Week 2 HW: DNA Read, Write, and Edit
Part 1: Benchling & In-silico Gel Art
See this week’s lab protocol “Gel Art: Restriction Digests and Gel Electrophoresis” for details. Overview:
Below are the steps I've done to finished this assignment.
Account at benchling.com √
 Have the .gb file of Lamdba_NEB saved.
Have the .gb file of Lamdba_NEB saved.
 Upload the .gb file to the platform.
Upload the .gb file to the platform.
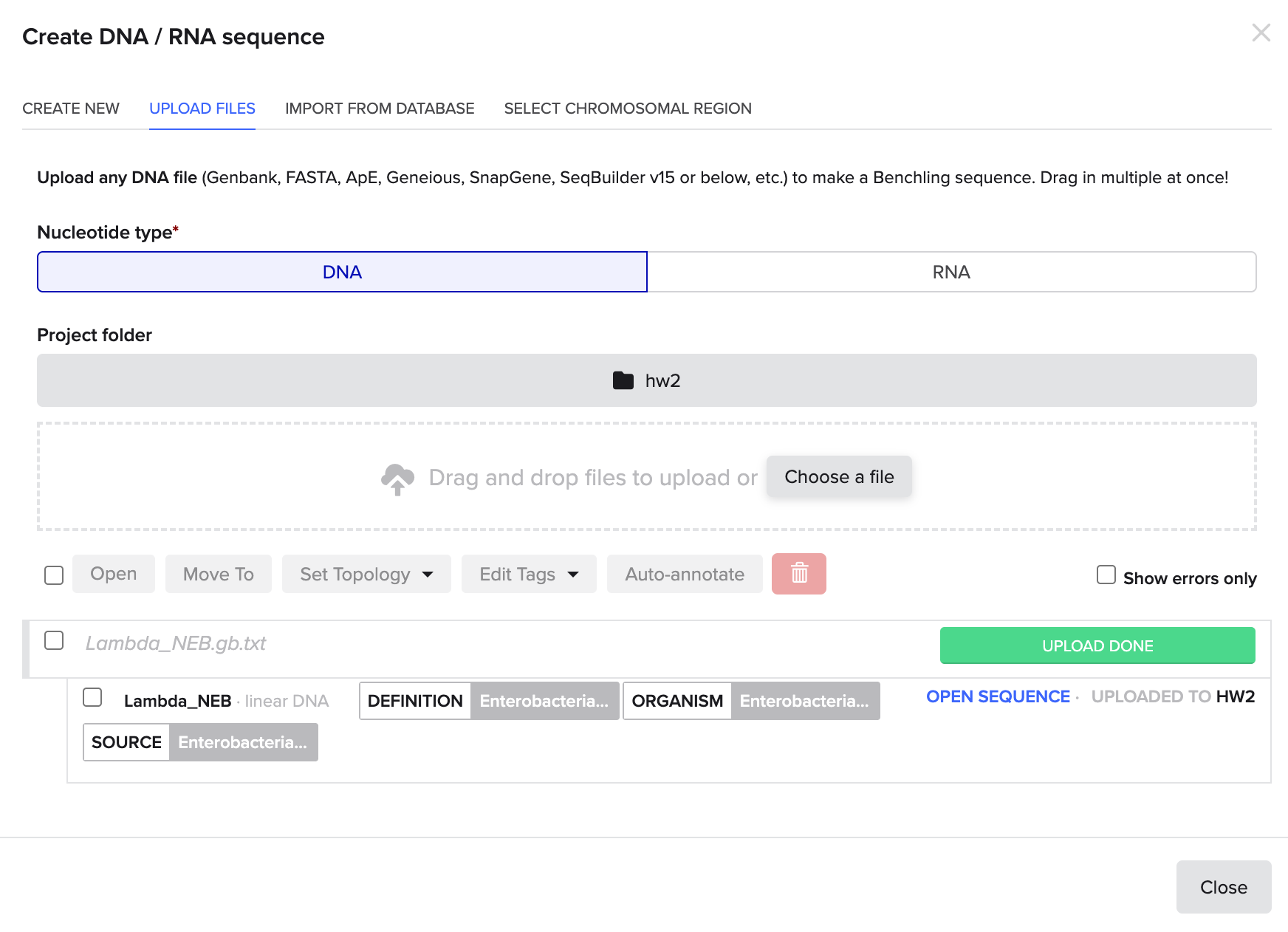 Visual display of the structure of the DNA sequence. Very interesting!
