Week 2 HW: DNA - read, write and edit
Part 1- Benchling & In-silico Gel Art
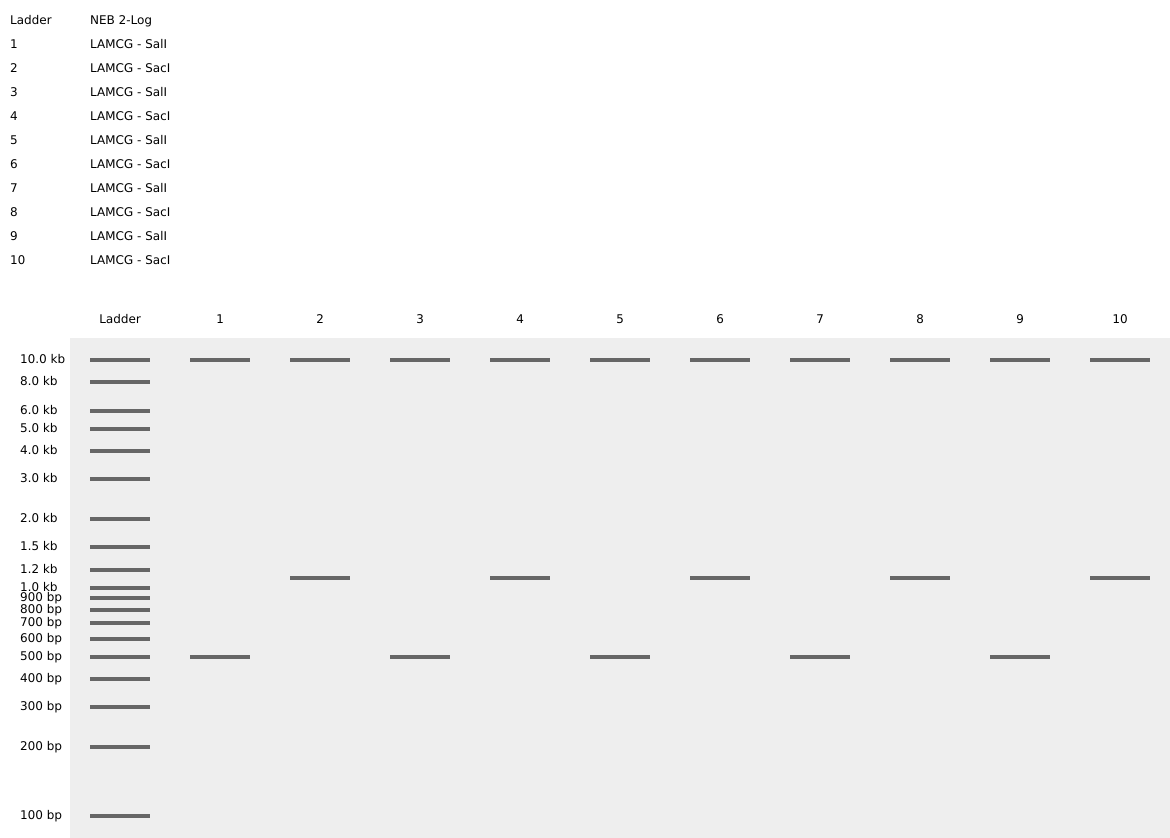
Simulating Restriction Enzyme Digestion with the following Enzymes:
.EcoRI
.HindIII
.BamHI
.KpnI
.EcoRV
.SacI
.SalI
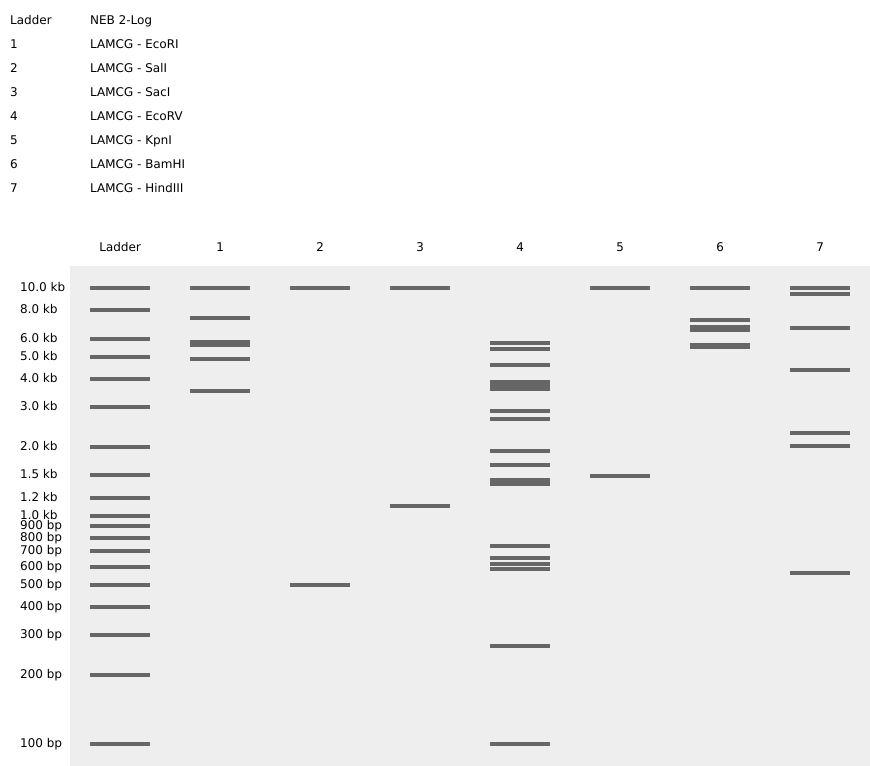


I had created a simple pattern in the style of Paul Vanouse’s Latent Figure Protocol artworks. The bands are arranged alternatively creating a horizontal alternative pattern.

