Subsections of Likhitha — HTGAA Spring 2026
Homework
Weekly homework submissions:
Week 1 HW: Principles and Practices
A Living Anti-Corrosion System for Ocean Infrastructure:
Week 2 pre HW: DNA Read, Write and Edit
Homework Questions from Professor Jacobson: Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy? Ans: DNA polymerase has an inherent error rate of approximately 1 in 10⁶ bases. Given the human genome size of about 3.2 billion base pairs, this would lead to thousands of mutations each time a cell divides if left uncorrected. To maintain genomic stability, cells use a multi-layered error-correction system. First, DNA polymerase performs immediate proofreading through its exonuclease activity. This is followed by post-replication mismatch repair (MMR) mechanisms. Together, these processes greatly enhance replication accuracy, reducing the final error rate to roughly 1 in 10⁹–10¹⁰, meaning fewer than one error typically occurs per genome duplication.
Week 2 HW: DNA - read, write and edit
Part 1- Benchling & In-silico Gel Art Simulating Restriction Enzyme Digestion with the following Enzymes: .EcoRI .HindIII .BamHI
My art for Opentrons Artwork Output of the python script. PAPER - Semiautomated Production of Cell-Free Biosensors Journal: ACS Synthetic Biology (2025) PMID: 40073441 Biosensors are biological systems that detect specific chemicals for example, if a substance is present, they might change color or glow. These can be used for:
Week 4 HW: Protein design part I
Part A. Conceptual Questions How many amino acid molecules are in 500 g of meat? Ans: Average amino acid ≈ 100 Daltons (100 g/mol) 500 g ÷ 100 g/mol = 5 moles 1 mole = 6.022 × 10²³ molecules So, 5 × 6.022 × 10²³ ≈ 3 × 10²⁴ amino acid molecules Why do humans eat beef but do not become a cow, eat fish but do not become fish? Ans: When we eat beef or fish, our body breaks proteins into amino acids during digestion. Then we rebuild them into human proteins, not cow proteins.
Week 7 HW: Genetic Circuits partII: Neuromorphic circuits
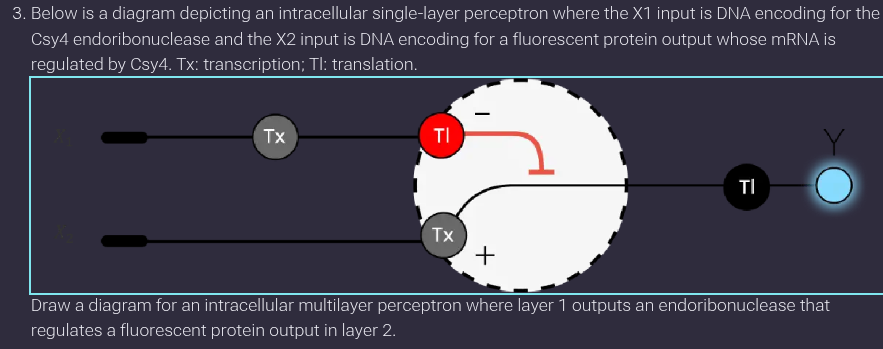
Assignment Part 1: Intracellular Artificial Neural Networks (IANNs)
- What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions? Traditional genetic circuits mostly behave like Boolean logic gates (ON/OFF). Intracellular Artificial Neural Networks (IANNs) are more flexible. Advantages: a. Analog (continuous) behavior -> Traditional circuits: only 0 or 1 (OFF/ON)
Homework Part A: General and Lecturer-Specific Questions
- Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production. Ans: Cell-free protein synthesis offers significant advantages over in vivo methods due to its open and controllable nature. It allows direct manipulation of reaction components, precise control over parameters such as pH and substrate concentration, and eliminates constraints related to cell viability. As a result, all system resources can be directed toward protein production, enabling rapid optimization and high-throughput experimentation.
Week 10 HW: Advanced Imaging & Measurement Technology
Homework: Final Project For your final project: Please identify at least one (ideally many) aspect(s) of your project that you will measure. It could be the mass or sequence of a protein, the presence, absence, or quantity of a biomarker, etc. Please describe all of the elements you would like to measure, and furthermore describe how you will perform these measurements. What are the technologies you will use (e.g., gel electrophoresis, DNA sequencing, mass spectrometry, etc.)? Describe in detail.
Week 11 HW: Bioproduction and Cloud Labs
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork Contribute at least one pixel to this global artwork experiment before the editing ends on Sunday 4/19 at 11:59 PM EST. A personalized URL was sent to the email address associated with your Discourse account, and you can discuss the artwork on the Discourse. If you did not have a chance to contribute, it’s okay, just make sure you become a TA this fall! 😉
Subsections of Homework
Week 1 HW: Principles and Practices


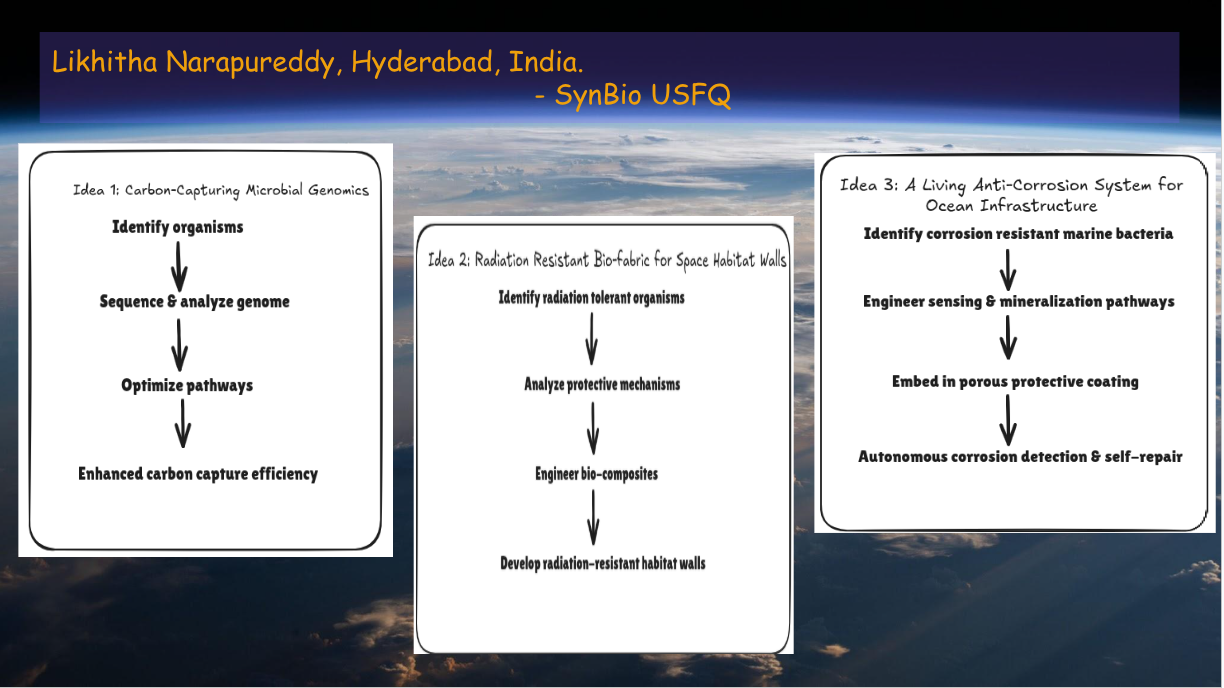
A Living Anti-Corrosion System for Ocean Infrastructure:
I propose a biologically engineered, self-healing anti-corrosion coating for offshore and ocean-energy infrastructure (tidal turbines, wave energy converters, offshore wind foundations).The system uses genetically engineered, non-pathogenic marine bacteria embedded in a sealed, porous coating. These microbes are designed to:
- Detect early corrosion signals (pH drop, Fe²⁺ ion release)
- Respond by precipitating protective minerals (e.g., calcium carbonate)
- Neutralize corrosive microenvironments
- Signal early warnings before structural failure
Governance Policy Goals To ensure this application contributes to an ethical future and prevents harm, governance should pursue the following goals:
- Ensure safety and security: a) Prevent environmental release or misuse of engineered organisms and b) Avoid ecological disruption or biosecurity risks
- Promote constructive and beneficial use a) Direct innovation toward public-interest infrastructure (renewable energy, climate resilience) and b)Prevent purely extractive or environmentally harmful deployment
Action 1: Mandatory Biological Containment & Kill-Switch Standards Purpose: What is done now: Chemical anti-corrosion coatings are regulated mainly for toxicity, not biological behavior.
Proposed change: Require all engineered microbes used in marine infrastructure to include:
- Genetic kill switches
- Nutrient dependency (cannot survive outside coating)
- Physical encapsulation in sealed matrices
- Maintain transparency and public trust Design : Needed - International biosafety certification for “contained-use marine bio-systems” Assumptions: Kill switches will function reliably in harsh marine environments What could be wrong:Evolutionary escape mechanisms
Action 2: Environmental impact and Community Consent framework Purpose What is done now: Environmental Impact Assessments (EIAs) often focus on physical structures, not biological agents.
Proposed change: Require Bio-Environmental Impact Assessments (Bio-EIAs) that include:
- Long-term microbial ecosystem modeling
- Transparent disclosure of organism function
- Consultation with coastal and fishing communities Design: Needed- Continuous post-deployment monitoring Assumptions: Communities can meaningfully engage with technical information What could be wrong:Information asymmetry and monitoring fatigue over time
Action 3: Restricted Use Licensing (Purpose-Bound Deployment) Purpose What is done now: Biotechnologies often spread from research into unintended domains (e.g., CRISPR kits, dual-use chemicals).
Proposed change: License this technology only for defined applications:
- Renewable energy infrastructure
- Public maritime assets Design: Needed- Purpose-specific approval, audits of deployment sites and clear penalties for misuse Assumptions: Clear boundaries between “civil” and “non-civil” uses exist What could be wrong: Commercial influence to expand scope
Next, score (from 1-3 with, 1 as the best, or n/a) each of your governance actions against your rubric of policy goals. The following is one framework but feel free to make your own:
| Governance Action | Safety & Security | Constructive & Beneficial Uses |
|---|---|---|
| Option 1: Bio-containment standards | 1 | 2 |
| Option 2: Voluntary guidelines | 3 | 3 |
| Option 3: Impact & consent reviews | 2 | 1 |
Based on the scores, I would prioritize Option 1 (Bio-containment standards) together with Option 3 (Impact and community consent reviews).
Option 1 comes first because safety has to be the foundation. When we introduce engineered biological systems into the ocean, the biggest risk is that something spreads, mutates, or behaves in ways we didn’t expect. Strong bio-containment rules like built-in kill switches or limits on survival outside controlled conditions help prevent accidents before they happen. Without this layer, even well-intended projects could cause long-term environmental harm.
Option 3 is equally important because it helps make sure the technology is actually used for good. Environmental impact checks and community consent force developers to think beyond the lab and consider real ocean ecosystems and the people who depend on them. This option scored highest for promoting constructive and beneficial uses because it guides innovation toward solutions that are socially and environmentally responsible, not just technically impressive.
I would not rely on Option 2 (Voluntary guidelines) on its own. While voluntary rules can encourage early innovation, they are easy to ignore and don’t offer strong protection when risks are high. For ocean systems, where damage can be difficult or impossible to reverse, voluntary measures are not enough.
Overall, combining strong safety rules with environmental and community oversight offers the most realistic and responsible way to move forward. It protects the ocean while still allowing beneficial innovation to happen.
References:
Jin, H., Wang, J., Tian, L., Gao, M., Zhao, J., & Ren, L. (2022). Recent advances in emerging integrated antifouling and anticorrosion coatings. Materials & Design, 213, 110307. https://doi.org/10.1016/j.matdes.2021.110307
Li, Y., & Ning, C. (2019). Latest research progress of marine microbiological corrosion and bio-fouling, and new approaches of marine anti-corrosion and anti-fouling. Bioactive Materials, 4, 189–195. https://doi.org/10.1016/j.bioactmat.2019.04.003
Week 2 pre HW: DNA Read, Write and Edit
Homework Questions from Professor Jacobson:
Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy? Ans: DNA polymerase has an inherent error rate of approximately 1 in 10⁶ bases. Given the human genome size of about 3.2 billion base pairs, this would lead to thousands of mutations each time a cell divides if left uncorrected. To maintain genomic stability, cells use a multi-layered error-correction system. First, DNA polymerase performs immediate proofreading through its exonuclease activity. This is followed by post-replication mismatch repair (MMR) mechanisms. Together, these processes greatly enhance replication accuracy, reducing the final error rate to roughly 1 in 10⁹–10¹⁰, meaning fewer than one error typically occurs per genome duplication.
How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest? Ans: An average human protein (~1036 bp) can be coded by many synonymous codons because the genetic code is redundant.In practice, many of these codes fail because they can create secondary structures that block translation, contain sequences that trigger RNA cleavage, or use “rare” codons that the host cell cannot efficiently process.
Homework Questions from Dr. LeProust:
What’s the most commonly used method for oligo synthesis currently? Ans: The phosphoramidite method is currently the most widely used chemistry for oligonucleotide synthesis. It involves a four-step cyclic process: coupling, capping, oxidation, and deblocking—to add nucleotides one by one onto a solid support, such as Controlled Pore Glass (CPG) or silicon chips.
Why is it difficult to make oligos longer than 200nt via direct synthesis? Ans: Oligonucleotide synthesis occurs by adding one nucleotide at a time, and each step has a small probability of error. As the length increases, these errors accumulate, reducing the yield of correct full-length oligos. In addition, longer oligos are more prone to incomplete reactions and strand loss during synthesis.
Why can’t you make a 2000bp gene via direct oligo synthesis? Ans: Directly synthesizing a 2000 bp gene is not feasible in practice. Errors accumulate with each step, resulting in low yield and incorrect sequences. Hence, long genes are constructed by assembling shorter, accurately synthesized oligos and then applying error-correction methods.
Homework Question from George Church: The question choosed by me
- [Using Google & Prof. Church’s slide #4] What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”? Ans: Essential amino acids in animals (10):
Histidine, Isoleucine, Leucine, Lysine, Methionine, Phenylalanine, Threonine, Tryptophan, Valine, and Arginine . These amino acids cannot be synthesized by animals and must be obtained from the diet
Lysine contingency : Lysine contingency refers to the idea that animal life is fundamentally dependent on external sources of lysine because animals cannot synthesize it themselves. Since lysine is an essential amino acid and often scarce in plant-based foods, growth and survival become contingent on its availability. This makes lysine a key metabolic bottleneck shaping nutrition, agriculture, and evolutionary constraints.
My views The fact that lysine is an essential amino acid for all animals reinforces the idea of the lysine contingency—that animal life is inherently dependent on external biological systems (plants, microbes, or other animals) to supply lysine.We are all metabolically fasten to the external world, relying on a constant “supply chain” of plants and microbes to build our bodies.
Even when we think about survival in extreme environments, like a colony on Mars. Instead of trying to “fix” human genetics to make us self-sufficient which is ethically messy and biologically complex it makes far more sense to master the environment around us. By engineering hardy, high-yield, lysine-producing plants or yeast, we solve the survival puzzle without ever touching a human strand of DNA. It’s a strategy that’s not only safer and more flexible but one that respects our natural biology by simply ensuring the “bio-battery” we’ve always relied on never runs dry. At last the lysine contingency shows that human survival depends on food systems, not genetic independence. For extreme environments, engineering plants and microbes is a safer and smarter solution than changing human biology or animal biology.
Reference Jurassic Park Wiki. (n.d.). Lysine contingency. Fandom. Retrieved from https://jurassicpark.fandom.com/wiki/Lysine_contingency
Week 2 HW: DNA - read, write and edit
Part 1- Benchling & In-silico Gel Art
Simulating Restriction Enzyme Digestion with the following Enzymes:
.EcoRI
.HindIII
.BamHI
.KpnI
.EcoRV
.SacI
.SalI
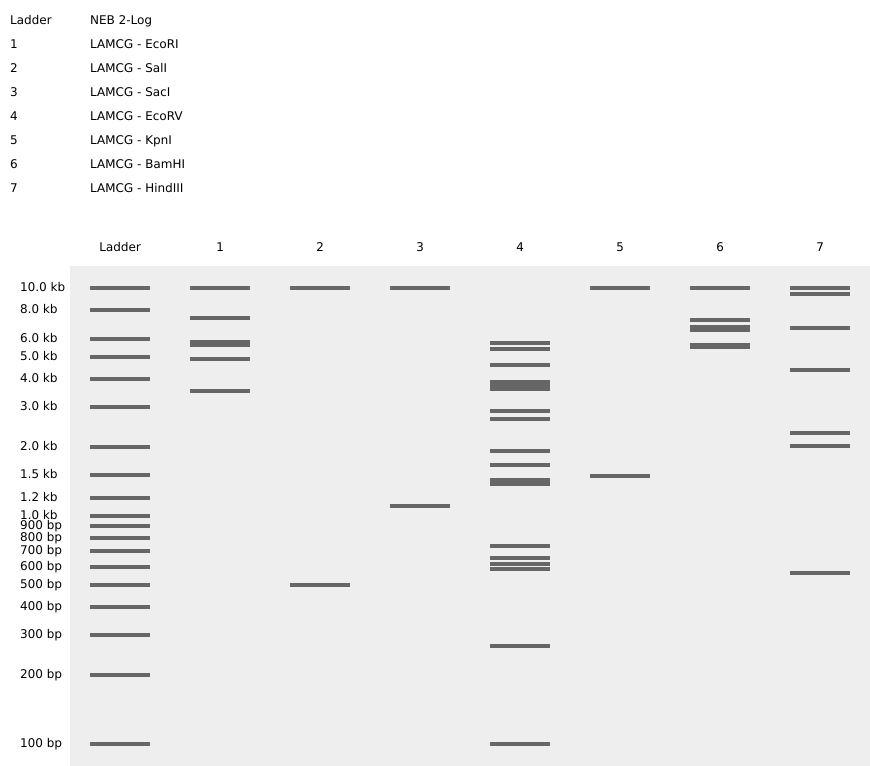


