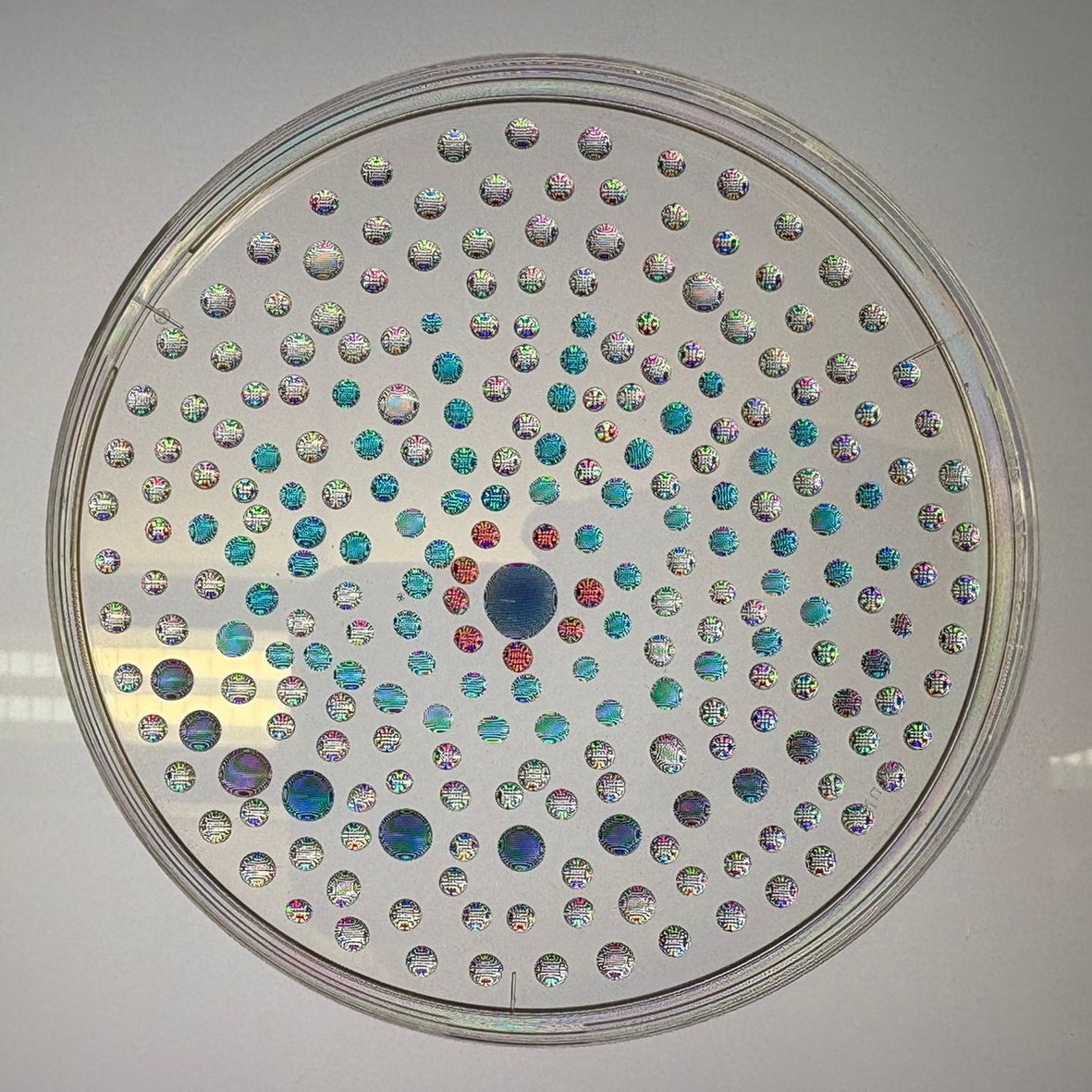
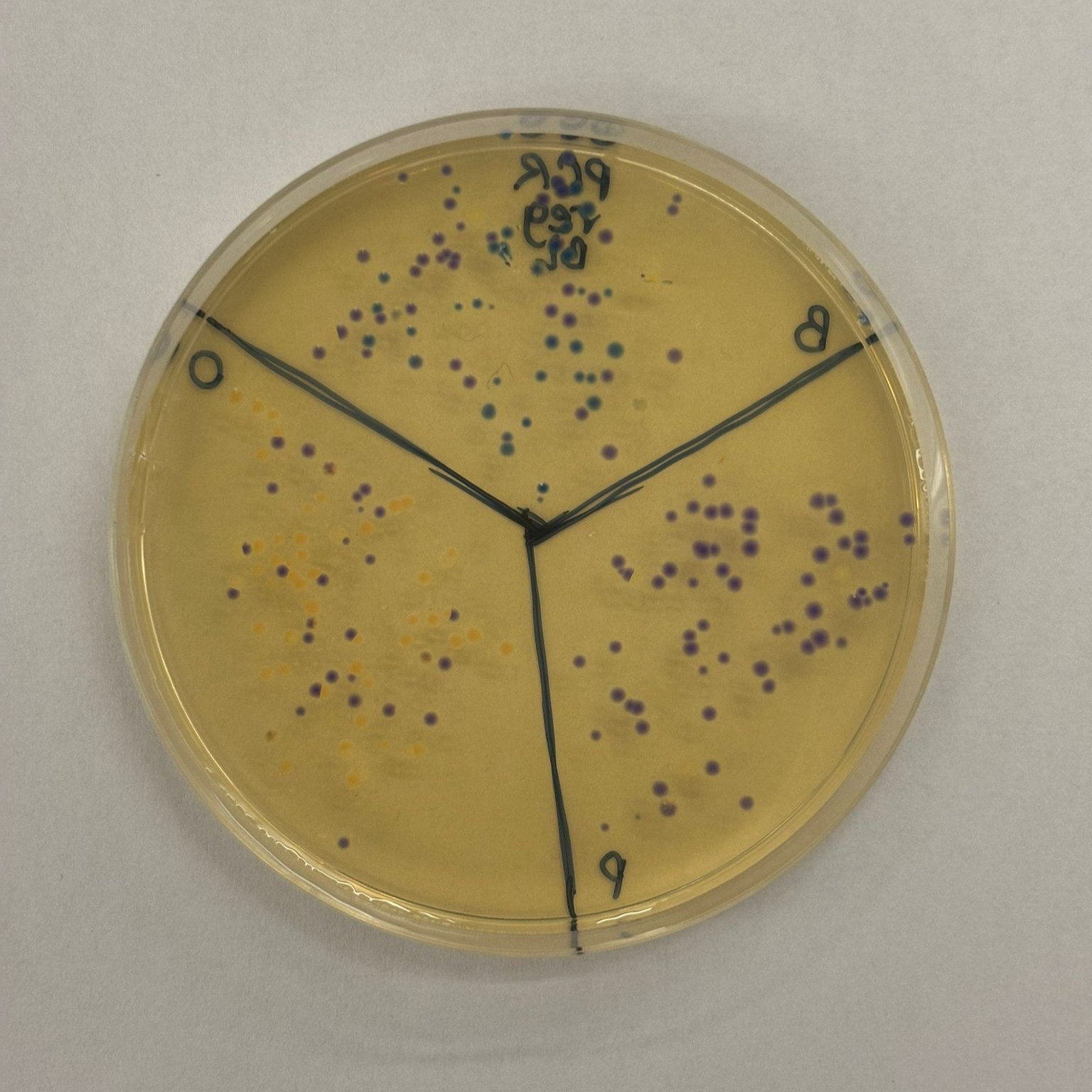
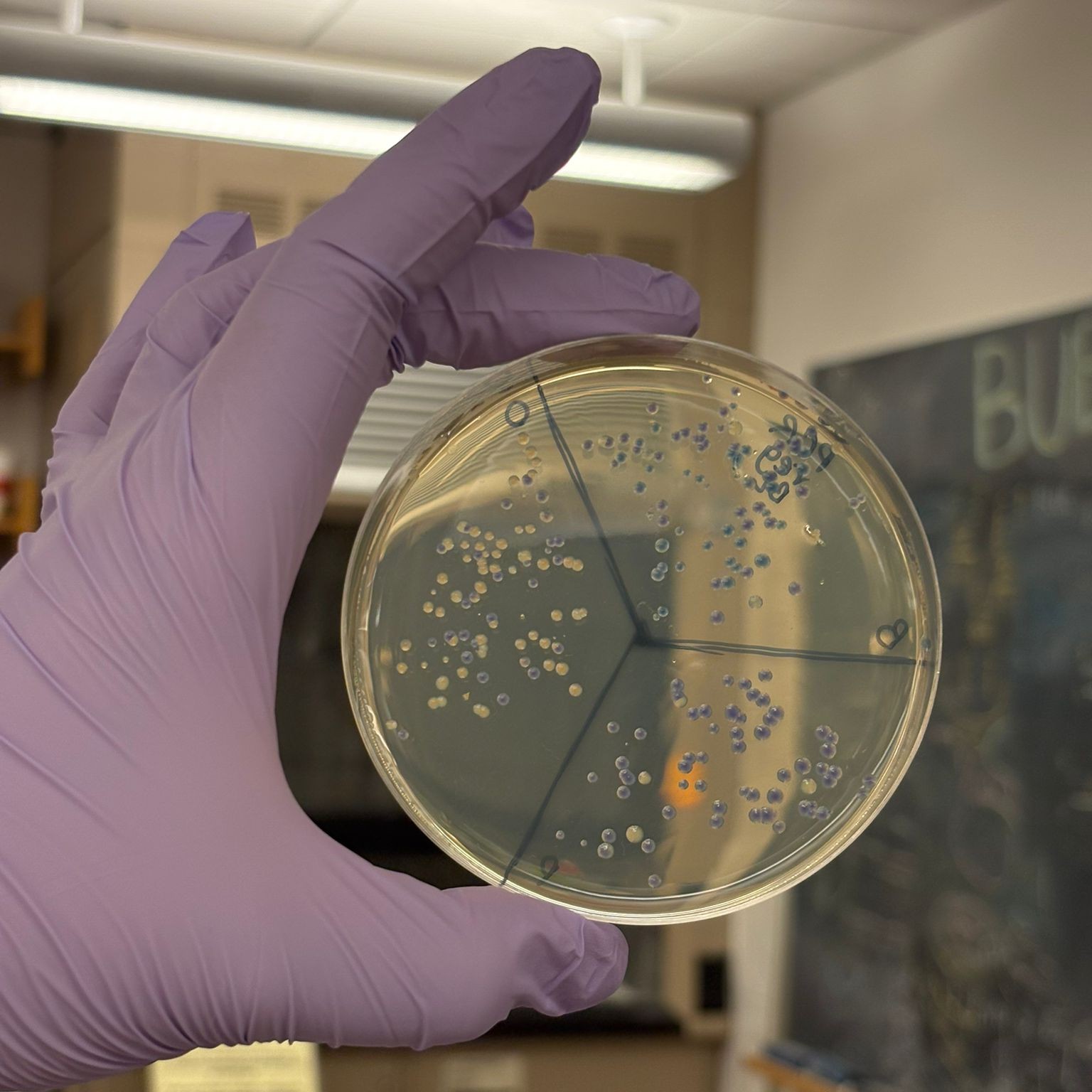
After “the print” was finished, we placed the agar plates in the incubator overnight and photographed them under UV light the following day.
On Saturday, I had the honor of visiting the Ginkgo Bioworks labs in Boston, where I saw how the Nebula (the RAC system at Ginkgo Bioworks) works, and even loaded another design of mine onto it. This time, I used a rectangular composition and painted the Israeli Sabra cactus - or in modern Hebrew, Tzabar - which reminds me of sweet childhood memories and my lovely country.
This definitely won’t be my last experiment with bio art - I’m already looking forward to bringing lab automation into my final project.
*Also, this cool idea reminded me of a game I used to play whenever I was bored. I won’t tell you anything about it —-except that it involves koalas. Here is the link to the game