Visual display of the structure of the DNA sequence. Very interesting!
 Start to add a new list of restriction enzymes.
Start to add a new list of restriction enzymes.
 Adding them one by one.
Adding them one by one.
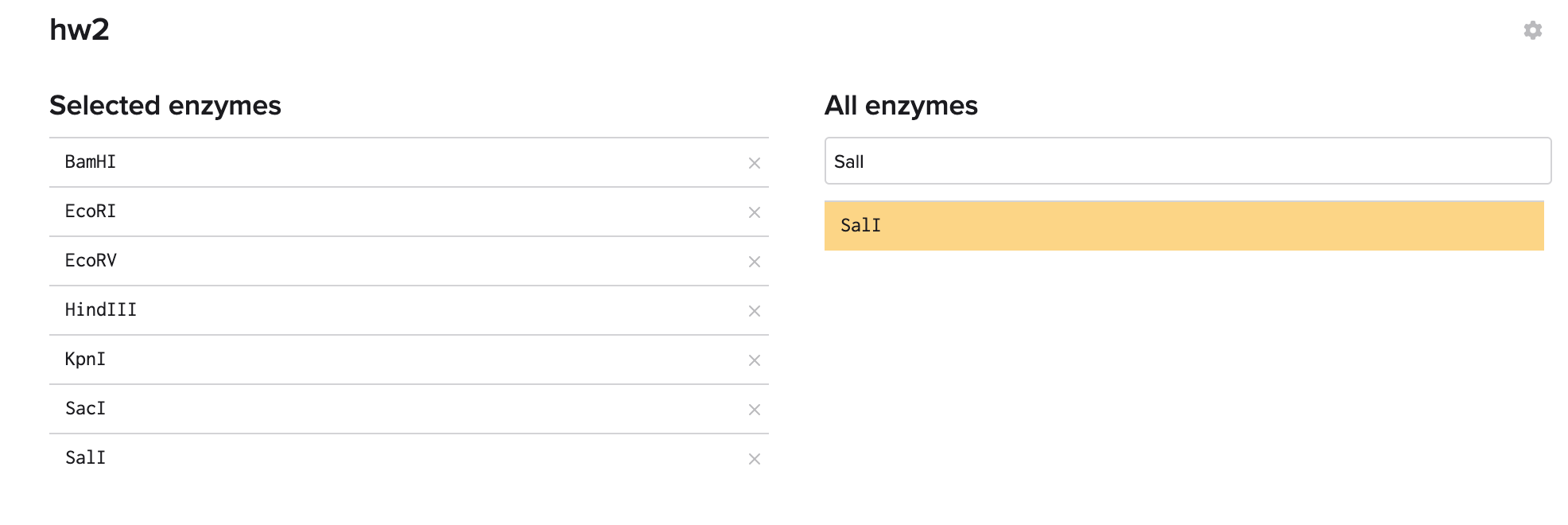 Using the analyze tool to the right to see the result(s).
Using the analyze tool to the right to see the result(s).