I had created a simple pattern in the style of Paul Vanouse’s Latent Figure Protocol artworks. The bands are arranged alternatively creating a horizontal alternative pattern.
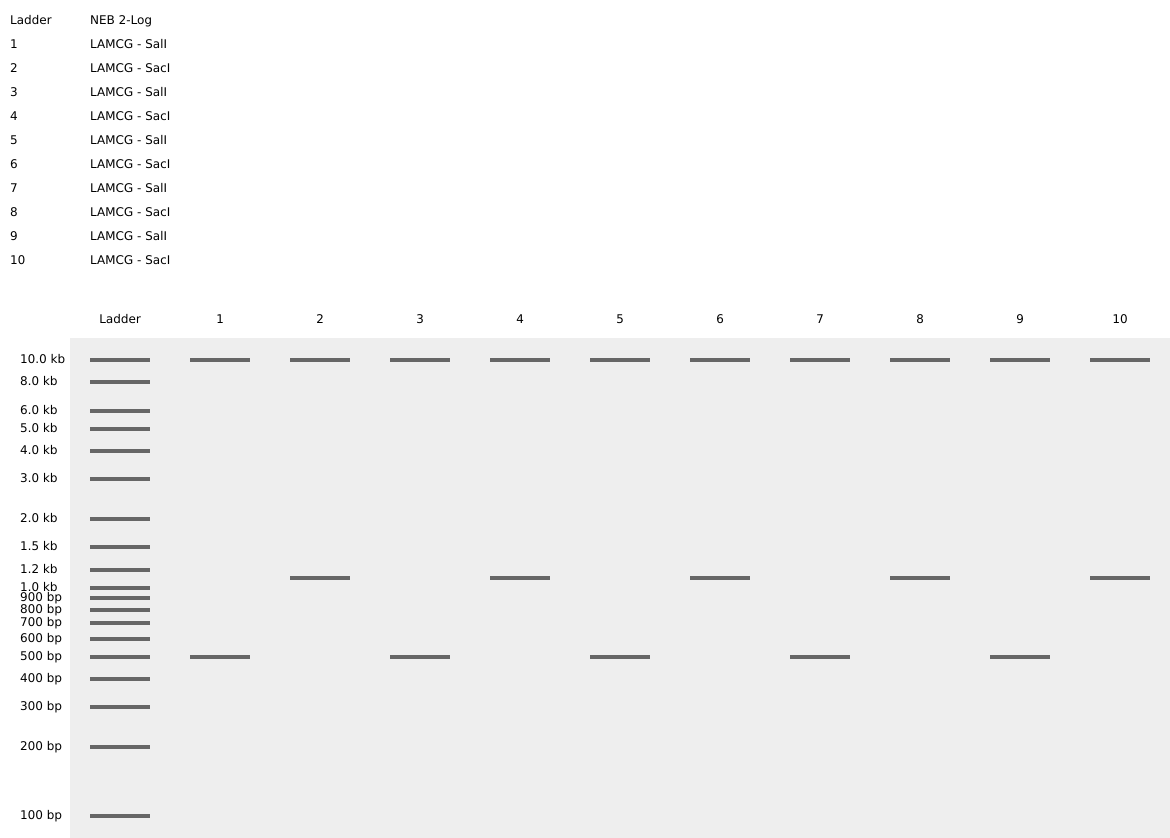