Part 3: DNA Design Challenge
3.1. Choose your protein The protein I had selected is Myosin, which is a motor protein responsible for muscle contraction. It binds to the actin filaments and uses ATP to generate force. This force pulls the actin filaments inwards causes the muscle fibres to shorten and contract. I am particularly interested in understanding how myosin behaves in microgravity conditions, where mechanical loading is absent and muscle atrophy occurs rapidly. I collected the protein from human at Uniprot- Q9Y2K3
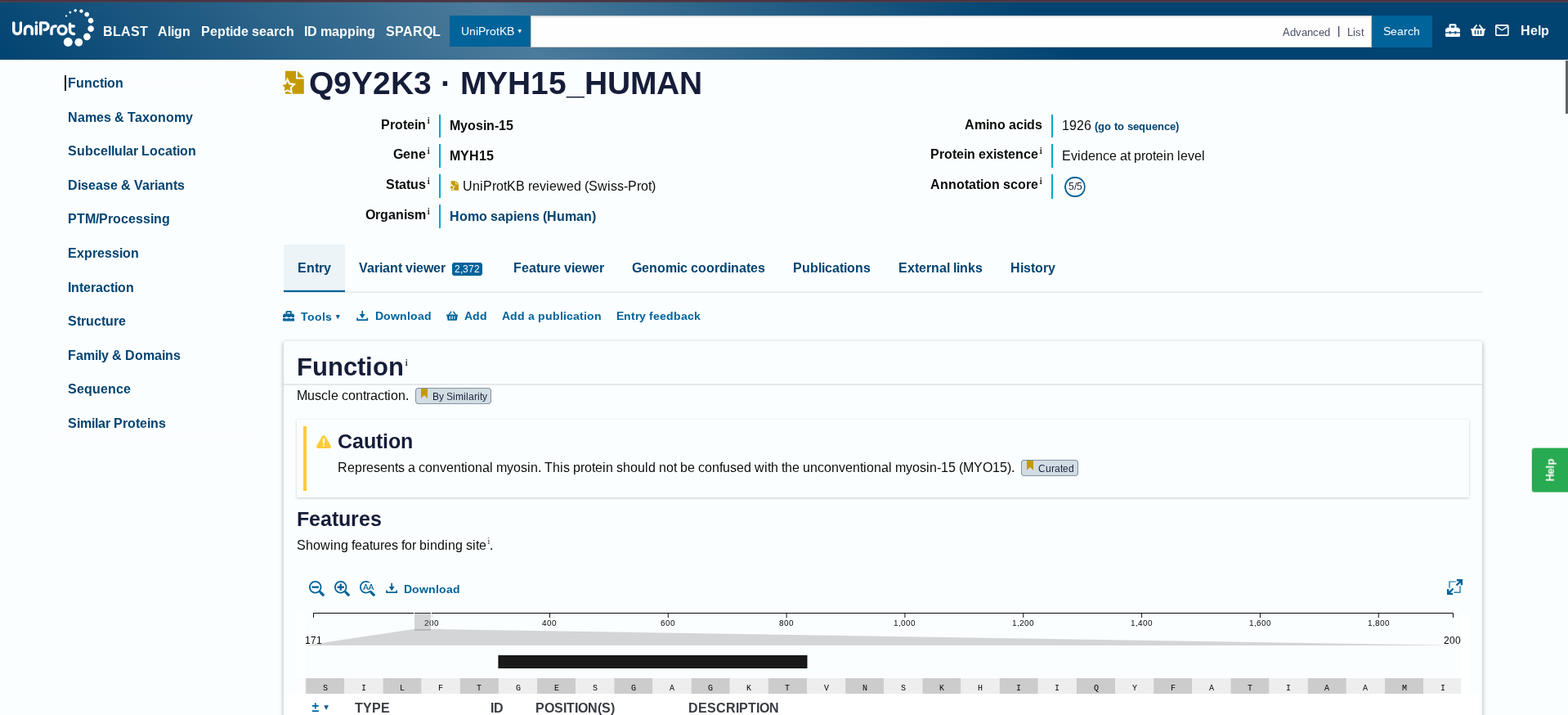
 Below is the protein sequence of Myosin-15 extracted from Homo sapien
Below is the protein sequence of Myosin-15 extracted from Homo sapien
sp|Q9Y2K3|MYH15_HUMAN Myosin-15 OS=Homo sapiens OX=9606 GN=MYH15 PE=1 SV=6 MDLSDLGEAAAFLRRSEAELLLLQATALDGKKKCWIPDGENAYIEAEVKGSEDDGTVIVE TADGESLSIKEDKIQQMNPPEFEMIEDMAMLTHLNEASVLHTLKRRYGQWMIYTYSGLFC VTINPYKWLPVYQKEVMAAYKGKRRSEAPPHIFAVANNAFQDMLHNRENQSILFTGESGA GKTVNSKHIIQYFATIAAMIESRKKQGALEDQIMQANTILEAFGNAKTLRNDNSSRFGKF
IRMHFGARGMLSSVDIDIYLLEKSRVIFQQAGERNYHIFYQILSGQKELHDLLLVSANPS DFHFCSCGAVTVESLDDAEELLATEQAMDILGFLPDEKYGCYKLTGAIMHFGNMKFKQKP REEQLEADGTENADKAAFLMGINSSELVKCLIHPRIKVGNEYVTRGQTIEQVTCAVGALS KSMYERMFKWLVARINRALDAKLSRQFFIGILDITGFEILEYNSLEQLCINFTNEKLQQF FNWHMFVLEQEEYKKESIEWVSIGFGLDLQACIDLIEKPMGILSILEEECMFPKATDLTF KTKLFDNHFGKSVHLQKPKPDKKKFEAHFELVHYAGVVPYNISGWLEKNKDLLNETVVAV FQKSSNRLLASLFENYMSTDSAIPFGEKKRKKGASFQTVASLHKENLNKLMTNLKSTAPH FVRCINPNVNKIPGILDPYLVLQQLRCNGVLEGTRICREGFPNRLQYADFKQRYCILNPR TFPKSKFVSSRKAAEELLGSLEIDHTQYRFGITKVFFKAGFLGQLEAIRDERLSKVFTLF QARAQGKLMRIKFQKILEERDALILIQWNIRAFMAVKNWPWMRLFFKIKPLVKSSEVGEE VAGLKEECAQLQKALEKSEFQREELKAKQVSLTQEKNDLILQLQAEQETLANVEEQCEWL IKSKIQLEARVKELSERVEEEEEINSELTARGRKLEDECFELKKEIDDLETMLVKSEKEK RTTEHKVKNLTEEVEFLNEDISKLNRAAKVVQEAHQQTLDDLHMEEEKLSSLSKANLKLE QQVDELEGALEQERKARMNCERELHKLEGNLKLNRESMENLESSQRHLAEELRKKELELS QMNSKVENEKGLVAQLQKTVKELQTQIKDLKEKLEAERTTRAKMERERADLTQDLADLNE RLEEVGGSSLAQLEITKKQETKFQKLHRDMEEATLHFETTSASLKKRHADSLAELEGQVE NLQQVKQKLEKDKSDLQLEVDDLLTRVEQMTRAKANAEKLCTLYEERLHEATAKLDKVTQ LANDLAAQKTKLWSESGEFLRRLEEKEALINQLSREKSNFTRQIEDLRGQLEKETKSQSA LAHALQKAQRDCDLLREQYEEEQEVKAELHRTLSKVNAEMVQWRMKYENNVIQRTEDLED AKKELAIRLQEAAEAMGVANARNASLERARHQLQLELGDALSDLGKVRSAAARLDQKQLQ SGKALADWKQKHEESQALLDASQKEVQALSTELLKLKNTYEESIVGQETLRRENKNLQEE