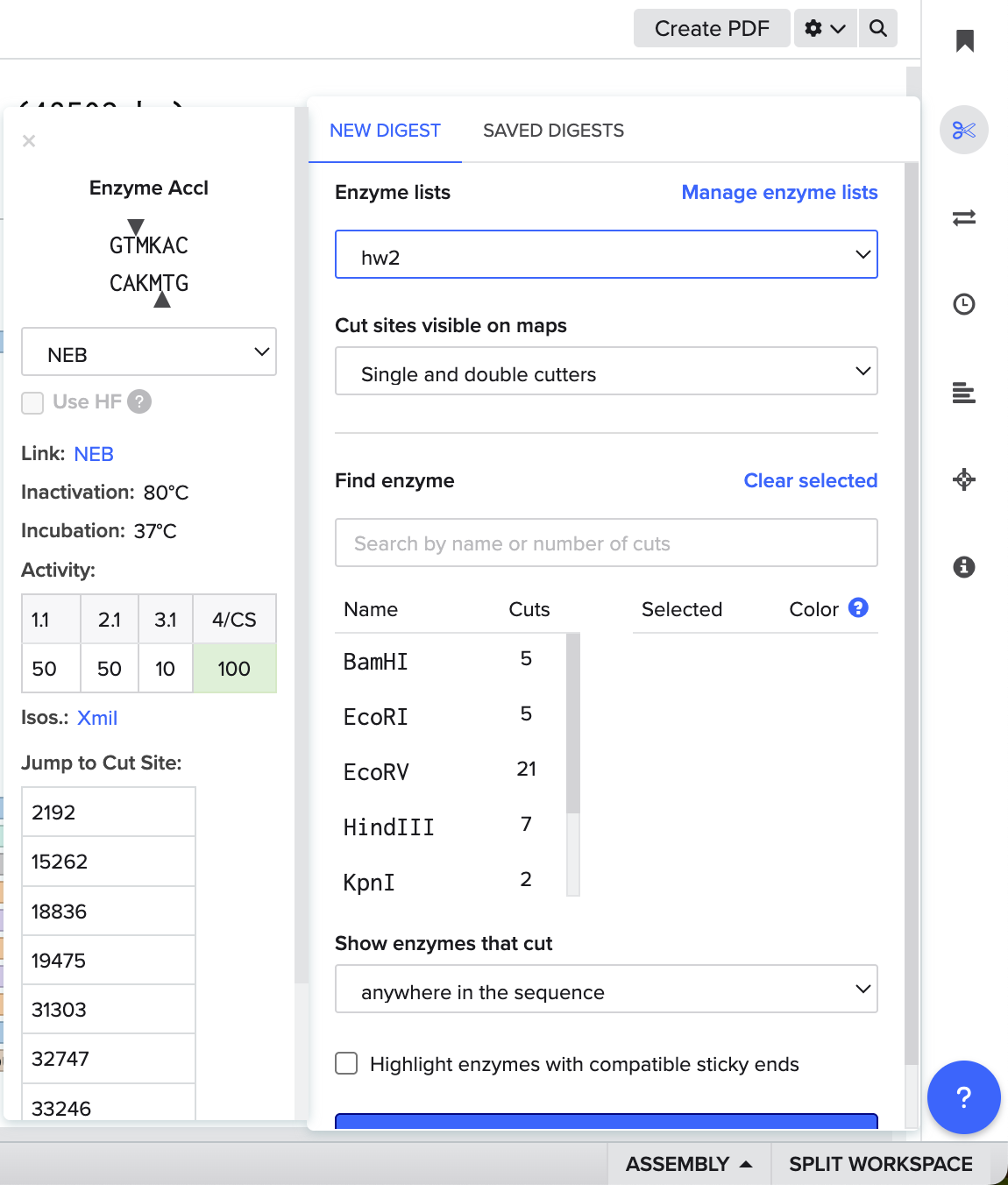 Patterns I got from each enzyme and all enzymes together.
Patterns I got from each enzyme and all enzymes together.
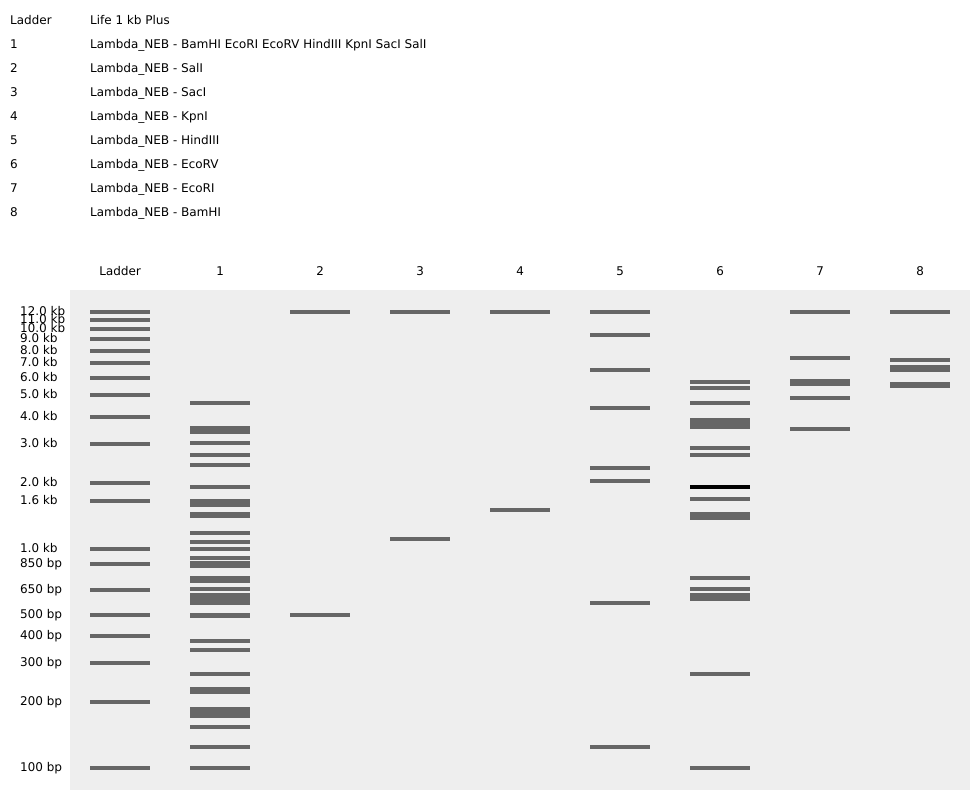 I tried to get a Lychee out of them. Can you see it?