Part 3: DNA Design Challenge
3.1. Choose your protein The protein I had selected is Myosin, which is a motor protein responsible for muscle contraction. It binds to the actin filaments and uses ATP to generate force. This force pulls the actin filaments inwards causes the muscle fibres to shorten and contract. I am particularly interested in understanding how myosin behaves in microgravity conditions, where mechanical loading is absent and muscle atrophy occurs rapidly. I collected the protein from human at Uniprot- Q9Y2K3
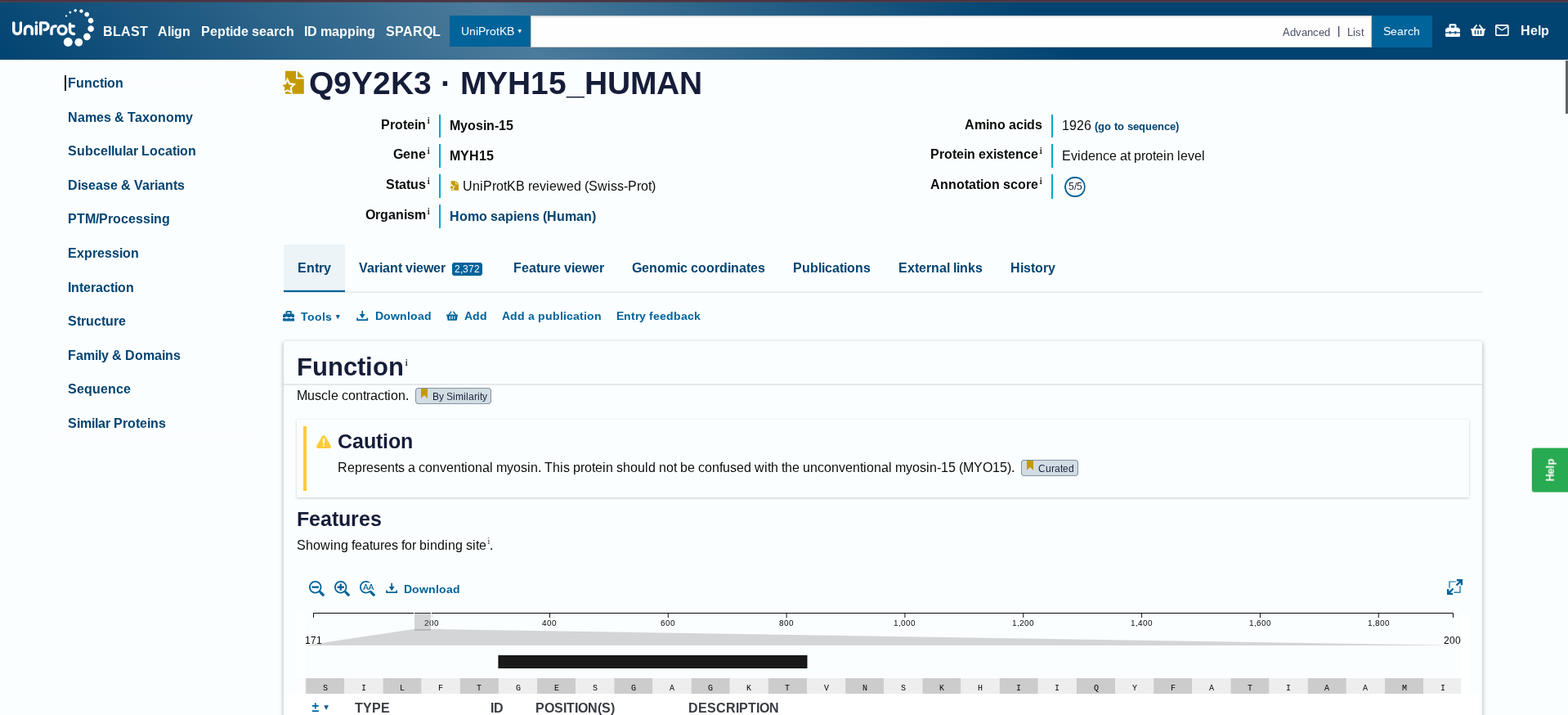
 Below is the protein sequence of Myosin-15 extracted from Homo sapien
Below is the protein sequence of Myosin-15 extracted from Homo sapien
sp|Q9Y2K3|MYH15_HUMAN Myosin-15 OS=Homo sapiens OX=9606 GN=MYH15 PE=1 SV=6 MDLSDLGEAAAFLRRSEAELLLLQATALDGKKKCWIPDGENAYIEAEVKGSEDDGTVIVE TADGESLSIKEDKIQQMNPPEFEMIEDMAMLTHLNEASVLHTLKRRYGQWMIYTYSGLFC VTINPYKWLPVYQKEVMAAYKGKRRSEAPPHIFAVANNAFQDMLHNRENQSILFTGESGA GKTVNSKHIIQYFATIAAMIESRKKQGALEDQIMQANTILEAFGNAKTLRNDNSSRFGKF
IRMHFGARGMLSSVDIDIYLLEKSRVIFQQAGERNYHIFYQILSGQKELHDLLLVSANPS DFHFCSCGAVTVESLDDAEELLATEQAMDILGFLPDEKYGCYKLTGAIMHFGNMKFKQKP REEQLEADGTENADKAAFLMGINSSELVKCLIHPRIKVGNEYVTRGQTIEQVTCAVGALS KSMYERMFKWLVARINRALDAKLSRQFFIGILDITGFEILEYNSLEQLCINFTNEKLQQF FNWHMFVLEQEEYKKESIEWVSIGFGLDLQACIDLIEKPMGILSILEEECMFPKATDLTF KTKLFDNHFGKSVHLQKPKPDKKKFEAHFELVHYAGVVPYNISGWLEKNKDLLNETVVAV FQKSSNRLLASLFENYMSTDSAIPFGEKKRKKGASFQTVASLHKENLNKLMTNLKSTAPH FVRCINPNVNKIPGILDPYLVLQQLRCNGVLEGTRICREGFPNRLQYADFKQRYCILNPR TFPKSKFVSSRKAAEELLGSLEIDHTQYRFGITKVFFKAGFLGQLEAIRDERLSKVFTLF QARAQGKLMRIKFQKILEERDALILIQWNIRAFMAVKNWPWMRLFFKIKPLVKSSEVGEE VAGLKEECAQLQKALEKSEFQREELKAKQVSLTQEKNDLILQLQAEQETLANVEEQCEWL IKSKIQLEARVKELSERVEEEEEINSELTARGRKLEDECFELKKEIDDLETMLVKSEKEK RTTEHKVKNLTEEVEFLNEDISKLNRAAKVVQEAHQQTLDDLHMEEEKLSSLSKANLKLE QQVDELEGALEQERKARMNCERELHKLEGNLKLNRESMENLESSQRHLAEELRKKELELS QMNSKVENEKGLVAQLQKTVKELQTQIKDLKEKLEAERTTRAKMERERADLTQDLADLNE RLEEVGGSSLAQLEITKKQETKFQKLHRDMEEATLHFETTSASLKKRHADSLAELEGQVE NLQQVKQKLEKDKSDLQLEVDDLLTRVEQMTRAKANAEKLCTLYEERLHEATAKLDKVTQ LANDLAAQKTKLWSESGEFLRRLEEKEALINQLSREKSNFTRQIEDLRGQLEKETKSQSA LAHALQKAQRDCDLLREQYEEEQEVKAELHRTLSKVNAEMVQWRMKYENNVIQRTEDLED AKKELAIRLQEAAEAMGVANARNASLERARHQLQLELGDALSDLGKVRSAAARLDQKQLQ SGKALADWKQKHEESQALLDASQKEVQALSTELLKLKNTYEESIVGQETLRRENKNLQEE ISNLTNQVREGTKNLTEMEKVKKLIEEEKTEVQVTLEETEGALERNESKILHFQLELLEA KAELERKLSEKDEEIENFRRKQQCTIDSLQSSLDSEAKSRIEVTRLKKKMEEDLNEMELQ LSCANRQVSEATKSLGQLQIQIKDLQMQLDDSTQLNSDLKEQVAVAERRNSLLQSELEDL RSLQEQTERGRRLSEEELLEATERINLFYTQNTSLLSQKKKLEADVARMQKEAEEVVQEC QNAEEKAKKAAIEAANLSEELKKKQDTIAHLERTRENMEQTITDLQKRLAEAEQMALMGS RKQIQKLESRVRELEGELEGEIRRSAEAQRGARRLERCIKELTYQAEEDKKNLSRMQTQM DKLQLKVQNYKQQVEVAETQANQYLSKYKKQQHELNEVKERAEVAESQVNKLKIKAREFG KKVQEE
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
The Central Dogma gives us the framework to work backwards from a given protein sequence and infer the DNA sequence that the protein is derived from. Reverse Translate results Results for 1926 residue sequence “sp|Q9Y2K3|MYH15_HUMAN Myosin-15 OS=Homo sapiens OX=9606 GN=MYH15 PE=1 SV=6” starting “MDLSDLGEAA”
reverse translation of sp|Q9Y2K3|MYH15_HUMAN Myosin-15 OS=Homo sapiens OX=9606 GN=MYH15 PE=1 SV=6 to a 5778 base sequence of most likely codons. atggatctgagcgatctgggcgaagcggcggcgtttctgcgccgcagcgaagcggaactg ctgctgctgcaggcgaccgcgctggatggcaaaaaaaaatgctggattccggatggcgaa aacgcgtatattgaagcggaagtgaaaggcagcgaagatgatggcaccgtgattgtggaa accgcggatggcgaaagcctgagcattaaagaagataaaattcagcagatgaacccgccg gaatttgaaatgattgaagatatggcgatgctgacccatctgaacgaagcgagcgtgctg cataccctgaaacgccgctatggccagtggatgatttatacctatagcggcctgttttgc gtgaccattaacccgtataaatggctgccggtgtatcagaaagaagtgatggcggcgtat aaaggcaaacgccgcagcgaagcgccgccgcatatttttgcggtggcgaacaacgcgttt caggatatgctgcataaccgcgaaaaccagagcattctgtttaccggcgaaagcggcgcg ggcaaaaccgtgaacagcaaacatattattcagtattttgcgaccattgcggcgatgatt gaaagccgcaaaaaacagggcgcgctggaagatcagattatgcaggcgaacaccattctg gaagcgtttggcaacgcgaaaaccctgcgcaacgataacagcagccgctttggcaaattt attcgcatgcattttggcgcgcgcggcatgctgagcagcgtggatattgatatttatctg ctggaaaaaagccgcgtgatttttcagcaggcgggcgaacgcaactatcatattttttat cagattctgagcggccagaaagaactgcatgatctgctgctggtgagcgcgaacccgagc gattttcatttttgcagctgcggcgcggtgaccgtggaaagcctggatgatgcggaagaa ctgctggcgaccgaacaggcgatggatattctgggctttctgccggatgaaaaatatggc tgctataaactgaccggcgcgattatgcattttggcaacatgaaatttaaacagaaaccg cgcgaagaacagctggaagcggatggcaccgaaaacgcggataaagcggcgtttctgatg ggcattaacagcagcgaactggtgaaatgcctgattcatccgcgcattaaagtgggcaac gaatatgtgacccgcggccagaccattgaacaggtgacctgcgcggtgggcgcgctgagc aaaagcatgtatgaacgcatgtttaaatggctggtggcgcgcattaaccgcgcgctggat gcgaaactgagccgccagttttttattggcattctggatattaccggctttgaaattctg gaatataacagcctggaacagctgtgcattaactttaccaacgaaaaactgcagcagttt tttaactggcatatgtttgtgctggaacaggaagaatataaaaaagaaagcattgaatgg gtgagcattggctttggcctggatctgcaggcgtgcattgatctgattgaaaaaccgatg ggcattctgagcattctggaagaagaatgcatgtttccgaaagcgaccgatctgaccttt aaaaccaaactgtttgataaccattttggcaaaagcgtgcatctgcagaaaccgaaaccg gataaaaaaaaatttgaagcgcattttgaactggtgcattatgcgggcgtggtgccgtat aacattagcggctggctggaaaaaaacaaagatctgctgaacgaaaccgtggtggcggtg tttcagaaaagcagcaaccgcctgctggcgagcctgtttgaaaactatatgagcaccgat agcgcgattccgtttggcgaaaaaaaacgcaaaaaaggcgcgagctttcagaccgtggcg agcctgcataaagaaaacctgaacaaactgatgaccaacctgaaaagcaccgcgccgcat tttgtgcgctgcattaacccgaacgtgaacaaaattccgggcattctggatccgtatctg gtgctgcagcagctgcgctgcaacggcgtgctggaaggcacccgcatttgccgcgaaggc tttccgaaccgcctgcagtatgcggattttaaacagcgctattgcattctgaacccgcgc acctttccgaaaagcaaatttgtgagcagccgcaaagcggcggaagaactgctgggcagc ctggaaattgatcatacccagtatcgctttggcattaccaaagtgttttttaaagcgggc tttctgggccagctggaagcgattcgcgatgaacgcctgagcaaagtgtttaccctgttt caggcgcgcgcgcagggcaaactgatgcgcattaaatttcagaaaattctggaagaacgc gatgcgctgattctgattcagtggaacattcgcgcgtttatggcggtgaaaaactggccg tggatgcgcctgttttttaaaattaaaccgctggtgaaaagcagcgaagtgggcgaagaa gtggcgggcctgaaagaagaatgcgcgcagctgcagaaagcgctggaaaaaagcgaattt cagcgcgaagaactgaaagcgaaacaggtgagcctgacccaggaaaaaaacgatctgatt ctgcagctgcaggcggaacaggaaaccctggcgaacgtggaagaacagtgcgaatggctg attaaaagcaaaattcagctggaagcgcgcgtgaaagaactgagcgaacgcgtggaagaa gaagaagaaattaacagcgaactgaccgcgcgcggccgcaaactggaagatgaatgcttt gaactgaaaaaagaaattgatgatctggaaaccatgctggtgaaaagcgaaaaagaaaaa cgcaccaccgaacataaagtgaaaaacctgaccgaagaagtggaatttctgaacgaagat attagcaaactgaaccgcgcggcgaaagtggtgcaggaagcgcatcagcagaccctggat gatctgcatatggaagaagaaaaactgagcagcctgagcaaagcgaacctgaaactggaa cagcaggtggatgaactggaaggcgcgctggaacaggaacgcaaagcgcgcatgaactgc gaacgcgaactgcataaactggaaggcaacctgaaactgaaccgcgaaagcatggaaaac ctggaaagcagccagcgccatctggcggaagaactgcgcaaaaaagaactggaactgagc cagatgaacagcaaagtggaaaacgaaaaaggcctggtggcgcagctgcagaaaaccgtg aaagaactgcagacccagattaaagatctgaaagaaaaactggaagcggaacgcaccacc cgcgcgaaaatggaacgcgaacgcgcggatctgacccaggatctggcggatctgaacgaa cgcctggaagaagtgggcggcagcagcctggcgcagctggaaattaccaaaaaacaggaa accaaatttcagaaactgcatcgcgatatggaagaagcgaccctgcattttgaaaccacc agcgcgagcctgaaaaaacgccatgcggatagcctggcggaactggaaggccaggtggaa aacctgcagcaggtgaaacagaaactggaaaaagataaaagcgatctgcagctggaagtg gatgatctgctgacccgcgtggaacagatgacccgcgcgaaagcgaacgcggaaaaactg tgcaccctgtatgaagaacgcctgcatgaagcgaccgcgaaactggataaagtgacccag ctggcgaacgatctggcggcgcagaaaaccaaactgtggagcgaaagcggcgaatttctg cgccgcctggaagaaaaagaagcgctgattaaccagctgagccgcgaaaaaagcaacttt acccgccagattgaagatctgcgcggccagctggaaaaagaaaccaaaagccagagcgcg ctggcgcatgcgctgcagaaagcgcagcgcgattgcgatctgctgcgcgaacagtatgaa gaagaacaggaagtgaaagcggaactgcatcgcaccctgagcaaagtgaacgcggaaatg gtgcagtggcgcatgaaatatgaaaacaacgtgattcagcgcaccgaagatctggaagat gcgaaaaaagaactggcgattcgcctgcaggaagcggcggaagcgatgggcgtggcgaac gcgcgcaacgcgagcctggaacgcgcgcgccatcagctgcagctggaactgggcgatgcg ctgagcgatctgggcaaagtgcgcagcgcggcggcgcgcctggatcagaaacagctgcag agcggcaaagcgctggcggattggaaacagaaacatgaagaaagccaggcgctgctggat gcgagccagaaagaagtgcaggcgctgagcaccgaactgctgaaactgaaaaacacctat gaagaaagcattgtgggccaggaaaccctgcgccgcgaaaacaaaaacctgcaggaagaa attagcaacctgaccaaccaggtgcgcgaaggcaccaaaaacctgaccgaaatggaaaaa gtgaaaaaactgattgaagaagaaaaaaccgaagtgcaggtgaccctggaagaaaccgaa ggcgcgctggaacgcaacgaaagcaaaattctgcattttcagctggaactgctggaagcg aaagcggaactggaacgcaaactgagcgaaaaagatgaagaaattgaaaactttcgccgc aaacagcagtgcaccattgatagcctgcagagcagcctggatagcgaagcgaaaagccgc attgaagtgacccgcctgaaaaaaaaaatggaagaagatctgaacgaaatggaactgcag ctgagctgcgcgaaccgccaggtgagcgaagcgaccaaaagcctgggccagctgcagatt cagattaaagatctgcagatgcagctggatgatagcacccagctgaacagcgatctgaaa gaacaggtggcggtggcggaacgccgcaacagcctgctgcagagcgaactggaagatctg cgcagcctgcaggaacagaccgaacgcggccgccgcctgagcgaagaagaactgctggaa gcgaccgaacgcattaacctgttttatacccagaacaccagcctgctgagccagaaaaaa aaactggaagcggatgtggcgcgcatgcagaaagaagcggaagaagtggtgcaggaatgc cagaacgcggaagaaaaagcgaaaaaagcggcgattgaagcggcgaacctgagcgaagaa ctgaaaaaaaaacaggataccattgcgcatctggaacgcacccgcgaaaacatggaacag accattaccgatctgcagaaacgcctggcggaagcggaacagatggcgctgatgggcagc cgcaaacagattcagaaactggaaagccgcgtgcgcgaactggaaggcgaactggaaggc gaaattcgccgcagcgcggaagcgcagcgcggcgcgcgccgcctggaacgctgcattaaa gaactgacctatcaggcggaagaagataaaaaaaacctgagccgcatgcagacccagatg gataaactgcagctgaaagtgcagaactataaacagcaggtggaagtggcggaaacccag gcgaaccagtatctgagcaaatataaaaaacagcagcatgaactgaacgaagtgaaagaa cgcgcggaagtggcggaaagccaggtgaacaaactgaaaattaaagcgcgcgaatttggc aaaaaagtgcaggaagaa
I used https://www.bioinformatics.org/sms2/rev_trans.html to reverse translate the protein sequence back to the nucleotides.
3.3. Codon optimization Codon optimization is the process of changing the DNA sequence of a gene so that it is expressed more efficiently in a specific organism without changing the protein it produces. I used https://www.idtdna.com/CodonOpt to do codon optimization. I used the reverse translated sequence from 3.2 step, because I was interested in that sequence. I want to continue the process and went for codon optimizaton of the reverse translated sequence.
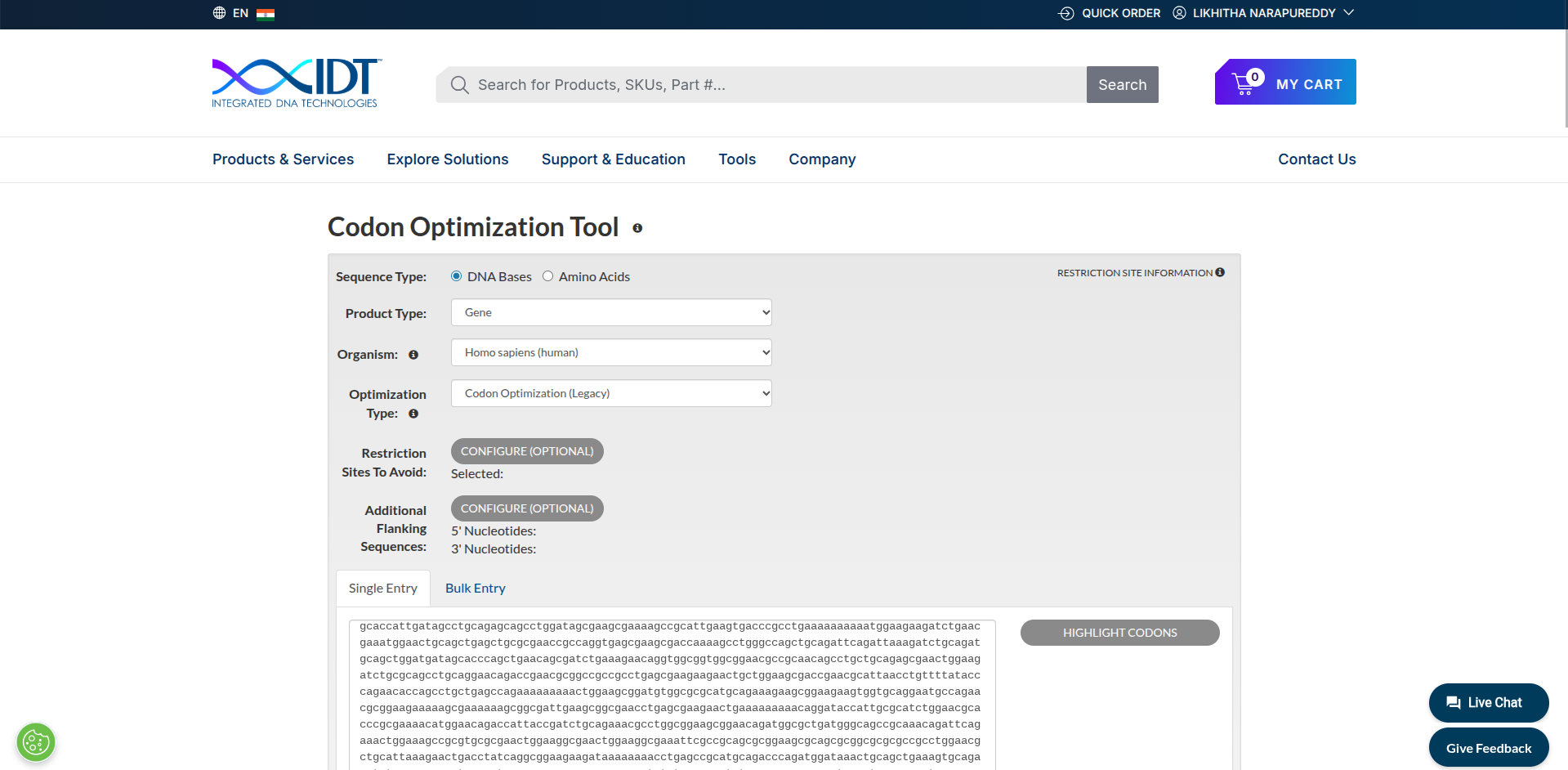

Myosin protein DNA-seq with codon optimization


3.4. You have a sequence! Now what?
After obtaining the myosin protein sequence from UniProt and reverse translating it into a DNA coding sequence, the gene can be optimized for expression in a suitable host such as Escherichia coli or mammalian cells. The optimized gene is inserted into an expression plasmid under a strong promoter. Once introduced into the host cells, RNA polymerase transcribes the DNA into mRNA. The ribosome then translates the mRNA into the myosin polypeptide by reading codons and assembling the corresponding amino acids. After translation, the protein folds into its functional three-dimensional structure and can be purified using affinity chromatography techniques.
Part 4: Prepare a Twist DNA Synthesis Order
I successfully logged in both Twist and benchling. I followed the steps mentioned on the week2 homework site, mapped the sequence, and then completed the annotation(4.2)
Here are my results for the step 4.2:
The image depicts the linear map of the mapped sequence
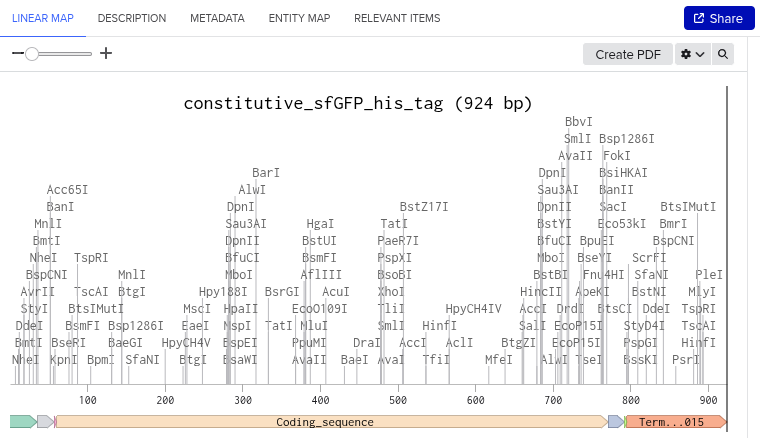

Subsequently, I uploaded the downloaded data in FASTA format to Twist. I selected the clonal genes option and uploaded the file. I then chose the pTwist Amp High Copy vector and dowloaded the resulting sequence. Later, I uploaded the downloaded data from Twist into Benchling. Resulted in creation of a ‘beautiful’ plasmid construct
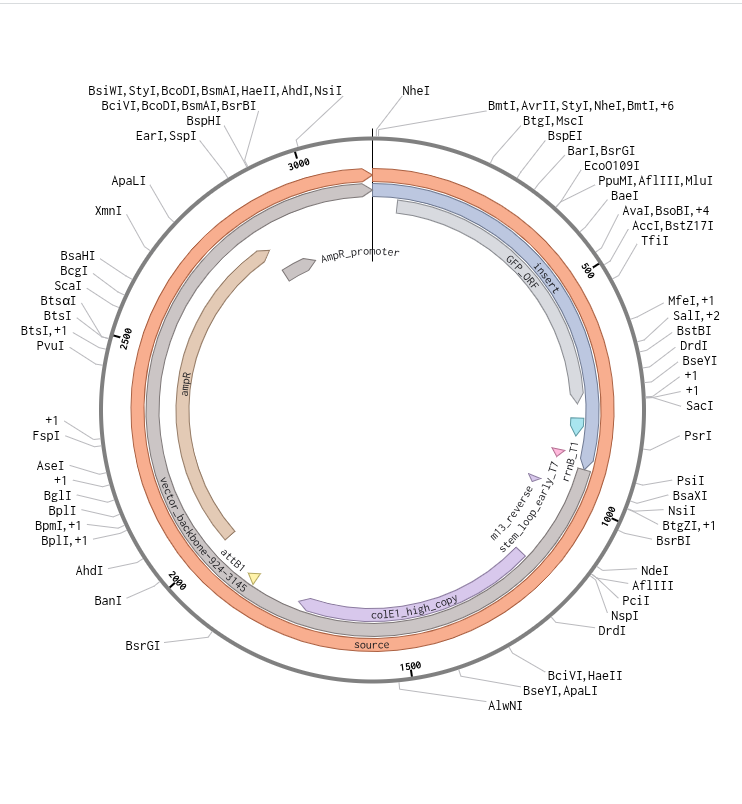

Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why?
For DNA sequencing, I would focus on microalgae and fungi that have strong potential for carbon capture and air purification. Species such as Chlorella vulgaris, Spirulina platensis, and Aspergillus niger are promising because they can absorb carbon dioxide, tolerate environmental stress, and in some cases degrade pollutants.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why? To perform sequencing, I would use Next-Generation Sequencing (NGS), specifically short-read sequencing developed by Illumina.
Also answer the following questions:
1.Is your method first-, second- or third-generation or other? How so?
This is a second-generation sequencing technology because it performs massively parallel sequencing of millions of DNA fragments simultaneously using sequencing-by-synthesis chemistry.
2.What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps.
- DNA extraction
- Fragmentation
- Adapter ligation
- PCR amplification
- Load library onto flow cell
- Software converts color signals → A, T, C, G (base calling)
3.What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)?
- DNA fragments bind to flow cell
- Bridge amplification creates clusters
- Fluorescently labeled nucleotides are added
- Each base emits a specific color signal
- A camera records the fluorescence
- Software converts color signals → A, T, C, G (base calling)
4.What is the output of your chosen sequencing technology?
- FASTQ files (raw reads with quality scores)
- Millions of short reads (150–300 bp)
- After assembly → full genome sequence
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?
For DNA synthesis, I would design a synthetic gene cassette to enhance carbon capture efficiency in microalgae. The construct would include a strong promoter, ribosome binding site, an optimized RuBisCO gene, and a terminator. Additional genes for stress tolerance or pollutant degradation could also be incorporated. The goal would be to create a genetic circuit that increases CO₂ fixation and improves survival in polluted environments.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
DNA synthesis would be performed using commercial gene synthesis services such as Twist Bioscience. The process involves digital DNA design, chemical synthesis of short oligonucleotides, assembly into a full-length gene, error correction, cloning into a plasmid, and sequence verification. Limitations include higher costs for long sequences, challenges with GC-rich regions, and potential synthesis errors. However, synthetic DNA enables precise control over gene design and optimization.
5.3 DNA Edit
(i) What DNA would you want to edit and why?
To further improve air filtration capabilities, I would edit the genomes of selected algae or fungi to enhance carbon fixation, increase pollutant tolerance, or remove metabolic bottlenecks. Genome editing could allow insertion of stronger promoters, modification of enzyme efficiency, or deletion of growth-limiting genes.
(ii) What technology or technologies would you use to perform these DNA edits and why?
The preferred editing tool would be CRISPR-Cas9. This system uses a guide RNA to direct the Cas9 enzyme to a specific DNA sequence, where it introduces a double-strand break.
Also answer the following questions:
- How does your technology of choice edit DNA? What are the essential steps?
a.Design guide RNA (gRNA) targeting gene b.Deliver Cas9 + gRNA into cell c.Cas9 creates double-strand break d.Cell repairs break: NHEJ → knockout and HDR → precise insertion
- What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
Inputs Required a.Guide RNA b.Cas9 protein or plasmid c.Donor DNA template (for HDR) d.Host cells e.Transformation method
- What are the limitations of your editing methods (if any) in terms of efficiency or precision?
Limitations include potential off-target mutations, variable editing efficiency, and delivery challenges in certain microalgal species. Ecological and regulatory considerations must also be addressed before environmental deployment.
Conclusion By integrating DNA sequencing, synthesis, and genome editing, it is possible to design and engineer enhanced biological air filtration systems. Sequencing reveals the natural genetic toolkit of algae and fungi, synthetic DNA enables rational design of improved pathways, and CRISPR-based editing allows precise genome modifications. Together, these technologies provide a powerful framework for developing sustainable, living solutions to air pollution and climate change mitigation.
References
1.Kumar, P., Arora, K., Chanana, I., Kulshreshtha, S., Thakur, V., & Choi, K.-Y. (2023). Comparative study on conventional and microalgae-based air purifiers: Paving the way for sustainable green spaces. Journal of Environmental Chemical Engineering, 11(6), 111046. https://doi.org/10.1016/j.jece.2023.111046
2.Marycz, M., Brillowska-Dąbrowska, A., Muñoz, R., Gębicki, J., et al. (2021). A state of the art review on the use of fungi in biofiltration to remove volatile hydrophobic pollutants. Reviews in Environmental Science and Bio/Technology. https://doi.org/10.1007/s11157-021-09608-7
Week 3 HW: lab automation
My art for Opentrons Artwork
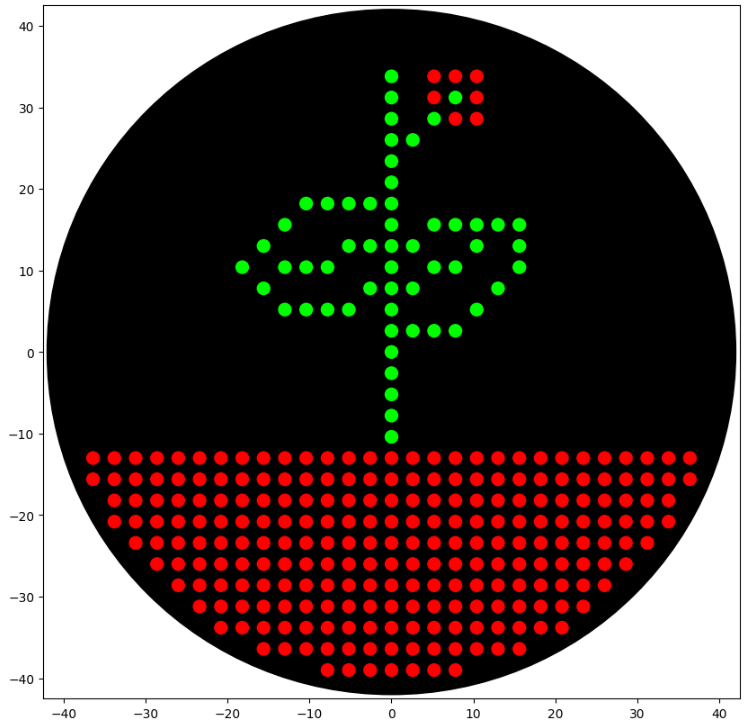

Output of the python script.
PAPER - Semiautomated Production of Cell-Free Biosensors
Journal: ACS Synthetic Biology (2025)
PMID: 40073441
Biosensors are biological systems that detect specific chemicals for example, if a substance is present, they might change color or glow. These can be used for:
- Environmental detection (e.g., fluoride in water)
- Health diagnostics
- Rapid point-of-need testing
But traditionally, making lots of biosensor reactions by hand is slow and inconsistent. Different people might mix things slightly differently, which leads to variability in performance.
Instead of assembling all the biosensor reactions manually, the researchers used a robotic liquid-handling platform (like Opentrons OT-2) to semi-automate the process:
- They wrote a protocol so the robot could prepare many reactions systematically
- They tested this by building a full 384-well plate of biosensors that detect fluoride
- They compared how well these robot-assembled reactions worked compared with manually assembled one
The robot-assembled biosensors worked as expected
The perks of the automated robot:OT-2
- Using robots makes it possible to produce many biosensors quickly and reliably
- This reduces human error when preparing them
- It helps scale up manufacturing or testing so the sensors can be widely deployed
Idea 1: Carbon Capturing Microbial Genomics
What I would automate: High Throughput Strain Screening I might have:
- Environmental isolates
- Engineered variants
- Promoter strengths
Total = n number of combinations
By using automation I can :
- Screen many strains simultaneously
- Maintain equal CO₂ exposure conditions
- Reduce pipetting variation
- Generate reproducible comparative data
Idea 2: Radiation Resistant Bio-Fabric for Space Habitat Walls
What I Would Automate: Stress Testing Simulation Automation could:
- Dispense engineered strains into plates
- Add oxidative stress chemicals (radiation mimic)
- Add ROS indicators
- Incubate
- Measure fluorescence or survival
This allows parallel radiation resistance testing.
Idea 3: Living Anti-Corrosion System for Ocean Infrastructure
What I Would Automate: Corrosion Sensor Screening
- Test combinations of: -> Iron responsive promoters -> pH sensitive promoters -> Mineral producing enzymes
- Salt Stress Optimization Automation could: -> Dispense increasing NaCl concentrations -> Add corrosion mimics -> Measure biofilm formation -> Quantify calcium carbonate output
Week 4 HW: Protein design part I
Part A. Conceptual Questions
- How many amino acid molecules are in 500 g of meat?
Ans: Average amino acid ≈ 100 Daltons (100 g/mol)
500 g ÷ 100 g/mol = 5 moles
1 mole = 6.022 × 10²³ molecules
So,
5 × 6.022 × 10²³ ≈ 3 × 10²⁴ amino acid molecules
- Why do humans eat beef but do not become a cow, eat fish but do not become fish?
Ans: When we eat beef or fish, our body breaks proteins into amino acids during digestion. Then we rebuild them into human proteins, not cow proteins.
- Why are there only 20 natural amino acids?
Ans: There are only 20 amino acids because-
-> The genetic code evolved to encode these efficiently
-> They provide enough chemical diversity (charge, size, polarity)
-> Evolution kept what worked best.
- Can you make other non-natural amino acids? Design some new amino acids.
Ans: Yes, scientists can synthesize new amino acids. Example design:
-> Add a fluorescent group → to track proteins.
-> Add a metal-binding group → to create catalytic proteins.
- Where did amino acids come from before enzymes that make them, and before life started?
Ans: They likely formed:
->In the early Earth atmosphere (like in the Miller–Urey experiment)
->In hydrothermal vents
->Delivered by meteorites
Amino acids can form naturally without life.
- If you make an α-helix using D-amino acids, what handedness (right or left) would you expect?
Ams: Natural L-amino acids form right-handed helices.
D-amino acids would form a left-handed α-helix.
- Why are most molecular helices right-handed?
Ans: Most molecular helices are right-handed because life uses L-amino acids, and their 3D geometry makes right-handed helices the most stable structure.
- Why do β-sheets tend to aggregate?
Ans: They naturally stack into large sheets. Because:
-> They form many hydrogen bonds
-> Hydrophobic regions stick together
- Why do many amyloid diseases form β-sheets? Can you use amyloid β-sheets as materials?
Ans: In diseases like Alzheimer’s disease and Parkinson’s disease:
->Normal proteins misfold.
->Instead of staying flexible, they rearrange into β-sheet structures.
->β-sheets are very stable because they form many hydrogen bonds.
->These sheets stack together into long fibers called amyloids.
Yes, one can use amyloid β-sheets as materials. Although harmful in disease, amyloid β-sheets have useful properties:
-> Very strong (like silk)
-> Self-assembling
-> Chemically stable
-> Nanoscale fibers
Part B: Protein Analysis and Visualization
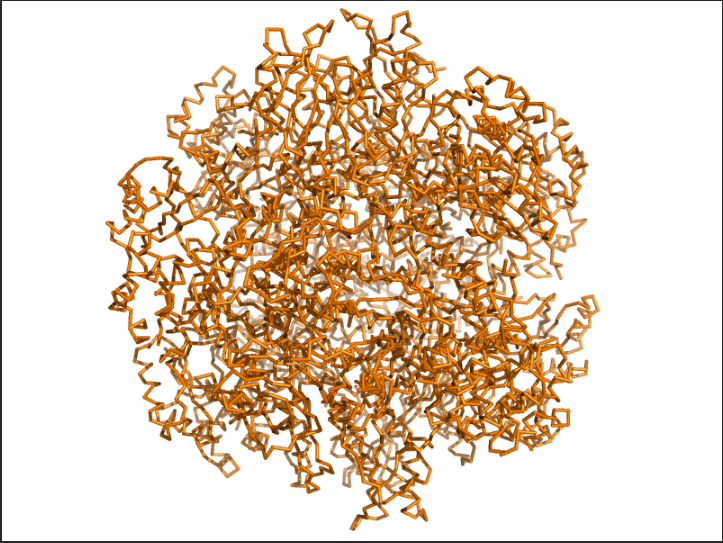
- Briefly describe the protein you selected and why you selected it.
The protein I selected was RuBisCo from Thermosynechococcus vestitus . RuBisCO (Ribulose-1,5-bisphosphate carboxylase/oxygenase) is the key enzyme responsible for carbon fixation in photosynthesis.RuBisCO catalyzes the first major step of the Calvin cycle:
-> It adds carbon dioxide (CO₂) to ribulose-1,5-bisphosphate (RuBP).
-> This reaction produces two molecules of 3-phosphoglycerate (3-PGA).
This process allows plants, algae, and cyanobacteria to convert atmospheric CO₂ into organic molecules (sugars).
Identify the amino acid sequence of your protein.
How long is it? What is the most frequent amino acid? You can use this Colab notebook to count the frequency of amino acids.
Ans: The length is 475 amino acids , the most frequent amino acid is G. G occured 74 times in the sequence.
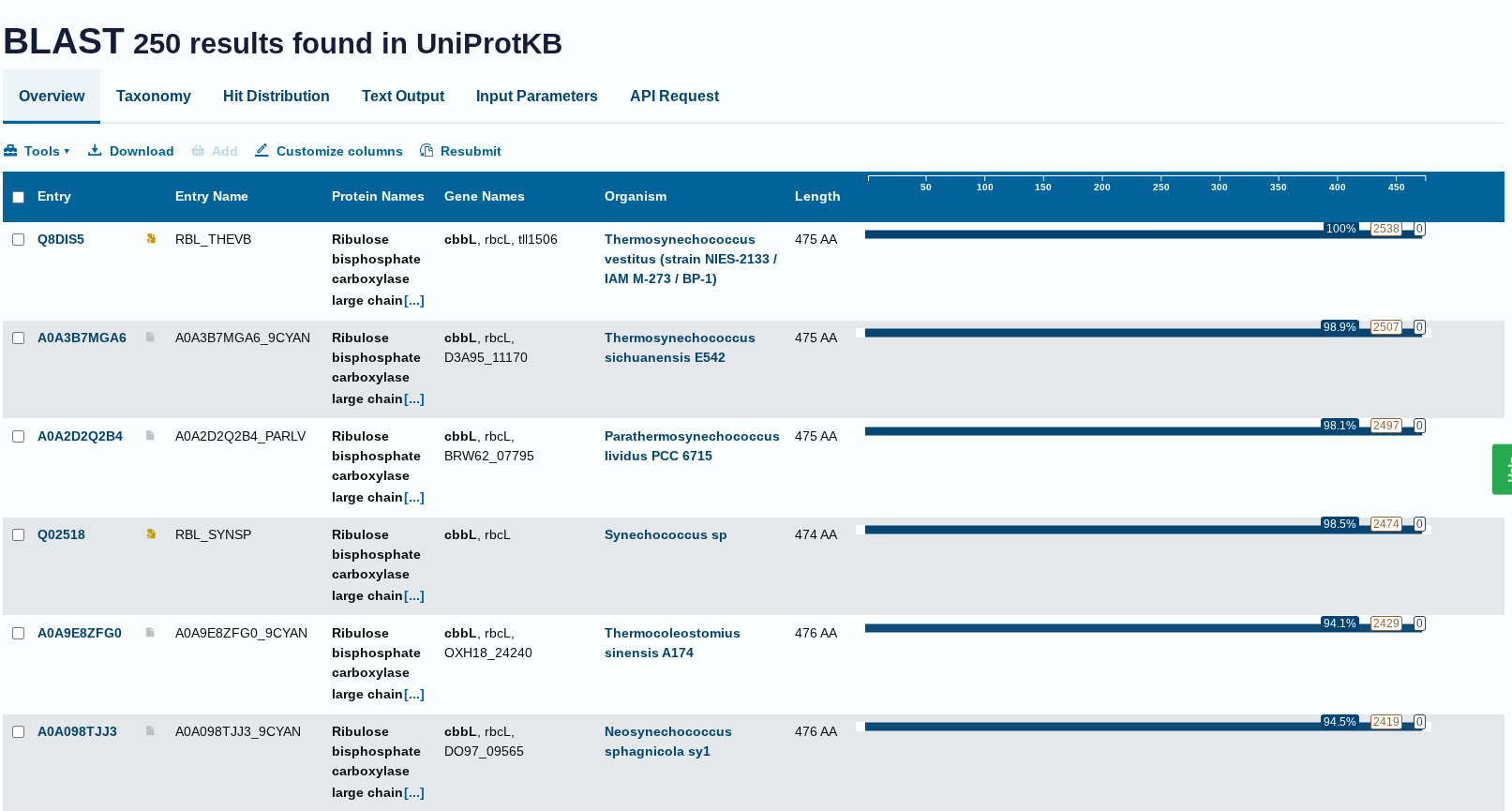
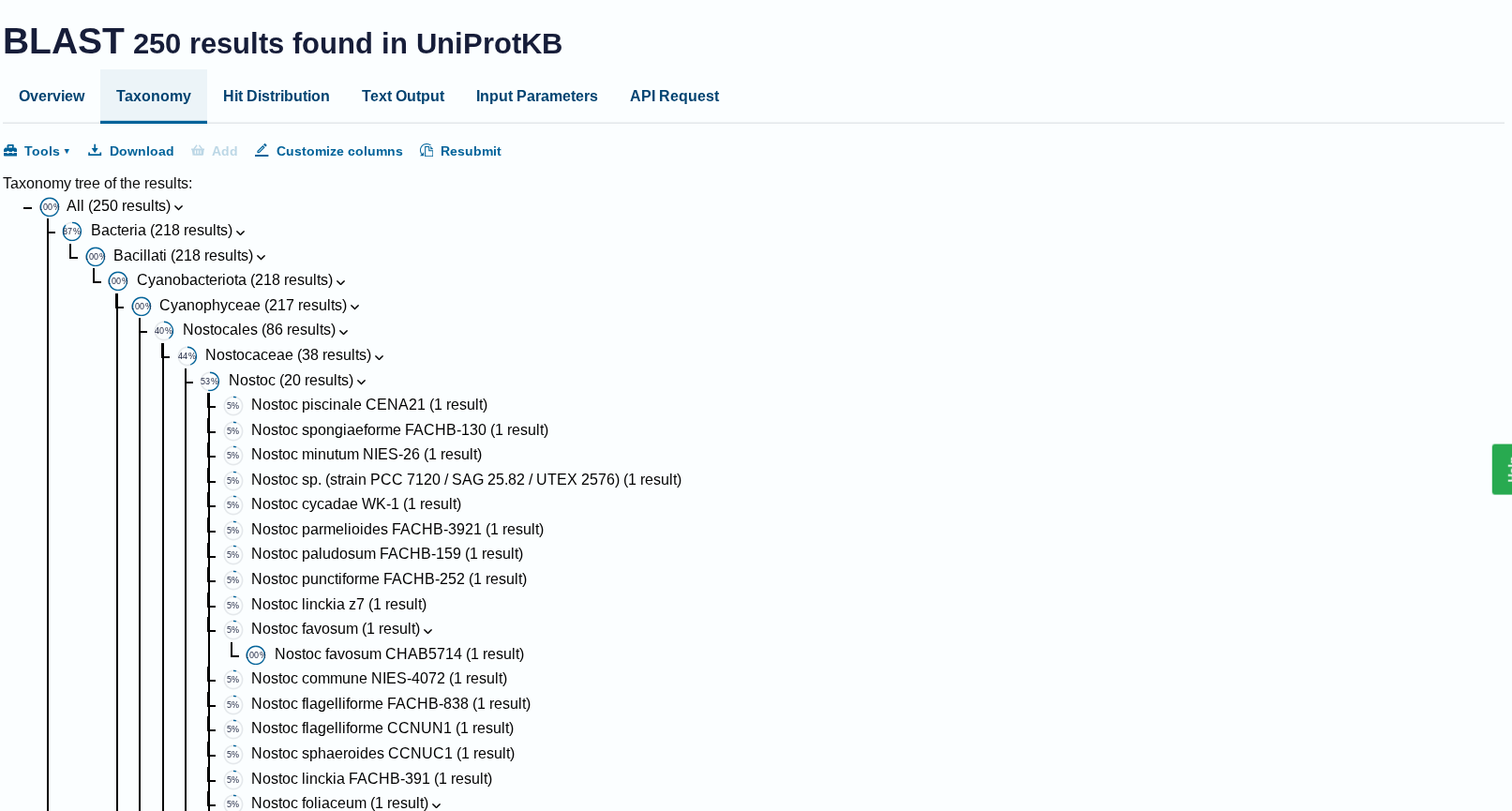
How many protein sequence homologs are there for your protein? Hint: Use Uniprot’s BLAST tool to search for homologs.
Ans: There 250 homologs to the protein- RuBISCo
Does your protein belong to any protein family?
Ans: Belongs to the RuBisCO large chain family. Type I subfamily.
Identify the structure page of your protein in RCSB
When was the structure solved? Is it a good quality structure? Good quality structure is the one with good resolution. Smaller the better (Resolution: 2.70 Å)
Ans: The structure was experimentally solved using X-ray diffraction data collected on February 4, 2007.
So the timeline is:
->Data collected (structure solved): 2007-02-04
->Deposited to Protein Data Bank: 2011-03-10
->Released publicly: 2012-03-28
Yes, it is a good quality structure, the resolution is 2.30 Å
Are there any other molecules in the solved structure apart from protein?
Ans: Yes, there are Cl ligand present in the solved structure. Along with the ligand water molecules and ions are present, these comes under hetero molecules.
Does your protein belong to any structure classification family?
Ans: Yes, my protein is structurally classified and is part of the RuBisCO small subunit structural family.
Open the structure of your protein in any 3D molecule visualization software:
PyMol Tutorial Here (hint: ChatGPT is good at PyMol commands)
Visualize the protein as “cartoon”, “ribbon” and “ball and stick”.
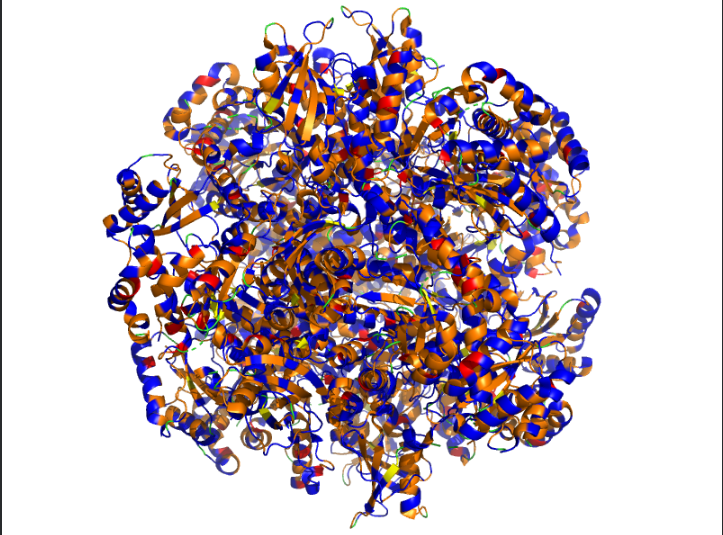
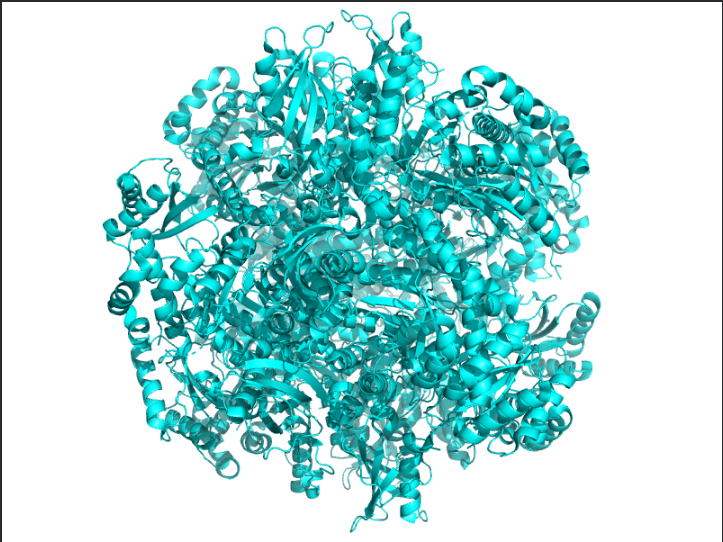
Ans: The protein structure was visualized using cartoon, ribbon, and ball-and-stick representations. The cartoon and ribbon models highlight the overall fold and secondary structural elements, while the ball-and-stick model shows atomic-level details of the protein structure.
Color the protein by secondary structure. Does it have more helices or sheets?
Ans: The protein is predominantly β-sheet(yellow) rich with fewer α-helices(red).
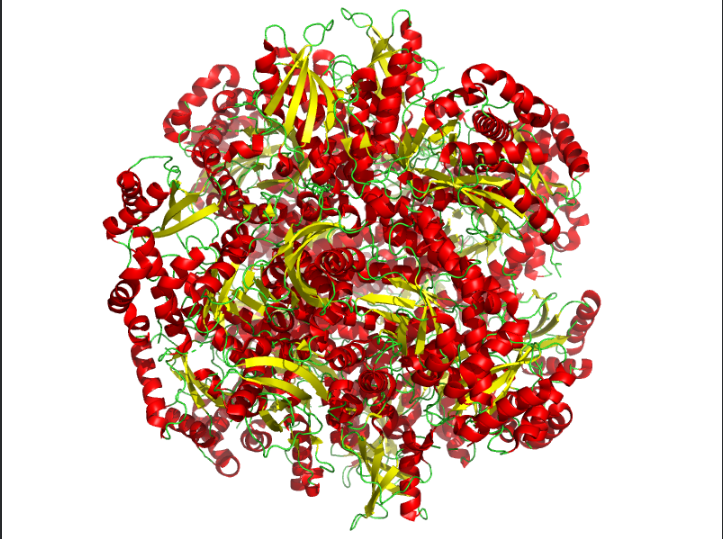