ISNLTNQVREGTKNLTEMEKVKKLIEEEKTEVQVTLEETEGALERNESKILHFQLELLEA KAELERKLSEKDEEIENFRRKQQCTIDSLQSSLDSEAKSRIEVTRLKKKMEEDLNEMELQ LSCANRQVSEATKSLGQLQIQIKDLQMQLDDSTQLNSDLKEQVAVAERRNSLLQSELEDL RSLQEQTERGRRLSEEELLEATERINLFYTQNTSLLSQKKKLEADVARMQKEAEEVVQEC QNAEEKAKKAAIEAANLSEELKKKQDTIAHLERTRENMEQTITDLQKRLAEAEQMALMGS RKQIQKLESRVRELEGELEGEIRRSAEAQRGARRLERCIKELTYQAEEDKKNLSRMQTQM DKLQLKVQNYKQQVEVAETQANQYLSKYKKQQHELNEVKERAEVAESQVNKLKIKAREFG KKVQEE
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
The Central Dogma gives us the framework to work backwards from a given protein sequence and infer the DNA sequence that the protein is derived from. Reverse Translate results Results for 1926 residue sequence “sp|Q9Y2K3|MYH15_HUMAN Myosin-15 OS=Homo sapiens OX=9606 GN=MYH15 PE=1 SV=6” starting “MDLSDLGEAA”
reverse translation of sp|Q9Y2K3|MYH15_HUMAN Myosin-15 OS=Homo sapiens OX=9606 GN=MYH15 PE=1 SV=6 to a 5778 base sequence of most likely codons. atggatctgagcgatctgggcgaagcggcggcgtttctgcgccgcagcgaagcggaactg ctgctgctgcaggcgaccgcgctggatggcaaaaaaaaatgctggattccggatggcgaa aacgcgtatattgaagcggaagtgaaaggcagcgaagatgatggcaccgtgattgtggaa accgcggatggcgaaagcctgagcattaaagaagataaaattcagcagatgaacccgccg gaatttgaaatgattgaagatatggcgatgctgacccatctgaacgaagcgagcgtgctg cataccctgaaacgccgctatggccagtggatgatttatacctatagcggcctgttttgc gtgaccattaacccgtataaatggctgccggtgtatcagaaagaagtgatggcggcgtat aaaggcaaacgccgcagcgaagcgccgccgcatatttttgcggtggcgaacaacgcgttt caggatatgctgcataaccgcgaaaaccagagcattctgtttaccggcgaaagcggcgcg ggcaaaaccgtgaacagcaaacatattattcagtattttgcgaccattgcggcgatgatt gaaagccgcaaaaaacagggcgcgctggaagatcagattatgcaggcgaacaccattctg gaagcgtttggcaacgcgaaaaccctgcgcaacgataacagcagccgctttggcaaattt attcgcatgcattttggcgcgcgcggcatgctgagcagcgtggatattgatatttatctg ctggaaaaaagccgcgtgatttttcagcaggcgggcgaacgcaactatcatattttttat cagattctgagcggccagaaagaactgcatgatctgctgctggtgagcgcgaacccgagc gattttcatttttgcagctgcggcgcggtgaccgtggaaagcctggatgatgcggaagaa ctgctggcgaccgaacaggcgatggatattctgggctttctgccggatgaaaaatatggc tgctataaactgaccggcgcgattatgcattttggcaacatgaaatttaaacagaaaccg cgcgaagaacagctggaagcggatggcaccgaaaacgcggataaagcggcgtttctgatg ggcattaacagcagcgaactggtgaaatgcctgattcatccgcgcattaaagtgggcaac gaatatgtgacccgcggccagaccattgaacaggtgacctgcgcggtgggcgcgctgagc aaaagcatgtatgaacgcatgtttaaatggctggtggcgcgcattaaccgcgcgctggat gcgaaactgagccgccagttttttattggcattctggatattaccggctttgaaattctg