I tried to get a Lychee out of them. Can you see it?
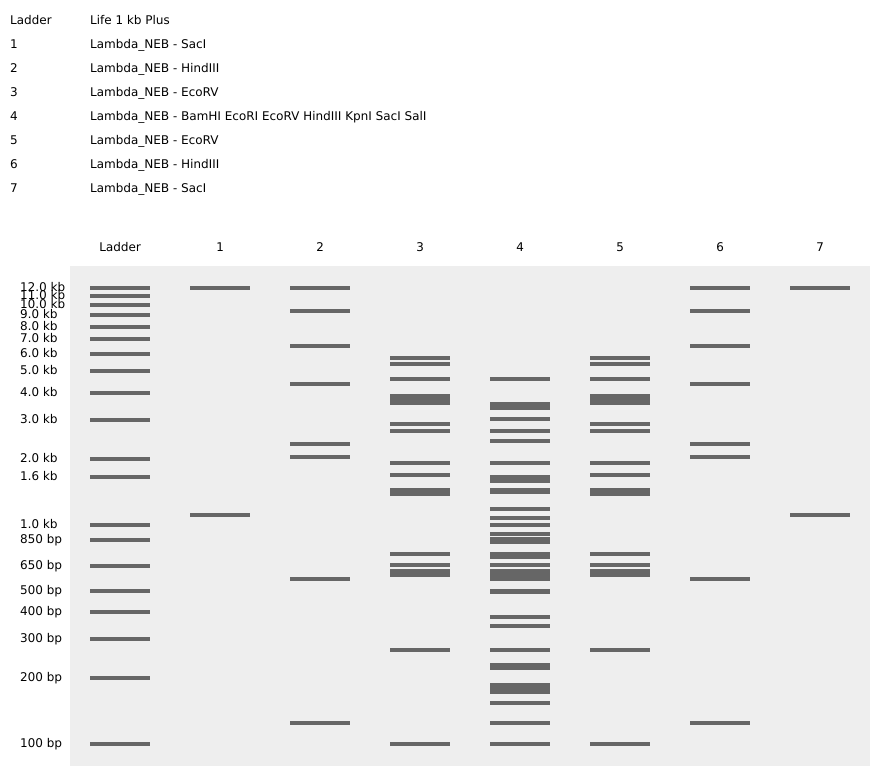

Part 3: DNA Design Challenge
3.1. Choose your protein.
For this assignment, I chose UDP-glucose pyrophosphorylase (UGPase) from Komagataeibacter xylinus.
I selected this protein because it plays a central role in bacterial cellulose biosynthesis.
UDP-glucose pyrophosphorylase catalyzes the conversion of glucose-1-phosphate and UTP into UDP-glucose, which is the immediate activated precursor used by cellulose synthase to polymerize β-1,4-glucan chains.
Since my final project focuses on improving bacterial cellulose production for biofilm-based food packaging materials, understanding and potentially enhancing the precursor supply through UGPase is directly relevant to my research goals.