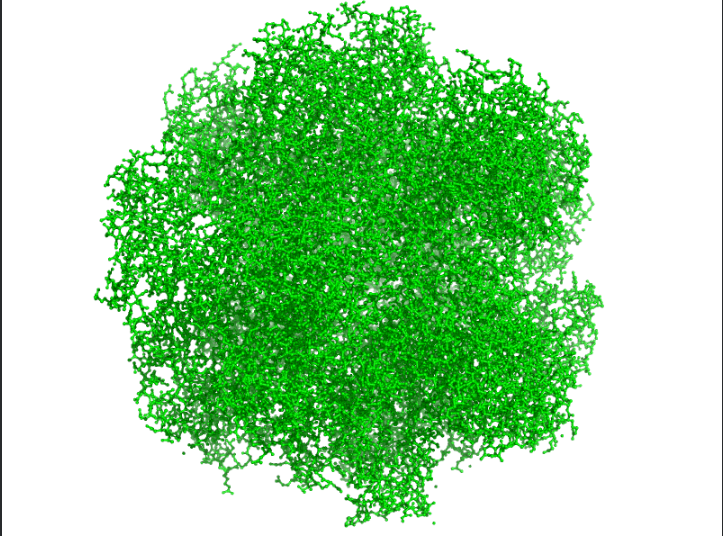
Color the protein by residue type. What can you tell about the distribution of hydrophobic vs hydrophilic residues?
Ans: Hydrophobic(orange) residues are buried in the core, while hydrophilic(yellow) residues are exposed on the surface, consistent with a soluble enzyme.
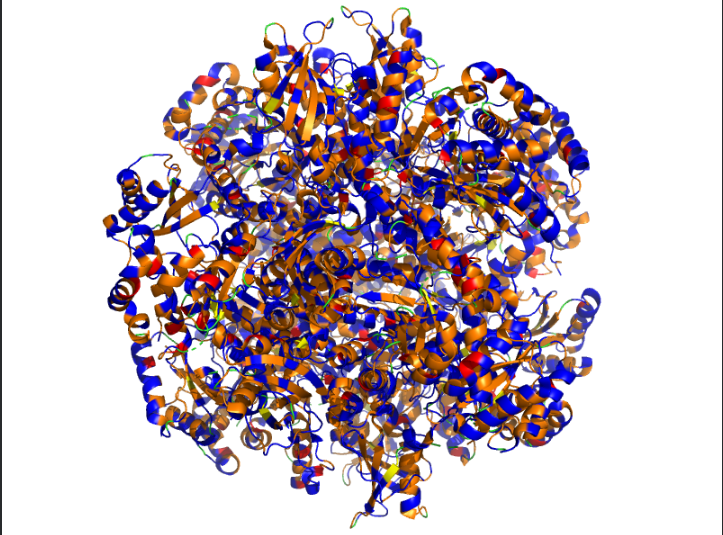


Visualize the surface of the protein. Does it have any “holes” (aka binding pockets)?
Ans: Surface representation reveals shallow binding pockets and structural clefts that likely contribute to substrate binding and subunit interaction.

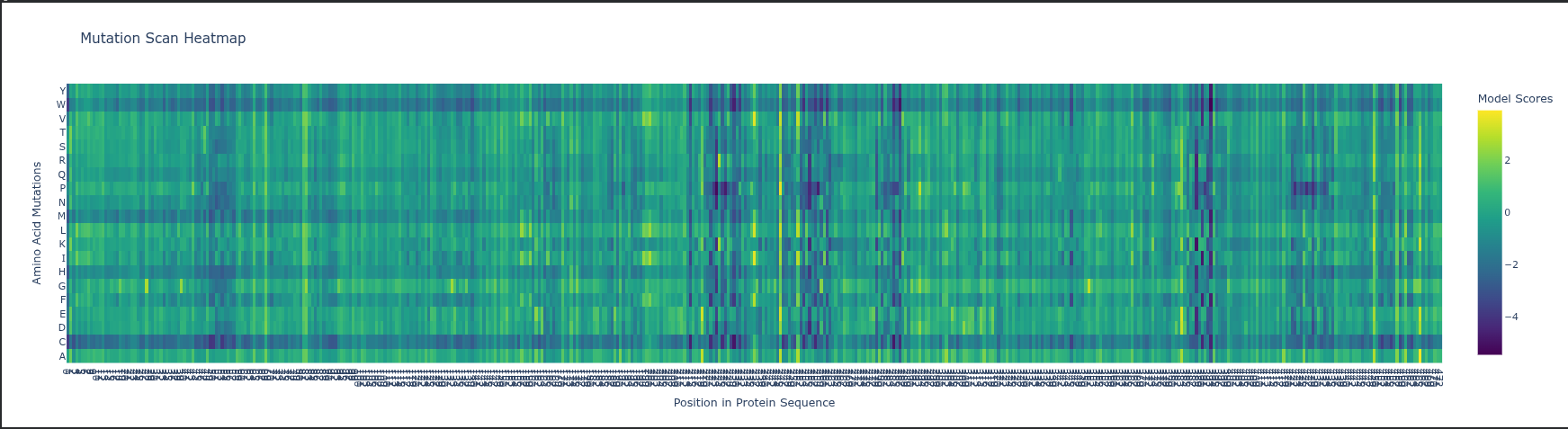
Part C. Using ML-Based Protein Design Tools
C1. Protein Language Modeling
Deep Mutational Scan (DMS) ESM2 Likelihoods: The heatmap predicts how a mutation affects protein fitness by calculating how the model is by a change; high negative values (darker colors) indicate mutations that likely disrupt the protein’s structure or function.
Specific Pattern: Look at Glycine (G) or Proline (P) residues in the sequence; mutations at these sites usually stand out as highly deleterious (darker) because these amino acids have unique structural roles (flexibility or rigid kinks) that other residues cannot easily replace.
Experimental Comparison: In RuBisCO, ESM2 predictions generally correlate strongly with experimental data in the catalytic core, but the model may “under-predict” the impact of mutations in surface loops that are functionally important for protein-protein interactions (like with RuBisCO activase) but less evolutionarily conserved.