gaatataacagcctggaacagctgtgcattaactttaccaacgaaaaactgcagcagttt tttaactggcatatgtttgtgctggaacaggaagaatataaaaaagaaagcattgaatgg gtgagcattggctttggcctggatctgcaggcgtgcattgatctgattgaaaaaccgatg ggcattctgagcattctggaagaagaatgcatgtttccgaaagcgaccgatctgaccttt aaaaccaaactgtttgataaccattttggcaaaagcgtgcatctgcagaaaccgaaaccg gataaaaaaaaatttgaagcgcattttgaactggtgcattatgcgggcgtggtgccgtat aacattagcggctggctggaaaaaaacaaagatctgctgaacgaaaccgtggtggcggtg tttcagaaaagcagcaaccgcctgctggcgagcctgtttgaaaactatatgagcaccgat agcgcgattccgtttggcgaaaaaaaacgcaaaaaaggcgcgagctttcagaccgtggcg agcctgcataaagaaaacctgaacaaactgatgaccaacctgaaaagcaccgcgccgcat tttgtgcgctgcattaacccgaacgtgaacaaaattccgggcattctggatccgtatctg gtgctgcagcagctgcgctgcaacggcgtgctggaaggcacccgcatttgccgcgaaggc tttccgaaccgcctgcagtatgcggattttaaacagcgctattgcattctgaacccgcgc acctttccgaaaagcaaatttgtgagcagccgcaaagcggcggaagaactgctgggcagc ctggaaattgatcatacccagtatcgctttggcattaccaaagtgttttttaaagcgggc tttctgggccagctggaagcgattcgcgatgaacgcctgagcaaagtgtttaccctgttt caggcgcgcgcgcagggcaaactgatgcgcattaaatttcagaaaattctggaagaacgc gatgcgctgattctgattcagtggaacattcgcgcgtttatggcggtgaaaaactggccg tggatgcgcctgttttttaaaattaaaccgctggtgaaaagcagcgaagtgggcgaagaa gtggcgggcctgaaagaagaatgcgcgcagctgcagaaagcgctggaaaaaagcgaattt cagcgcgaagaactgaaagcgaaacaggtgagcctgacccaggaaaaaaacgatctgatt ctgcagctgcaggcggaacaggaaaccctggcgaacgtggaagaacagtgcgaatggctg attaaaagcaaaattcagctggaagcgcgcgtgaaagaactgagcgaacgcgtggaagaa gaagaagaaattaacagcgaactgaccgcgcgcggccgcaaactggaagatgaatgcttt gaactgaaaaaagaaattgatgatctggaaaccatgctggtgaaaagcgaaaaagaaaaa cgcaccaccgaacataaagtgaaaaacctgaccgaagaagtggaatttctgaacgaagat attagcaaactgaaccgcgcggcgaaagtggtgcaggaagcgcatcagcagaccctggat gatctgcatatggaagaagaaaaactgagcagcctgagcaaagcgaacctgaaactggaa cagcaggtggatgaactggaaggcgcgctggaacaggaacgcaaagcgcgcatgaactgc gaacgcgaactgcataaactggaaggcaacctgaaactgaaccgcgaaagcatggaaaac ctggaaagcagccagcgccatctggcggaagaactgcgcaaaaaagaactggaactgagc cagatgaacagcaaagtggaaaacgaaaaaggcctggtggcgcagctgcagaaaaccgtg aaagaactgcagacccagattaaagatctgaaagaaaaactggaagcggaacgcaccacc cgcgcgaaaatggaacgcgaacgcgcggatctgacccaggatctggcggatctgaacgaa cgcctggaagaagtgggcggcagcagcctggcgcagctggaaattaccaaaaaacaggaa accaaatttcagaaactgcatcgcgatatggaagaagcgaccctgcattttgaaaccacc agcgcgagcctgaaaaaacgccatgcggatagcctggcggaactggaaggccaggtggaa