To obtain the protein sequence, I used the UniProt database. I searched for “UDP-glucose pyrophosphorylase Komagataeibacter xylinus” and identified the corresponding UniProt entry (Accession: P27897). The database provides the complete annotated protein sequence in FASTA format.
Protein Information
Protein name: UDP-glucose pyrophosphorylase Organism: Komagataeibacter xylinus UniProt accession: P27897 Function: Catalyzes the formation of UDP-glucose, the direct precursor for cellulose biosynthesis.
Protein Sequence (FASTA format)
sp|P27897|UGP_KOMXY UDP-glucose pyrophosphorylase OS=Komagataeibacter xylinus OX=28448 GN=galU PE=1 SV=1
MSTVKILANVAGVKADGVVVPTGDLAKAGWVIVGGDGSLS
ETVRVGKLLEEAQLRASRDPAEVIVALTPEGHILGDAQTV
VIGAGGTGKSGYEGLARILPDDSLSVPLGITVEKARDAFL
RNPIIDALGKVMGKDAVNLVDQGELVDLDALGVSR AALID
AGGGTRGHTLAVAAAGANARGLDRLKAGADKAKLGGVEIL
DKSVGAAHGLQALRGLGIDSDGAAVILSRKLGSYEKLGAG
TIVAPLALLAEAVGAKGMVYGEARLITNGEGQTIVVAGAG
NLVGADTIVVTEGYDRGILSGYEGAHGLRVGIEGVVQPIG
VNAGEATDLGVLGVDL
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
The Central Dogma explains that DNA is transcribed into RNA and translated into protein. Because the genetic code is degenerate (most amino acids are encoded by multiple codons), a given protein sequence can correspond to multiple possible DNA sequences. Therefore, reverse translation tools generate a plausible coding DNA sequence based on a selected codon usage table.
To infer the nucleotide sequence corresponding to my chosen protein (UDP-glucose pyrophosphorylase, UGPase), I used an online reverse translation tool. I kept the default codon usage table, which the tool states was generated from all E. coli coding sequences in GenBank (obtained from the Codon Usage Database). Since E. coli is a representative bacterial organism and my protein is from a bacterium, this provides a reasonable codon-usage reference for generating a plausible coding sequence.
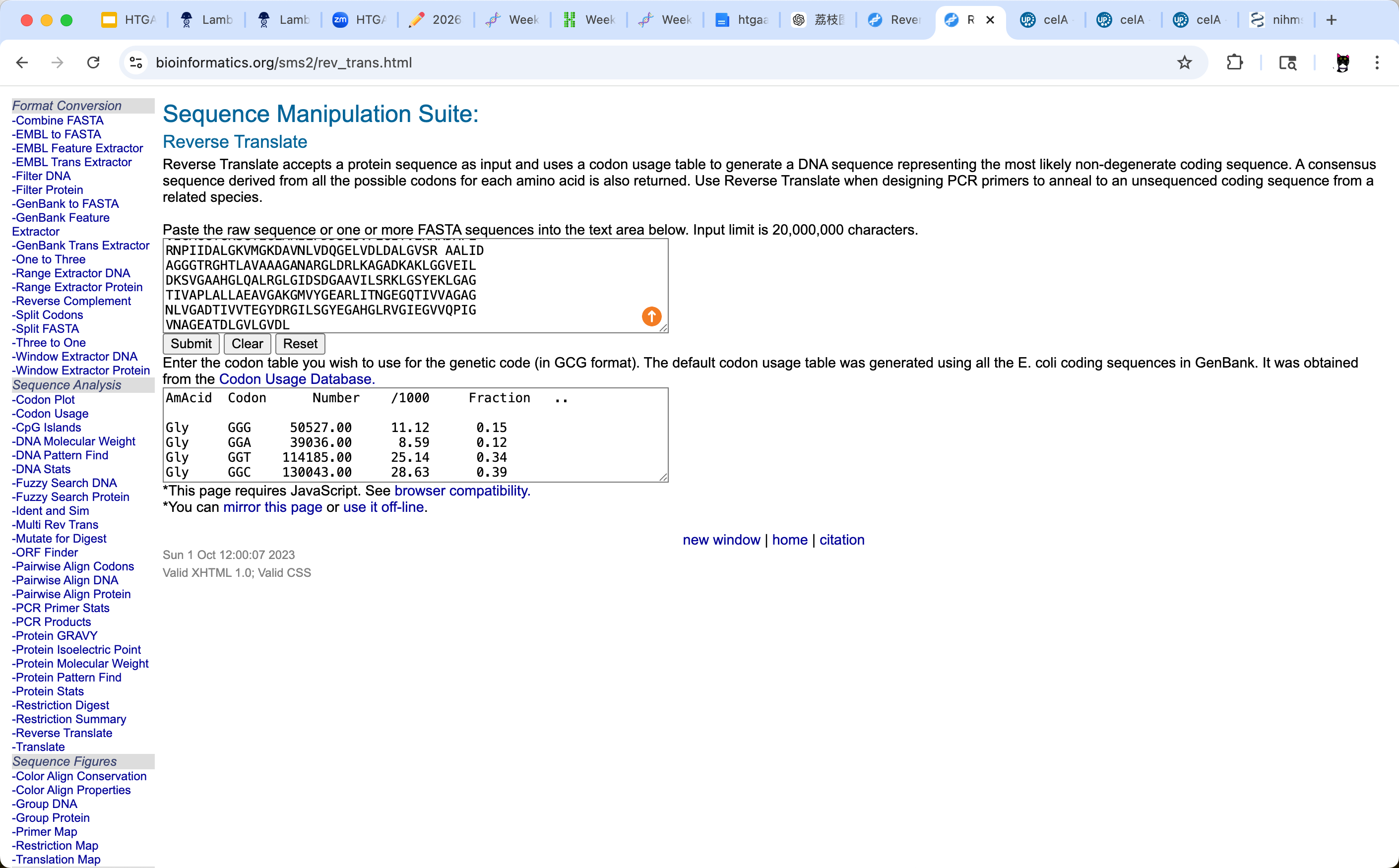
