Week 11 HW: Bioproduction and Cloud Labs
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
- Contribute at least one pixel to this global artwork experiment before the editing ends on Sunday 4/19 at 11:59 PM EST.
A personalized URL was sent to the email address associated with your Discourse account, and you can discuss the artwork on the Discourse.
If you did not have a chance to contribute, it’s okay, just make sure you become a TA this fall! 😉
- Make a note on your HTGAA webpages including:
what you contributed to the community bioart project (e.g., “I made part of the DNA on the bottom right plate”)
what you liked about the project, and
what about this collaborative art experiment could be made better for next year.
Ans: I made a character red from the game Among Us at the right corner of Q1. This project gave freedom to everyone to create any form of art and I liked it.
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
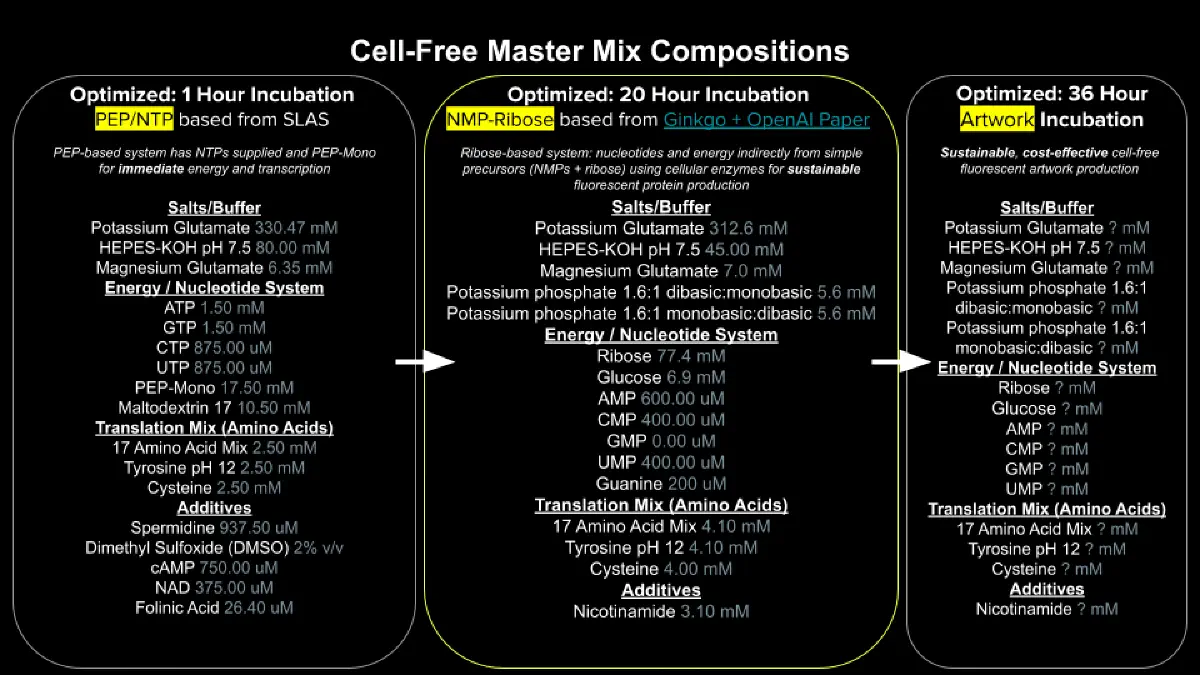

- Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
Ans: Components & Their Roles
E. coli Lysate
BL21 (DE3) Star Lysate (includes T7 RNA Polymerase) Provides the core cellular machinery—ribosomes, tRNAs, enzymes for transcription and translation; the built-in T7 RNA polymerase drives high-level transcription from T7 promoters.
Salts / Buffer
Potassium Glutamate Mimics intracellular ionic conditions and stabilizes ribosomes and enzymes for efficient protein synthesis.
HEPES-KOH pH 7.5 Maintains a stable physiological pH, which is critical for enzyme activity during transcription and translation.
Magnesium Glutamate Essential cofactor for ribosomes and polymerases; directly impacts translation efficiency and RNA stability.
Potassium Phosphate (Monobasic & Dibasic) Provides buffering capacity and phosphate ions, helping maintain pH balance and supporting energy metabolism.
Energy / Nucleotide System
Ribose Serves as a carbon source for nucleotide regeneration through metabolic pathways.
Glucose Fuels ATP regeneration via glycolytic enzymes present in the lysate, enabling longer reaction lifetimes.
AMP, CMP, GMP, UMP Nucleotide monophosphates that are enzymatically converted into triphosphates (ATP, GTP, etc.) required for transcription and translation.
Guanine A nucleobase precursor that can be salvaged into GMP, contributing to nucleotide pool replenishment.
Translation Mix (Amino Acids)
17 Amino Acid Mix Supplies most of the building blocks required for protein synthesis.
Tyrosine & Cysteine Added separately because they are less stable or more reactive, ensuring sufficient availability during translation.
Additives
Nicotinamide Supports redox balance and metabolic reactions (via NAD⁺/NADH systems), improving energy regeneration and reaction longevity.
Backfill
Nuclease-Free Water Adjusts the final reaction volume while preventing degradation of nucleic acids.
- Describe the main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the Google Slide above. (2-3 sentences)
Ans: The PEP-NTP (1-hour) system uses high-energy molecules like phosphoenolpyruvate and directly supplied nucleotide triphosphates, enabling rapid and high protein production but with short reaction lifetimes. The NMP-Ribose-Glucose (20-hour) system relies on slower metabolic regeneration of energy and nucleotides from simpler precursors, resulting in longer-lasting but lower-rate protein synthesis.
- Bonus question: How can transcription occur if GMP is not included but Guanine is?
Ans: Transcription can still occur because guanine can be salvaged into GMP via enzymatic pathways present in the lysate (nucleotide salvage pathways). Once converted to GMP, it can then be phosphorylated to GTP, which is the actual substrate used by RNA polymerase.
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
- Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each)
sfGFP
mRFP1
mKO2
mTurquoise2
mScarlet_I
Electra2
Ans: sfGFP
Key property: Robust folding efficiency sfGFP is engineered to fold efficiently even under suboptimal conditions, making it highly reliable in cell-free systems where chaperones may be limited.
mRFP1
Key property: Slow maturation time
mRFP1 takes longer to form its fluorescent chromophore, so signal appears delayed even if protein expression is occurring.
mTurquoise2
Key property: High quantum yield (brightness)
This cyan protein is extremely bright and efficient at emitting light, allowing sensitive detection even at low expression levels.
mScarlet-I
Key property: Rapid maturation and high brightness
mScarlet-I combines fast chromophore formation with strong fluorescence, making it one of the best-performing red proteins in cell-free systems.
Electra2
Key property: Oxygen dependent chromophore formation
Like most fluorescent proteins, Electra2 requires oxygen for chromophore maturation, so limited oxygen in cell-free reactions can reduce or delay fluorescence.
- Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
Ans: Hypothesis for Electra2 (maximize fluorescence over 36 hours)
Reducing the glucose concentration in the cell-free mastermix will decrease metabolic oxygen consumption, thereby increasing dissolved oxygen availability for Electra2 chromophore maturation and improving fluorescence over a 36-hour incubation.
Reagent to adjust:
Glucose (energy source in the mastermix)
Expected effect: Lower glucose levels will slow ATP-generating metabolism, reducing oxygen depletion in the reaction. This preserves oxygen needed for Electra2’s oxygen-dependent chromophore formation, leading to a higher fraction of properly matured fluorescent protein and increased overall fluorescence signal after 36 hours.