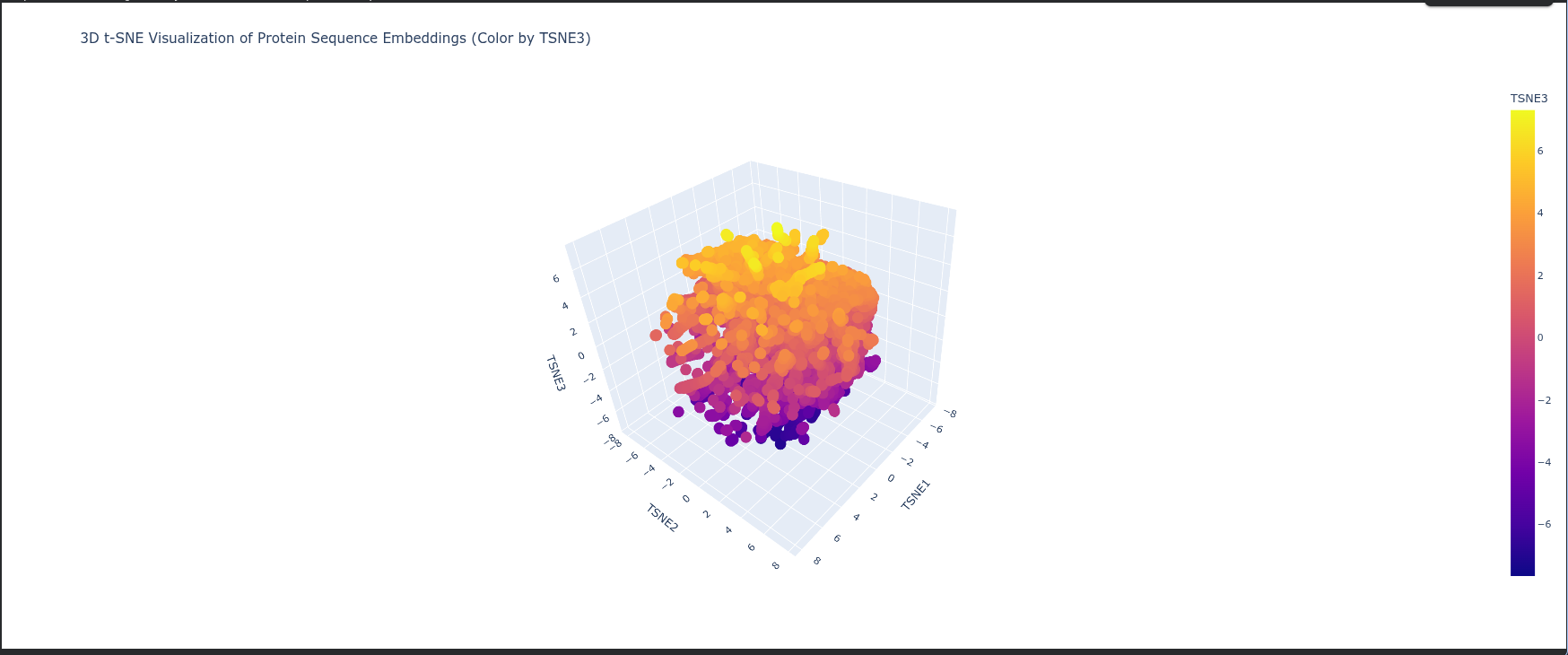
Latent Space Analysis Neighborhoods: The clusters in t-SNE plot represent groups of proteins with similar structural folds and evolutionary origins, meaning proteins in the same neighborhood likely share the same biological pathway or enzymatic mechanism.
Protein Position: The protein is positioned based on its high-dimensional embedding; it likely sits in a dense neighborhood of Type I RuBisCO enzymes, indicating it shares a highly conserved sequence identity and 3D architecture with other photosynthetic large subunits.
Similarity: Proximity to neighbors suggests that ESM2 has successfully captured “hidden” biological rules—such as hydrophobic packing and electrostatic networks—placing the protein near those with the most similar functional constraints.

C2. Protein Folding
Protein Folding Analysis Coordinate Matching: ESMFold predictions generally match original structures closely for well-defined domains, though disordered regions show higher variance between predicted and experimental coordinates.
Structural Resilience: The protein appears highly resilient to single mutations, as most of the heatmap is green (neutral), indicating that the language model expects the overall fold to remain stable despite small changes.
Segment Impact: Large segment deletions or radical mutations in the “dark blue” caused structural collapse, as these regions represent the core stability of the protein.
The image depicts the structure of the protein RuBISCo

C3. Protein Generation
The predicted sequence has low score when compared to the original sequence. Below images show structural difference between predicted sequence and original sequence.


Image depicts the structure of original sequence
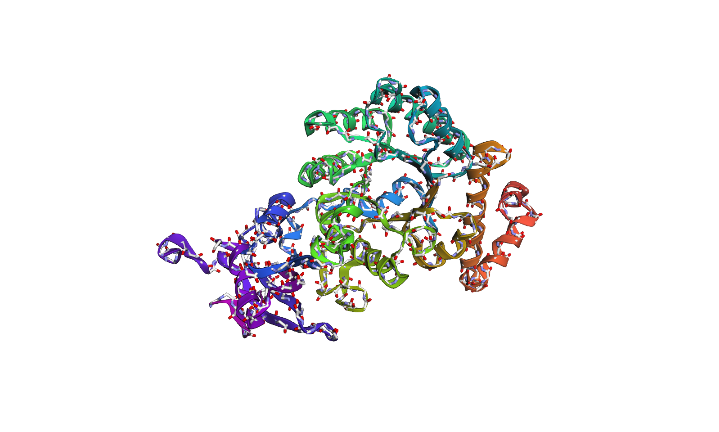

Image depicts the structure of predicted sequence.
Week 5: Protein design part II
Week 6 HW: Genetic Circuits Part I - Assembly Technologies
Week 7 HW: Genetic Circuits partII: Neuromorphic circuits
Assignment Part 1: Intracellular Artificial Neural Networks (IANNs)
1. What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
Traditional genetic circuits mostly behave like Boolean logic gates (ON/OFF). Intracellular Artificial Neural Networks (IANNs) are more flexible.
Advantages:
a. Analog (continuous) behavior
-> Traditional circuits: only 0 or 1 (OFF/ON)
-> IANNs: can process graded inputs (like protein concentration levels). More similar to real biological systems
b. Ability to learn complex patterns
-> Boolean circuits struggle with complex relationships
-> IANNs can approximate nonlinear functions.Useful for detecting subtle biological signals
c. Multivariate decision-making
-> Traditional: limited number of inputs
-> IANNs: integrate multiple inputs simultaneously. Example: detecting disease based on multiple biomarkers
d. Noise tolerance
-> Biological systems are noisy
-> Neural-like circuits can be designed to be robust to fluctuations
e. Scalability
-> Hard to scale Boolean circuits without complexity exploding
->IANNs naturally scale into layers (like perceptrons)
2.Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
Application of an IANN: Microplastic Detection and Filtering System
Description:
An intracellular artificial neural network (IANN) can be engineered in microbial cells to detect and respond to the presence of microplastics in aquatic environments. Since microplastics are not directly sensed biologically, the system relies on indirect chemical and physical signals associated with plastic contamination.
Input Behavior:
The system takes multiple inputs encoded as DNA sensors:
X1: Detects plastic-associated chemicals (e.g., bisphenol-like compounds released from plastics)
X2: Detects hydrophobic particle interactions (via surface-binding proteins)
X3: Detects oxidative stress caused by microplastic exposure
X4: Detects co-contaminants that commonly adsorb onto microplastics
Each input produces a graded transcriptional response, resulting in varying levels of regulatory proteins inside the cell.
These inputs are weighted and integrated by the IANN through regulatory elements such as transcription factors or endoribonucleases, allowing the system to compute the overall likelihood of microplastic contamination.
Output Behavior
The output depends on the combined input signal:
-> When the integrated signal is below threshold → minimal or no response
-> When the integrated signal is above threshold → activation of output genes
Possible outputs: a. Fluorescent protein expression
b. Indicates presence of microplastics (detection mode)
c. Expression of plastic-binding proteins or enzymes (e.g., PET-degrading enzymes)
d. Enables capture or partial degradation of microplastics (filtering mode)
This allows the system to act as a smart biosensor and response unit, activating only when contamination is significant.
Limitations: -> Indirect detection: Microplastics are not directly sensed; accuracy depends on proxy signals
-> Biological noise: Variability in gene expression may affect reliability
-> Slow response time: Transcription and translation processes delay output
-> Environmental safety concerns: Release of engineered microbes into natural ecosystems poses risks
-> Limited degradation efficiency: Biological breakdown of plastics is slow and incomplete
Layer 1 produces an endoribonuclease (Csy4) that negatively regulates fluorescent protein expression in Layer 2 by cleaving mRNA.
Assignment Part 2: Fungal Materials
- What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Ans: i) Examples of fungal materials & their uses
a. Mycelium-based materials
Example: Ecovative Design products
Uses: -> Packaging (alternative to Styrofoam)
-> Building materials (insulation, bricks)
-> Furniture
b. Fungal textiles
Uses:
-> Sustainable fabrics
-> Biodegradable fashion materials
ii) Advantages over traditional counterpart
a. Biodegradable: Break down naturally (unlike plastics)
b.Sustainable: Grown from agricultural waste
c.Low energy production: No high-temperature industrial processes
d.Carbon sequestration: Can store CO₂ during growth
e.Customizable growth: Shape materials during growth
iii) Disadvantages
a. Lower durability: Not as strong as metals or high-grade plastics
b. Moisture sensitivity: Can degrade in humid environments
c.Scaling challenges: Hard to mass-produce consistently
d.Slower production: Growth takes days vs instant manufacturing
e.Limited lifespan: Not ideal for long-term structural use
- What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
Ans: I want to genetically engineer the fungi for the following reasons:
a. Fungi can be engineered to produce materials with improved performance. Because This allows development of lightweight, biodegradable composites with mechanical properties closer to plastics or wood-based materials.
b. Fungi naturally produce extracellular enzymes capable of breaking down complex substrates.Because this expands fungal capability for bioremediation, enabling degradation of persistent materials such as plastics, dyes, and hydrocarbons under mild environmental conditions.
c. Fungal mycelium can be engineered to respond dynamically to environmental stimuli. Because This enables adaptive materials that can self-repair, respond to damage, or change properties in real time.
Advantages of using fungi for synthetic biology vs bacteria
-> Eukaryotic system: Capable of complex protein folding and post-translational modifications, unlike many bacteria such as Escherichia coli
-> Secretion capacity: Efficient export of enzymes and metabolites simplifies downstream processing
-> Mycelial structure: Naturally forms 3D networks, enabling direct fabrication of structured materials
-> Substrate flexibility: Can utilize low-cost feedstocks (e.g., lignocellulosic waste)
Week 9 HW: Cell-Free Systems
Homework Part A: General and Lecturer-Specific Questions
1. Explain the main advantages of cell-free protein synthesis over traditional in vivo methods, specifically in terms of flexibility and control over experimental variables. Name at least two cases where cell-free expression is more beneficial than cell production.
Ans: Cell-free protein synthesis offers significant advantages over in vivo methods due to its open and controllable nature. It allows direct manipulation of reaction components, precise control over parameters such as pH and substrate concentration, and eliminates constraints related to cell viability. As a result, all system resources can be directed toward protein production, enabling rapid optimization and high-throughput experimentation.
CFPS is especially beneficial in cases such as (1) expression of toxic or difficult-to-express proteins, where cellular systems fail, and (2) high-throughput screening and synthetic biology applications, where rapid prototyping without cloning is required.
2. Describe the main components of a cell-free expression system and explain the role of each component.
Ans: A cell-free protein synthesis (CFPS) system contains the essential molecular machinery required for transcription and translation outside living cells.
Key components and roles
a. Cell extract (lysate): Derived from organisms like E. coli, wheat germ, or rabbit reticulocytes
Contains:
Ribosomes → protein synthesis
tRNAs → amino acid delivery
Enzymes → transcription & translation
Role: Core machinery that performs protein production
b. DNA or mRNA template:Encodes the target protein
Can be plasmid DNA or PCR product
Role: Provides genetic instructions for protein synthesis
c. Amino acids
Role: Building blocks for protein formation
d. Energy source system:ATP, GTP + regeneration components
Role: Powers transcription and translation processes
e. Nucleotides (NTPs) ATP, GTP, CTP, UTP
Role: Required for mRNA synthesis during transcription
f. Cofactors and salts Mg²⁺, K⁺, etc.
Role: Maintain optimal enzyme activity and ribosome stability
3. Why is energy provision regeneration critical in cell-free systems? Describe a method you could use to ensure continuous ATP supply in your cell-free experiment.
Ans: Protein synthesis is energy-intensive:
->ATP → transcription + tRNA charging
->GTP → translation (elongation steps)
Without regeneration: a.ATP is rapidly depleted
b.Reaction stops prematurely
c.Protein yield becomes very low
CFPS lacks metabolism, so no natural ATP recycling occurs
Method to ensure continuous ATP supply
Phosphocreatine–creatine kinase system
Addition of Phosphocreatine (energy reservoir) and creatine kinase enzyme
Mechanism:
Phosphocreatine donates phosphate → regenerates ATP from ADP
4.Compare prokaryotic versus eukaryotic cell-free expression systems. Choose a protein to produce in each system and explain why.
Ans: Below table shows the difference between the prokaryotic and eukaryotic cell-free expression systems.
| Feature | Prokaryotic (e.g., E. coli) | Eukaryotic (e.g., wheat germ, rabbit reticulocyte) |
|---|---|---|
| Speed | Fast | Slower |
| Yield | High | Moderate |
| Cost | Low | Expensive |
| Folding | Limited | Better folding |
| PTMs (post-translational modifications) | Minimal | Present (e.g., glycosylation) |
Note: Included the major features for flexible comparison
Protein I choosed.
a. Prokaryotic system → GFP (Green Fluorescent Protein)
-> Simple, no complex modifications needed
-> High yield required
Reason: E. coli CFPS is fast, cheap, and efficient for simple proteins
b. Eukaryotic system → Antibodies
Production of antibodies requires:
-> Proper folding
-> Disulfide bonds
-> Sometimes post-translational processing
As Eukaryotic systems better mimic cellular conditions for complex proteins, one can use Eukaryotic system to produce antibodies.
5.How would you design a cell-free experiment to optimize the expression of a membrane protein? Discuss the challenges and how you would address them in your setup.
Ans: Challenges included in designing a cell-free experiment to optimize the expresion of a membrane protein:
Membrane proteins are:
-> Hydrophobic
-> Prone to aggregation
-> Difficult to fold correctly
Remedies of challenges:
a. Add membrane mimetics: Liposomes or nanodiscs - Detergents (mild, non-denaturing)
Purpose: Provide a membrane-like environment
b. Optimize reaction conditions : By adjusting Mg²⁺, temperature, and redox conditions.
c. Include chaperones: Assist folding and insertion
d. Continuous exchange system (dialysis CFPS): ->Removes toxic byproducts
-> Extends reaction time
6. Imagine you observe a low yield of your target protein in a cell-free system. Describe three possible reasons for this and suggest a troubleshooting strategy for each.
Ans: Low protein yield: causes and troubleshooting
Problem 1: Poor DNA template quality
Reason:
Degraded DNA or weak promoter
Solution:
a.Use high-quality plasmid
b.Optimize promoter and RBS
Problem 2: Energy depletion
Reason:
ATP runs out quickly
Solution:
a. Use efficient regeneration system (e.g., PEP or glucose-based)
b. Optimize energy substrate concentration
Problem 4 : Inhibitory byproducts
Reason:
Accumulation of phosphate or waste
Solution:
Use continuous exchange CFPS
Homework question from Kate Adamala
Week 10 HW: Advanced Imaging & Measurement Technology
Homework: Final Project
For your final project:
Please identify at least one (ideally many) aspect(s) of your project that you will measure. It could be the mass or sequence of a protein, the presence, absence, or quantity of a biomarker, etc.
Please describe all of the elements you would like to measure, and furthermore describe how you will perform these measurements.
What are the technologies you will use (e.g., gel electrophoresis, DNA sequencing, mass spectrometry, etc.)? Describe in detail.
Ans:The primary aspect measured in this project is the biosensor response, observed as a visible color change resulting from the activation of a reporter gene in the presence of plastic associated chemical signals. Additionally, the level of gene expression will be evaluated to understand the strength of the response under different signal conditions. These measurements will be performed using in silico simulation tools, where the behavior of the genetic circuit is modeled to predict activation and output. DNA construct design and validation will be carried out using Benchling, ensuring proper sequence structure and functionality. The simulation of circuit behavior will be conducted using Asimov Kernel.
In a practical setting, these measurements could be further validated using cell-free expression assays and techniques such as spectrophotometry or fluorescence analysis to quantify the output signal.The output of the biosensor is a visible color change, which can be measured quantitatively using spectrophotometry. This technique measures the absorbance of light at specific wavelengths corresponding to the produced color. The intensity of absorbance is directly proportional to the amount of reporter protein expressed.Fluorescence intensity can be measured using a fluorometer. The emitted light intensity corresponds to the level of gene expression. This method provides high sensitivity and allows precise quantification of the biosensor response, especially at low signal concentrations.
Homework: Waters Part I — Molecular Weight
We will analyze an eGFP standard on a Waters Xevo G3 QTof MS system to determine the molecular weight of intact eGFP and observe its charge state distribution in the native and denatured (unfolded) states. The conditions for LC-MS analysis of intact protein cause it to unfold and be detected in its denatured form (due to the solvents and pH used for analysis).
- Based on the predicted amino acid sequence of eGFP (see below) and any known modifications, what is the calculated molecular weight? You can use an online calculator like the one at https://web.expasy.org/compute_pi/ eGFP Sequence: MVSKGEELFTG VVPILVELDG DVNGHKFSVS GEGEGDATYG KLTLKFICTT GKLPVPWPTL VTTLTYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIFF KDDGNYKTRA EVKFEGDTLV NRIELKGIDF KEDGNILGHK LEYNYNSHNV YIMADKQKNG IKVNFKIRHN IEDGSVQLAD HYQQNTPIGD GPVLLPDNHY LSTQSALSKD PNEKRDHMVL LEFVTAAGIT LGMDELYKLE HHHHHH Note: This contains a His-purification tag (HHHHHH) and a linker (the LE before it).
Ans: The calculated molecular weight using https://web.expasy.org/compute_pi/ is 28006.60 Da
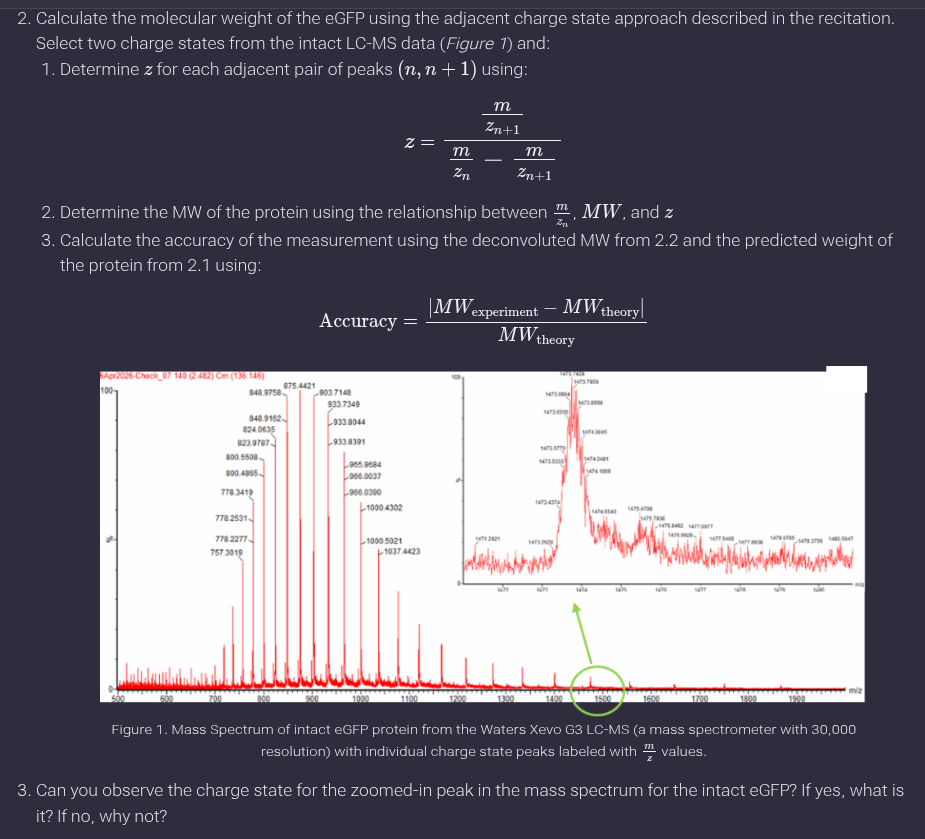