Reverse-translated coding DNA sequence (5’→3’):
reverse translation of Untitled to a 1008 base sequence of most likely codons. atgagcaccgtgaaaattctggcgaacgtggcgggcgtgaaagcggatggcgtggtggtg ccgaccggcgatctggcgaaagcgggctgggtgattgtgggcggcgatggcagcctgagc gaaaccgtgcgcgtgggcaaactgctggaagaagcgcagctgcgcgcgagccgcgatccg gcggaagtgattgtggcgctgaccccggaaggccatattctgggcgatgcgcagaccgtg gtgattggcgcgggcggcaccggcaaaagcggctatgaaggcctggcgcgcattctgccg gatgatagcctgagcgtgccgctgggcattaccgtggaaaaagcgcgcgatgcgtttctg cgcaacccgattattgatgcgctgggcaaagtgatgggcaaagatgcggtgaacctggtg gatcagggcgaactggtggatctggatgcgctgggcgtgagccgcgcggcgctgattgat gcgggcggcggcacccgcggccataccctggcggtggcggcggcgggcgcgaacgcgcgc ggcctggatcgcctgaaagcgggcgcggataaagcgaaactgggcggcgtggaaattctg gataaaagcgtgggcgcggcgcatggcctgcaggcgctgcgcggcctgggcattgatagc gatggcgcggcggtgattctgagccgcaaactgggcagctatgaaaaactgggcgcgggc accattgtggcgccgctggcgctgctggcggaagcggtgggcgcgaaaggcatggtgtat ggcgaagcgcgcctgattaccaacggcgaaggccagaccattgtggtggcgggcgcgggc aacctggtgggcgcggataccattgtggtgaccgaaggctatgatcgcggcattctgagc ggctatgaaggcgcgcatggcctgcgcgtgggcattgaaggcgtggtgcagccgattggc gtgaacgcgggcgaagcgaccgatctgggcgtgctgggcgtggatctg
3.3. Codon optimization.
After determining the nucleotide sequence corresponding to my chosen protein (UDP-glucose pyrophosphorylase, UGPase), codon optimization was performed to improve potential expression efficiency.
Although multiple codons can encode the same amino acid, different organisms exhibit codon usage bias, meaning they preferentially use certain synonymous codons over others. If a gene contains codons that are rare in the target host, translation efficiency may decrease due to limited availability of corresponding tRNAs. This can lead to reduced protein expression or translational stalling. Therefore, codon optimization modifies the nucleotide sequence while preserving the amino acid sequence, replacing rare codons with host-preferred synonymous codons.