aacctgcagcaggtgaaacagaaactggaaaaagataaaagcgatctgcagctggaagtg gatgatctgctgacccgcgtggaacagatgacccgcgcgaaagcgaacgcggaaaaactg tgcaccctgtatgaagaacgcctgcatgaagcgaccgcgaaactggataaagtgacccag ctggcgaacgatctggcggcgcagaaaaccaaactgtggagcgaaagcggcgaatttctg cgccgcctggaagaaaaagaagcgctgattaaccagctgagccgcgaaaaaagcaacttt acccgccagattgaagatctgcgcggccagctggaaaaagaaaccaaaagccagagcgcg ctggcgcatgcgctgcagaaagcgcagcgcgattgcgatctgctgcgcgaacagtatgaa gaagaacaggaagtgaaagcggaactgcatcgcaccctgagcaaagtgaacgcggaaatg gtgcagtggcgcatgaaatatgaaaacaacgtgattcagcgcaccgaagatctggaagat gcgaaaaaagaactggcgattcgcctgcaggaagcggcggaagcgatgggcgtggcgaac gcgcgcaacgcgagcctggaacgcgcgcgccatcagctgcagctggaactgggcgatgcg ctgagcgatctgggcaaagtgcgcagcgcggcggcgcgcctggatcagaaacagctgcag agcggcaaagcgctggcggattggaaacagaaacatgaagaaagccaggcgctgctggat gcgagccagaaagaagtgcaggcgctgagcaccgaactgctgaaactgaaaaacacctat gaagaaagcattgtgggccaggaaaccctgcgccgcgaaaacaaaaacctgcaggaagaa attagcaacctgaccaaccaggtgcgcgaaggcaccaaaaacctgaccgaaatggaaaaa gtgaaaaaactgattgaagaagaaaaaaccgaagtgcaggtgaccctggaagaaaccgaa ggcgcgctggaacgcaacgaaagcaaaattctgcattttcagctggaactgctggaagcg aaagcggaactggaacgcaaactgagcgaaaaagatgaagaaattgaaaactttcgccgc aaacagcagtgcaccattgatagcctgcagagcagcctggatagcgaagcgaaaagccgc attgaagtgacccgcctgaaaaaaaaaatggaagaagatctgaacgaaatggaactgcag ctgagctgcgcgaaccgccaggtgagcgaagcgaccaaaagcctgggccagctgcagatt cagattaaagatctgcagatgcagctggatgatagcacccagctgaacagcgatctgaaa gaacaggtggcggtggcggaacgccgcaacagcctgctgcagagcgaactggaagatctg cgcagcctgcaggaacagaccgaacgcggccgccgcctgagcgaagaagaactgctggaa gcgaccgaacgcattaacctgttttatacccagaacaccagcctgctgagccagaaaaaa aaactggaagcggatgtggcgcgcatgcagaaagaagcggaagaagtggtgcaggaatgc cagaacgcggaagaaaaagcgaaaaaagcggcgattgaagcggcgaacctgagcgaagaa ctgaaaaaaaaacaggataccattgcgcatctggaacgcacccgcgaaaacatggaacag accattaccgatctgcagaaacgcctggcggaagcggaacagatggcgctgatgggcagc cgcaaacagattcagaaactggaaagccgcgtgcgcgaactggaaggcgaactggaaggc gaaattcgccgcagcgcggaagcgcagcgcggcgcgcgccgcctggaacgctgcattaaa gaactgacctatcaggcggaagaagataaaaaaaacctgagccgcatgcagacccagatg gataaactgcagctgaaagtgcagaactataaacagcaggtggaagtggcggaaacccag gcgaaccagtatctgagcaaatataaaaaacagcagcatgaactgaacgaagtgaaagaa cgcgcggaagtggcggaaagccaggtgaacaaactgaaaattaaagcgcgcgaatttggc aaaaaagtgcaggaagaa
I used https://www.bioinformatics.org/sms2/rev_trans.html to reverse translate the protein sequence back to the nucleotides.