Ans: a. A common pair visible is approximately:
𝑚/𝑧𝑛 ≈ 933
𝑚/𝑧𝑛+1 ≈ 903
Determination of charge state (z):
Two adjacent peaks were selected at approximately 933 and 903 m/z. Using the given formula:
Z = 903/933−903 ≈ 30
Thus, the charge state is z ≈ +30.
b. Molecular weight calculation:
Using the relationship:
𝑀𝑊 = 𝑧(𝑚/𝑧)−𝑧
MW = 30×933−30=27,960 Da
Thus, the molecular weight of eGFP is approximately 27.96 kDa.
c.From earlier:
𝑀𝑊𝑒𝑥𝑝 = 27,960 Da
𝑀𝑊𝑡ℎ𝑒𝑜𝑟𝑦 = 28,006.60 Da
Accuracy calculation:
Accuracy =∣27960−28006.60∣/ 28006.60 ≈ 0.00166
Error ≈ 0.166%
- Charge state of zoomed-in peak:
No, the charge state cannot be directly determined from the zoomed-in peak because isotopic spacing is not resolved, which is necessary to assign charge states.
Homework: Waters Part III — Peptide Mapping - primary structure
- How many Lysines (K) and Arginines (R) are in eGFP? Please circle or highlight them in the eGFP sequence given in Waters Part I question 1 above. (Note: adding the sequence to Benchling as an amino acid file and clicking biochemical properties tab will show you a count for each amino acid).
Ans: Number of Lysines (K) present are 20 and number of Arginines (R) present = 6
MVSKGEELFTG VVPILVELDG DVNGH[K]FSVS GEGEGDATYG [K]LTL[K]FICTT G[K]LPVPWPTL VTTLTYGVQC FS[R]YPDHM[K]Q HDFF[K]SAMPE GYVQE[R]TIFF [K]DDGNY[K]T[R]A EV[K]FEGDTLV N[R]IEL[K]GIDF [K]EDGNILGH[K] LEYNYNSHNV YIMAD[K]Q[K]NG IKVNFKI[R]HN IEDGSVQLAD HYQQNTPIGD GPVLLPDNHY LSTQSALSKD PNE[K][R]DHMVL LEFVTAAGIT LGMDELY[K]LE HHHHHH
2.How many peptides will be generated from tryptic digestion of eGFP?
Navigate to https://web.expasy.org/peptide_mass/
Copy/paste the sequence above into the input box in the PeptideMass tool to generate expected list of peptides.
Use Figure 4 below as a guide for the relevant parameters to predict peptides from eGFP.
Click “Perform the Cleavage” button in the PeptideMass tool and report the number of peptides generated when using trypsin to perform the digest.
Ans: Using the PeptideMass tool with default settings shows 19 peptides because small peptides (<500 Da) are excluded.
- Based on the LC-MS data for the Peptide Map data generated in lab (please use Figure 5a as a reference) how many chromatographic peaks do you see in the eGFP peptide map between 0.5 and 6 minutes? You may count all peaks that are >10% relative abundance.
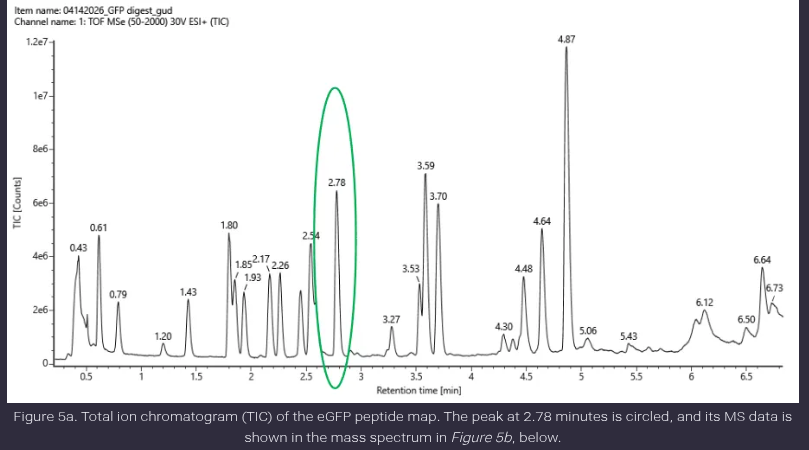

Ans: The number of peptide peaks observed in the chromatogram between 0.5 and 6 minutes, considering only peaks above 10% relative abundance, is approximately 12–14 peaks.
- Assuming all the peaks are peptides, does the number of peaks match the number of peptides predicted from question 2 above? Are there more peaks in the chromatogram or fewer?
Ans: The predicted number of peptides from tryptic digestion is 19 peptides (based on PeptideMass with filtering). The observed number of chromatographic peaks is similar but slightly lower or comparable.
This difference arises because:
Some peptides may not ionize efficiently
Very small or hydrophilic peptides may not be detected
Some peptides may co-elute
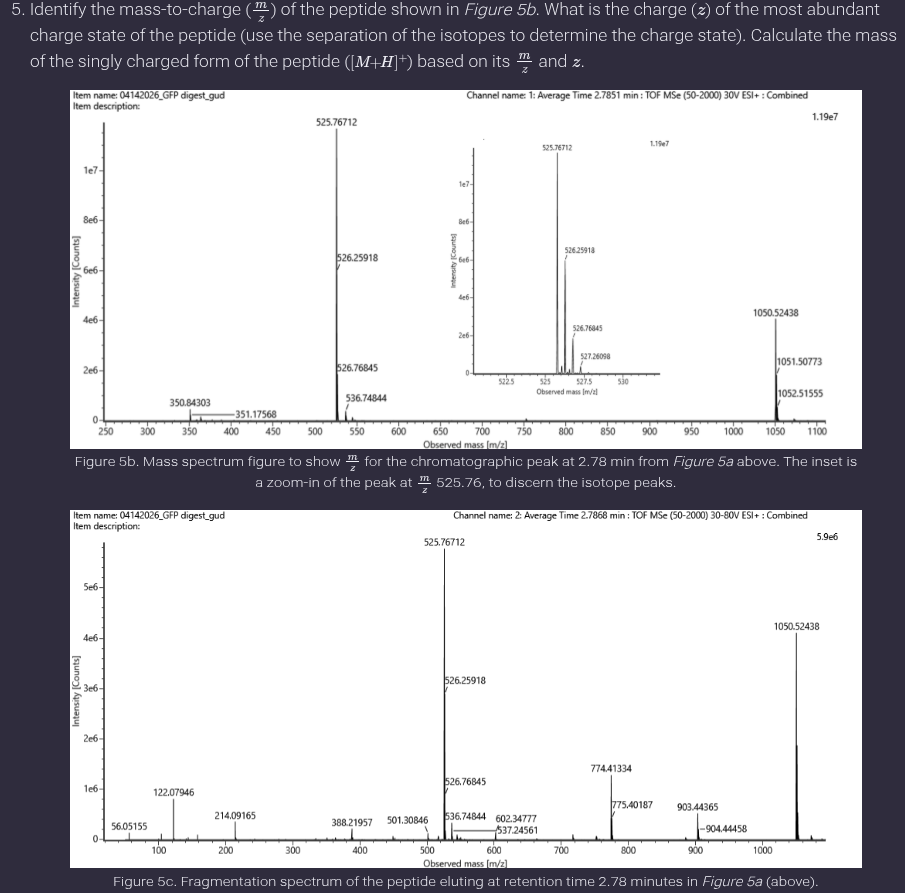

Ans: Charge state and peptide mass
The isotopic peak spacing for the peptide at 2.78 minutes is approximately 0.5 m/z, giving:
𝑧=1/0.5 = 2
Thus, the charge state is +2.
Using:
MW=z(m/z)−z
MW=2×526.27−2.0146 = 1050.525 Da
Therefore, the peptide has:
Charge state (z) = +2
Molecular weight ≈ 1050.5 Da
- Identify the peptide based on comparison to expected masses in the PeptideMass tool. What is mass accuracy of measurement? Please calculate the error in ppm. (Recall that Accuracy from part I)
Ans: Peptide Identification
The peptide is FEGDTLVNR with theoretical mass 1050.5214 Da.
The mass accuracy was calculated using:
1050.525−1050.5214 / 1050.5214 × 10 ^ 6 = 0.0036 / 1050.5214 × 10 ^ 6 ≈3.4ppm
- What is the percentage of the sequence that is confirmed by peptide mapping? (see Figure 6)
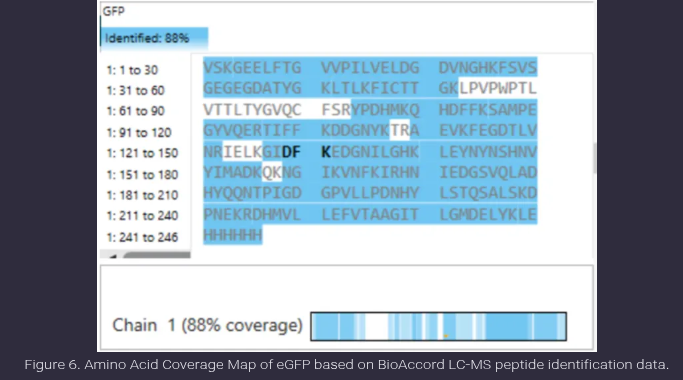


Ans: Peptide mapping confirms 88% of the eGFP sequence, as indicated in Figure 6. This value represents the proportion of the protein sequence that is covered by experimentally identified peptides in the LC-MS analysis. The highlighted regions correspond to detected peptides, while the unhighlighted regions indicate portions of the sequence that were not observed, likely due to limitations such as poor ionization or peptide size. This high coverage suggests successful and reliable protein identification.
Bonus Peptide Map Questions
- Can you determine the peptide sequence for the peptide fragmentation spectrum shown in Figure 5c? (HINT: Use your results from Question 2 above to match the peptide molecular weight that is closest to that shown in Figure 5b. Copy and paste its sequence into this tool online to predict the fragmentation pattern based on its amino acid sequence: http://db.systemsbiology.net/proteomicsToolkit/FragIonServlet.html. What is the sequence of the eGFP peptide that best matches the fragmentation spectrum in Figure 5c?
Ans: The peptide sequence that best matches the fragmentation spectrum in Figure 5c is FEGDTLVNR.This identification is based on matching the experimentally determined molecular weight (~1050.5 Da) with the theoretical peptide masses from the PeptideMass tool. The peptide FEGDTLVNR (1050.5214 Da) shows the closest agreement.
- Does the peptide map data make sense, i.e. do the results indicate the protein is the eGFP standard? Why or why not? Consult with Figure 6, which depicts the % amino acid coverage of peptides positively identified using their calculated mass and fragmentation pattern.
Ans: Yes, the peptide map data is consistent with the protein being the eGFP standard. Figure 6 shows approximately 88% amino acid sequence coverage, indicating that a large majority of the protein sequence has been experimentally confirmed through peptide identification. Additionally, multiple peptides across different regions of the sequence were identified and validated using both mass measurements and fragmentation patterns, providing strong evidence for correct protein identification. The remaining 12% of the sequence not covered is likely due to typical limitations such as poor ionization or peptide detectability and does not significantly affect the confidence of identification. Therefore, the results strongly support that the protein analyzed is eGFP.
Homework: Waters Part IV — Oligomers
We will determine Keyhole Limpet Hemocyanin (KLH)’s oligomeric states using charge detection mass spectrometry (CDMS). CDMS single-particle measurements of KLH allow us to make direct mass measurements to determine what oligomeric states (that is, how many protein subunits combine) are present in solution. Using the known masses of the polypeptide subunits (Table 1) for KLH, identify where the following oligomeric species are on the spectrum shown below from the CDMS (Figure 7):
7FU Decamer 8FU Didecamer 8FU 3-Decamer 8FU 4-Decamer
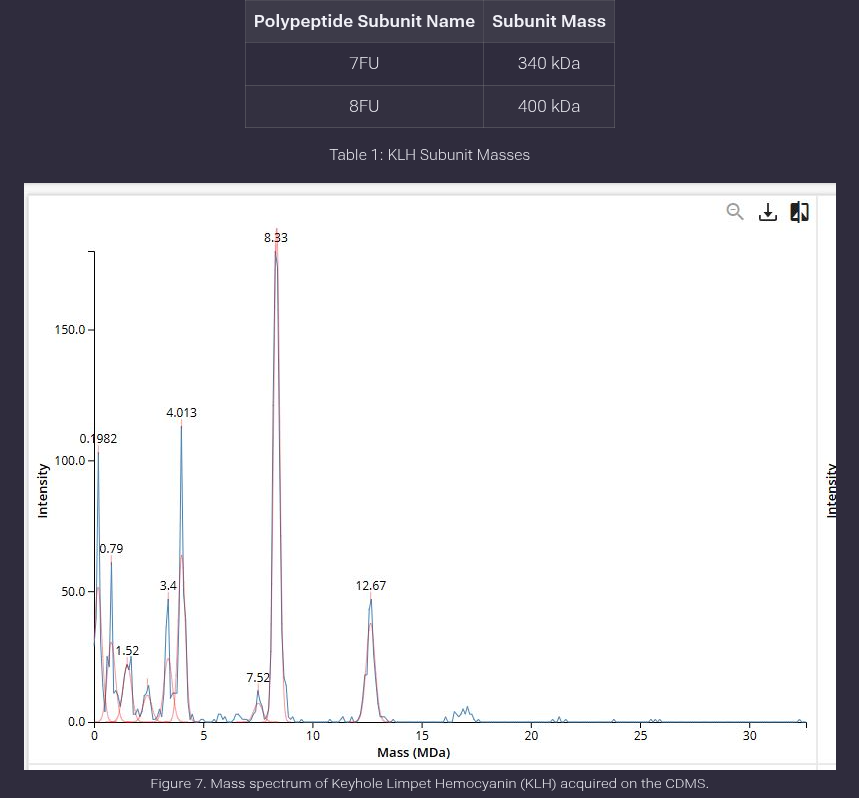