3.3. Codon optimization Codon optimization is the process of changing the DNA sequence of a gene so that it is expressed more efficiently in a specific organism without changing the protein it produces. I used https://www.idtdna.com/CodonOpt to do codon optimization. I used the reverse translated sequence from 3.2 step, because I was interested in that sequence. I want to continue the process and went for codon optimizaton of the reverse translated sequence.
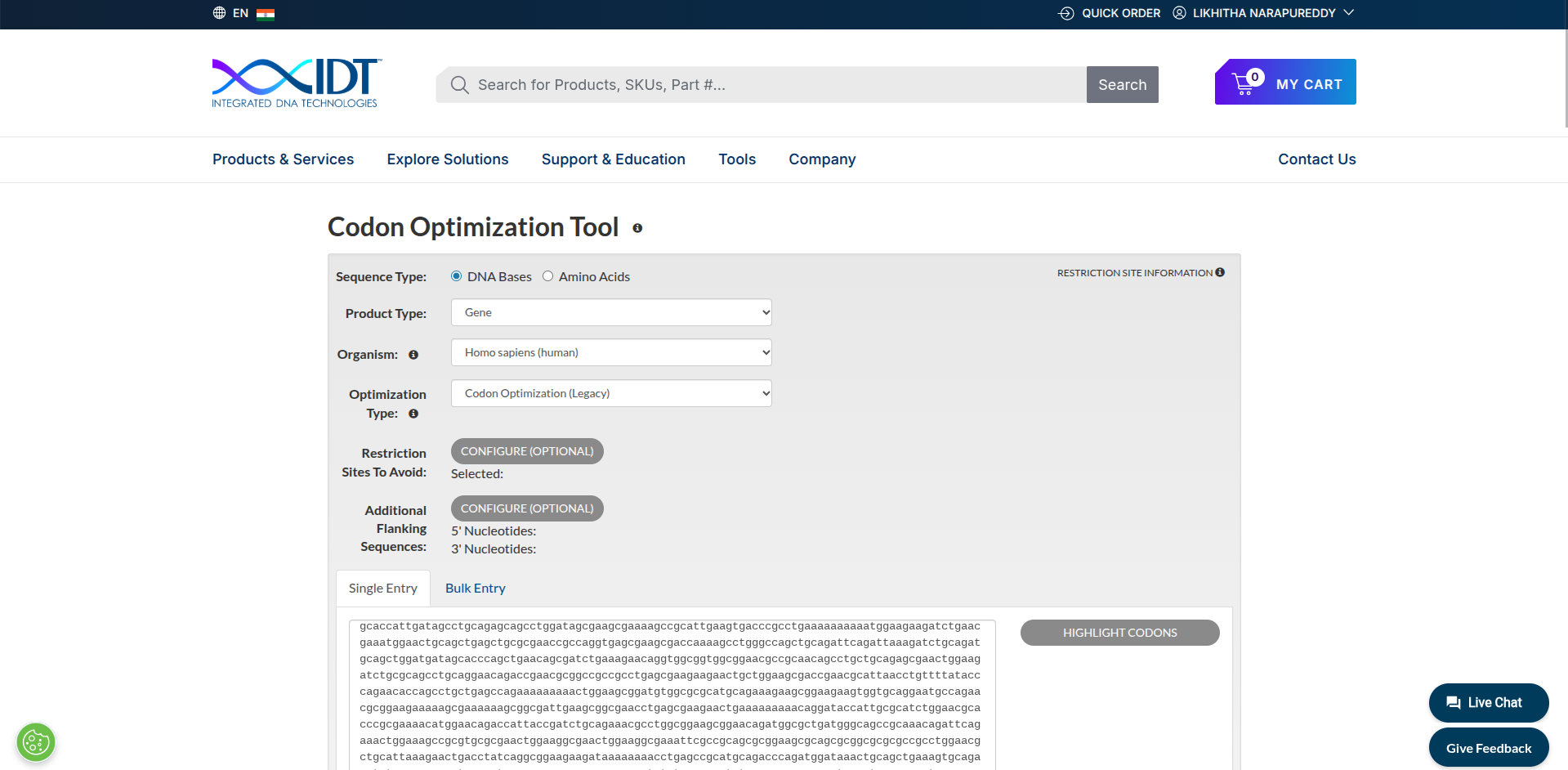

Myosin protein DNA-seq with codon optimization
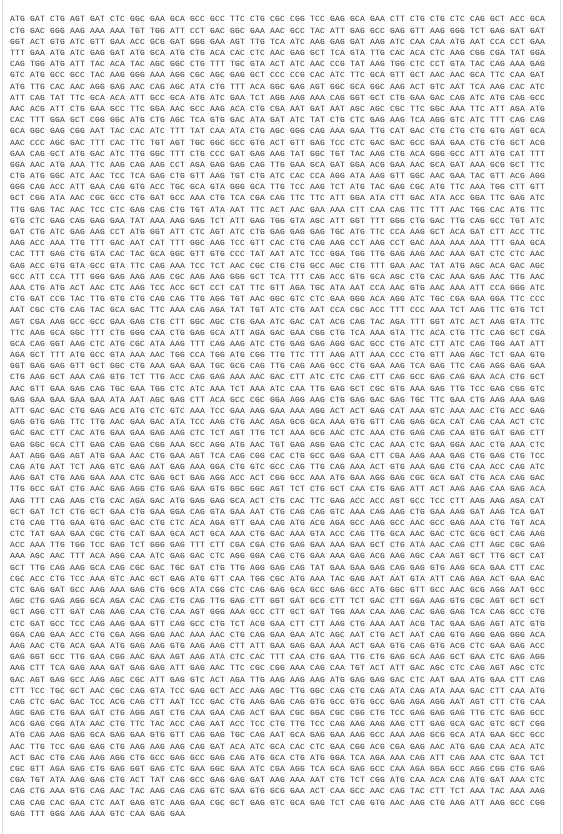


3.4. You have a sequence! Now what?
After obtaining the myosin protein sequence from UniProt and reverse translating it into a DNA coding sequence, the gene can be optimized for expression in a suitable host such as Escherichia coli or mammalian cells. The optimized gene is inserted into an expression plasmid under a strong promoter. Once introduced into the host cells, RNA polymerase transcribes the DNA into mRNA. The ribosome then translates the mRNA into the myosin polypeptide by reading codons and assembling the corresponding amino acids. After translation, the protein folds into its functional three-dimensional structure and can be purified using affinity chromatography techniques.
Part 4: Prepare a Twist DNA Synthesis Order
I successfully logged in both Twist and benchling. I followed the steps mentioned on the week2 homework site, mapped the sequence, and then completed the annotation(4.2)
Here are my results for the step 4.2:
The image depicts the linear map of the mapped sequence
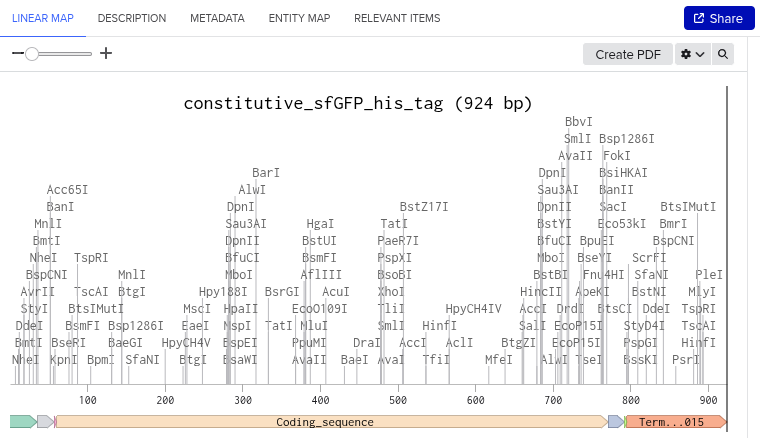

Subsequently, I uploaded the downloaded data in FASTA format to Twist. I selected the clonal genes option and uploaded the file. I then chose the pTwist Amp High Copy vector and dowloaded the resulting sequence. Later, I uploaded the downloaded data from Twist into Benchling. Resulted in creation of a ‘beautiful’ plasmid construct
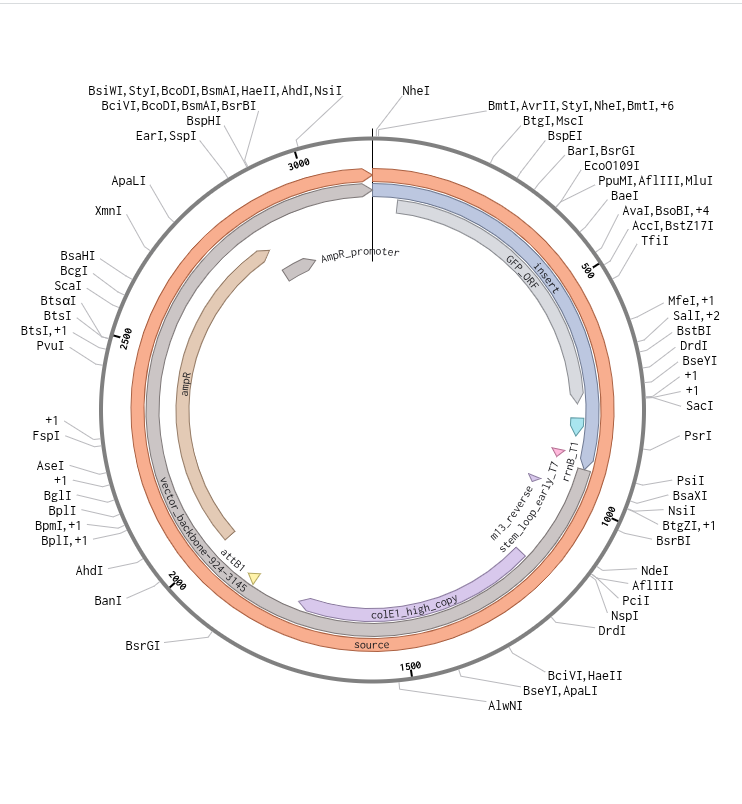

Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
For DNA sequencing, I would focus on microalgae and fungi that have strong potential for carbon capture and air purification. Species such as Chlorella vulgaris, Spirulina platensis, and Aspergillus niger are promising because they can absorb carbon dioxide, tolerate environmental stress, and in some cases degrade pollutants.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why? To perform sequencing, I would use Next-Generation Sequencing (NGS), specifically short-read sequencing developed by Illumina.