Ans: To identify the oligomeric species of Keyhole Limpet Hemocyanin (KLH) on the provided CDMS spectrum, we calculate the expected mass of each assembly and correlate it with the experimental peaks shown in Figure 7.Theoretical Mass CalculationsSince the x-axis of the spectrum is in Megadaltons (MDa) and the subunit masses are in kilodaltons (kDa), we use the conversion 1,000 kDa = 1 MDa
| Oligomeric Species | Composition | Calculation | Theoretical Mass |
|---|---|---|---|
| 7FU Decamer | 10 × 7FU subunits | 10 × 340 kDa | 3.40 MDa |
| 8FU Didecamer | 20 × 8FU subunits | 20 × 400 kDa | 8.00 MDa |
| 8FU 3-Decamer | 30 × 8FU subunits | 30 × 400 kDa | 12.00 MDa |
| 8FU 4-Decamer | 40 × 8FU subunits | 40 × 400 kDa | 16.00 MDa |
Species Identification on Spectrum (Figure 7) Based on the calculations above, the oligomeric species correspond to the following peaks labeled in the mass spectrum:
7FU Decamer: Assigned to the peak at 3.4 MDa. This matches the theoretical calculation exactly.
8FU Didecamer: Assigned to the highest intensity peak at 8.33 MDa. The slight shift from 8.00 MDa to 8.33 MDa is attributed to native glycosylation and adducts common in large KLH proteins.
8FU 3-Decamer: Assigned to the peak at 12.67 MDa. This represents the assembly of 30 8FU subunits
8FU 4-Decamer: Assigned to the low-intensity cluster of peaks between 16.00 and 17.00 MDa. This corresponds to the 40-subunit assembly.
Homework: Waters Part V — Did I make GFP?
Please fill out this table with the data you acquired from the lab work done at the Waters Immerse Lab in Cambridge, or else the data screenshots in this document if you were unable to have lab work done at Waters.
| Theoretical (kDa) | Observed /Measured on Intact LC-MS (kDa) | PPM Mass Error |
|---|---|---|
| 28.0066 | 27.960 | -1664 ppm |
The relatively high ppm error is due to approximate peak selection and lack of deconvolution in intact protein analysis.
Calculations:
Theoretical MW (from sequence )
28,006.60 Da = 28.0066 kDa
Observed MW (from LC-MS intact protein)
From your earlier intact MS calculation:
≈ 27,960 Da = 27.960 kDa
PPM error calculation
Formula:
PPM = 𝑀𝑊𝑜𝑏𝑠 − 𝑀𝑊𝑡ℎ𝑒𝑜𝑟𝑦 / 𝑀𝑊𝑡ℎ𝑒𝑜𝑟𝑦 × 10 ^ 6
PPM = 27960 −28006.60 / 28006.60 × 10^6
≈−1664ppm
Week 11 HW: Bioproduction and Cloud Labs
Part A: The 1,536 Pixel Artwork Canvas | Collective Artwork
- Contribute at least one pixel to this global artwork experiment before the editing ends on Sunday 4/19 at 11:59 PM EST.
A personalized URL was sent to the email address associated with your Discourse account, and you can discuss the artwork on the Discourse.
If you did not have a chance to contribute, it’s okay, just make sure you become a TA this fall! 😉
- Make a note on your HTGAA webpages including:
what you contributed to the community bioart project (e.g., “I made part of the DNA on the bottom right plate”)
what you liked about the project, and
what about this collaborative art experiment could be made better for next year.
Ans: I made a character red from the game Among Us at the right corner of Q1. This project gave freedom to everyone to create any form of art and I liked it.
Part B: Cell-Free Protein Synthesis | Cell-Free Reagents
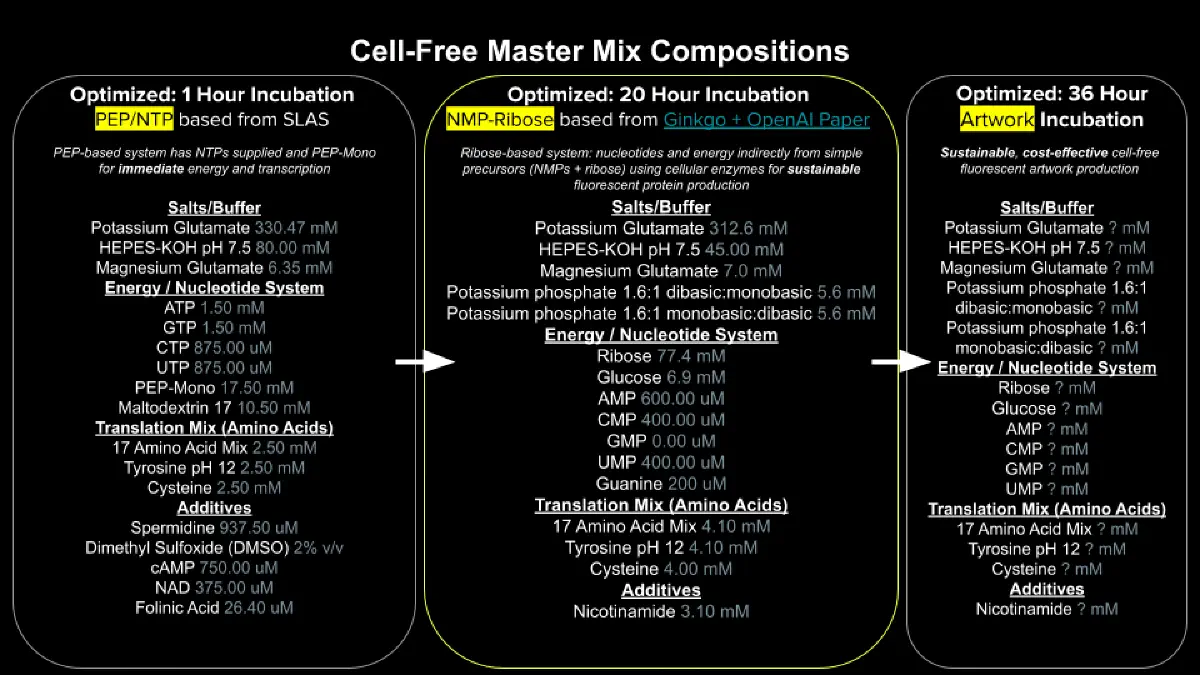

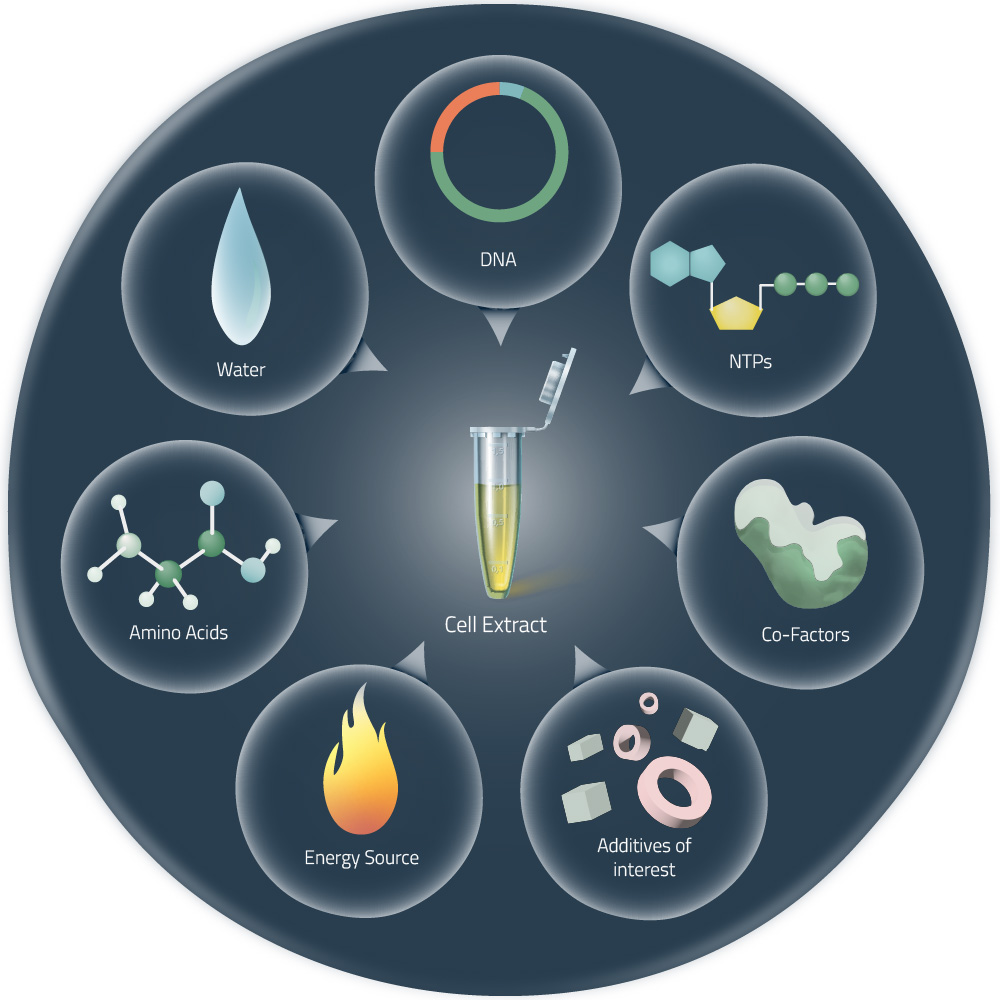
- Referencing the cell-free protein synthesis reaction composition (the middle box outlined in yellow on the image above, also listed below), provide a 1-2 sentence description of what each component’s role is in the cell-free reaction.
Ans: Components & Their Roles
E. coli Lysate
BL21 (DE3) Star Lysate (includes T7 RNA Polymerase) Provides the core cellular machinery—ribosomes, tRNAs, enzymes for transcription and translation; the built-in T7 RNA polymerase drives high-level transcription from T7 promoters.
Salts / Buffer
Potassium Glutamate Mimics intracellular ionic conditions and stabilizes ribosomes and enzymes for efficient protein synthesis.
HEPES-KOH pH 7.5 Maintains a stable physiological pH, which is critical for enzyme activity during transcription and translation.
Magnesium Glutamate Essential cofactor for ribosomes and polymerases; directly impacts translation efficiency and RNA stability.
Potassium Phosphate (Monobasic & Dibasic) Provides buffering capacity and phosphate ions, helping maintain pH balance and supporting energy metabolism.
Energy / Nucleotide System
Ribose Serves as a carbon source for nucleotide regeneration through metabolic pathways.
Glucose Fuels ATP regeneration via glycolytic enzymes present in the lysate, enabling longer reaction lifetimes.
AMP, CMP, GMP, UMP Nucleotide monophosphates that are enzymatically converted into triphosphates (ATP, GTP, etc.) required for transcription and translation.
Guanine A nucleobase precursor that can be salvaged into GMP, contributing to nucleotide pool replenishment.
Translation Mix (Amino Acids)
17 Amino Acid Mix Supplies most of the building blocks required for protein synthesis.
Tyrosine & Cysteine Added separately because they are less stable or more reactive, ensuring sufficient availability during translation.
Additives
Nicotinamide Supports redox balance and metabolic reactions (via NAD⁺/NADH systems), improving energy regeneration and reaction longevity.
Backfill
Nuclease-Free Water Adjusts the final reaction volume while preventing degradation of nucleic acids.
- Describe the main differences between the 1-hour optimized PEP-NTP master mix and the 20-hour NMP-Ribose-Glucose master mix shown in the Google Slide above. (2-3 sentences)
Ans: The PEP-NTP (1-hour) system uses high-energy molecules like phosphoenolpyruvate and directly supplied nucleotide triphosphates, enabling rapid and high protein production but with short reaction lifetimes. The NMP-Ribose-Glucose (20-hour) system relies on slower metabolic regeneration of energy and nucleotides from simpler precursors, resulting in longer-lasting but lower-rate protein synthesis.
- Bonus question: How can transcription occur if GMP is not included but Guanine is?
Ans: Transcription can still occur because guanine can be salvaged into GMP via enzymatic pathways present in the lysate (nucleotide salvage pathways). Once converted to GMP, it can then be phosphorylated to GTP, which is the actual substrate used by RNA polymerase.
Part C: Planning the Global Experiment | Cell-Free Master Mix Design
- Given the 6 fluorescent proteins we used for our collaborative painting, identify and explain at least one biophysical or functional property of each protein that affects expression or readout in cell-free systems. (Hint: options include maturation time, acid sensitivity, folding, oxygen dependence, etc) (1-2 sentences each)
sfGFP
mRFP1
mKO2
mTurquoise2
mScarlet_I
Electra2
Ans: sfGFP
Key property: Robust folding efficiency sfGFP is engineered to fold efficiently even under suboptimal conditions, making it highly reliable in cell-free systems where chaperones may be limited.
mRFP1
Key property: Slow maturation time
mRFP1 takes longer to form its fluorescent chromophore, so signal appears delayed even if protein expression is occurring.
mTurquoise2
Key property: High quantum yield (brightness)
This cyan protein is extremely bright and efficient at emitting light, allowing sensitive detection even at low expression levels.
mScarlet-I
Key property: Rapid maturation and high brightness
mScarlet-I combines fast chromophore formation with strong fluorescence, making it one of the best-performing red proteins in cell-free systems.
Electra2
Key property: Oxygen dependent chromophore formation
Like most fluorescent proteins, Electra2 requires oxygen for chromophore maturation, so limited oxygen in cell-free reactions can reduce or delay fluorescence.
- Create a hypothesis for how adjusting one or more reagents in the cell-free mastermix could improve a specific biophysical or functional property you identified above, in order to maximize fluorescence over a 36-hour incubation. Clearly state the protein, the reagent(s), and the expected effect.
Ans: Hypothesis for Electra2 (maximize fluorescence over 36 hours)
Reducing the glucose concentration in the cell-free mastermix will decrease metabolic oxygen consumption, thereby increasing dissolved oxygen availability for Electra2 chromophore maturation and improving fluorescence over a 36-hour incubation.
Reagent to adjust:
Glucose (energy source in the mastermix)
Expected effect: Lower glucose levels will slow ATP-generating metabolism, reducing oxygen depletion in the reaction. This preserves oxygen needed for Electra2’s oxygen-dependent chromophore formation, leading to a higher fraction of properly matured fluorescent protein and increased overall fluorescence signal after 36 hours.