Also answer the following questions:
1.Is your method first-, second- or third-generation or other? How so?
This is a second-generation sequencing technology because it performs massively parallel sequencing of millions of DNA fragments simultaneously using sequencing-by-synthesis chemistry.
2.What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
- DNA extraction
- Fragmentation
- Adapter ligation
- PCR amplification
- Load library onto flow cell
- Software converts color signals → A, T, C, G (base calling)
3.What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
- DNA fragments bind to flow cell
- Bridge amplification creates clusters
- Fluorescently labeled nucleotides are added
- Each base emits a specific color signal
- A camera records the fluorescence
- Software converts color signals → A, T, C, G (base calling)
4.What is the output of your chosen sequencing technology?
- FASTQ files (raw reads with quality scores)
- Millions of short reads (150–300 bp)
- After assembly → full genome sequence
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?
For DNA synthesis, I would design a synthetic gene cassette to enhance carbon capture efficiency in microalgae. The construct would include a strong promoter, ribosome binding site, an optimized RuBisCO gene, and a terminator. Additional genes for stress tolerance or pollutant degradation could also be incorporated. The goal would be to create a genetic circuit that increases CO₂ fixation and improves survival in polluted environments.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
DNA synthesis would be performed using commercial gene synthesis services such as Twist Bioscience. The process involves digital DNA design, chemical synthesis of short oligonucleotides, assembly into a full-length gene, error correction, cloning into a plasmid, and sequence verification. Limitations include higher costs for long sequences, challenges with GC-rich regions, and potential synthesis errors. However, synthetic DNA enables precise control over gene design and optimization.
5.3 DNA Edit
(i) What DNA would you want to edit and why?
To further improve air filtration capabilities, I would edit the genomes of selected algae or fungi to enhance carbon fixation, increase pollutant tolerance, or remove metabolic bottlenecks. Genome editing could allow insertion of stronger promoters, modification of enzyme efficiency, or deletion of growth-limiting genes.
(ii) What technology or technologies would you use to perform these DNA edits and why?
The preferred editing tool would be CRISPR-Cas9. This system uses a guide RNA to direct the Cas9 enzyme to a specific DNA sequence, where it introduces a double-strand break.
Also answer the following questions:
- How does your technology of choice edit DNA? What are the essential steps?
a.Design guide RNA (gRNA) targeting gene b.Deliver Cas9 + gRNA into cell c.Cas9 creates double-strand break d.Cell repairs break: NHEJ → knockout and HDR → precise insertion
- What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
Inputs Required a.Guide RNA b.Cas9 protein or plasmid c.Donor DNA template (for HDR) d.Host cells e.Transformation method
- What are the limitations of your editing methods (if any) in terms of efficiency or precision?
Limitations include potential off-target mutations, variable editing efficiency, and delivery challenges in certain microalgal species. Ecological and regulatory considerations must also be addressed before environmental deployment.
Conclusion By integrating DNA sequencing, synthesis, and genome editing, it is possible to design and engineer enhanced biological air filtration systems. Sequencing reveals the natural genetic toolkit of algae and fungi, synthetic DNA enables rational design of improved pathways, and CRISPR-based editing allows precise genome modifications. Together, these technologies provide a powerful framework for developing sustainable, living solutions to air pollution and climate change mitigation.
References
1.Kumar, P., Arora, K., Chanana, I., Kulshreshtha, S., Thakur, V., & Choi, K.-Y. (2023). Comparative study on conventional and microalgae-based air purifiers: Paving the way for sustainable green spaces. Journal of Environmental Chemical Engineering, 11(6), 111046. https://doi.org/10.1016/j.jece.2023.111046
2.Marycz, M., Brillowska-Dąbrowska, A., Muñoz, R., Gębicki, J., et al. (2021). A state of the art review on the use of fungi in biofiltration to remove volatile hydrophobic pollutants. Reviews in Environmental Science and Bio/Technology. https://doi.org/10.1007/s11157-021-09608-7