I used Benchling’s codon optimization tool to optimize the coding sequence. Since Komagataeibacter was not directly available in the organism list, I selected Escherichia coli as a representative bacterial host for codon optimization. E. coli is a well-characterized prokaryotic expression system and provides a reasonable approximation of bacterial codon usage bias.
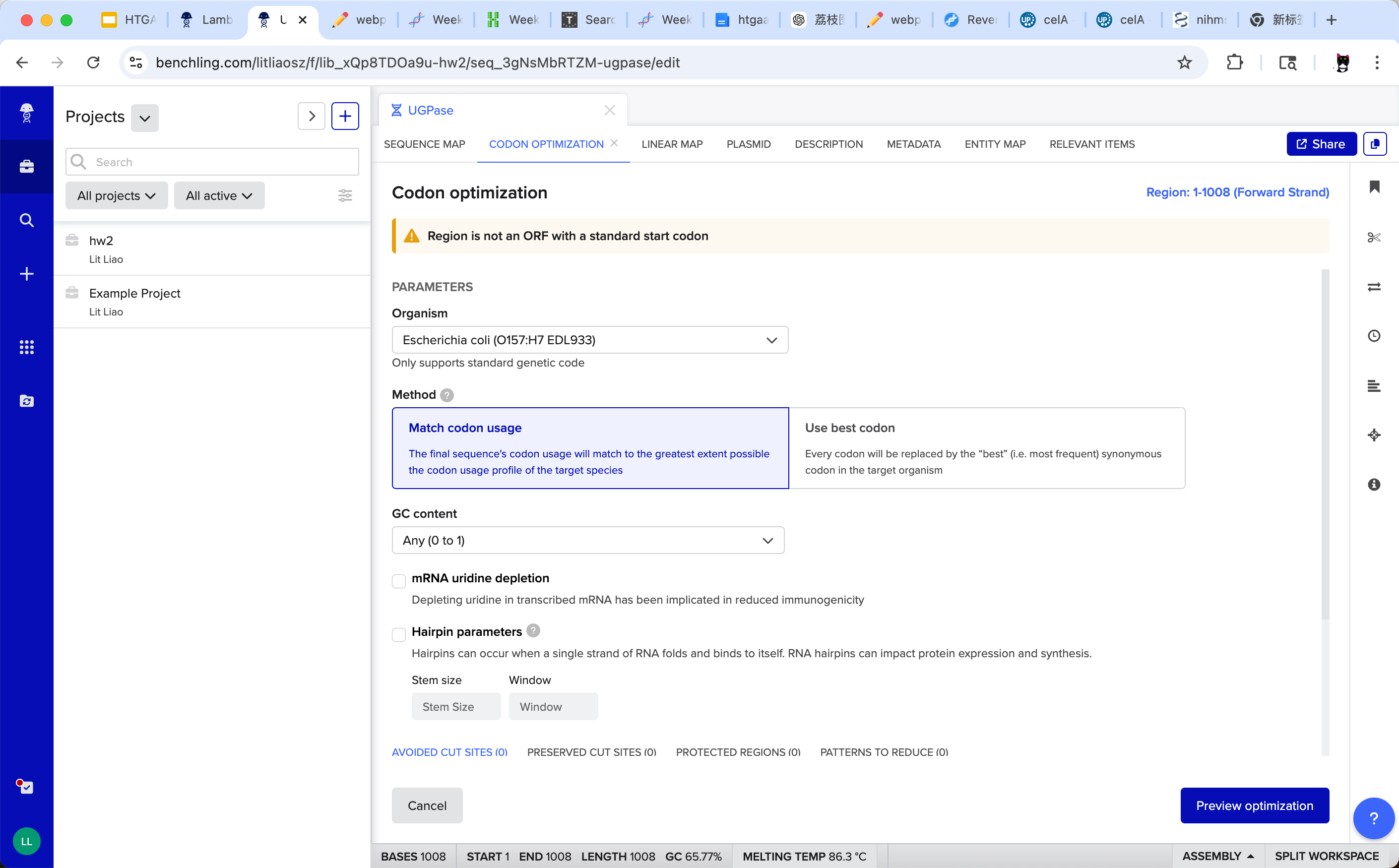
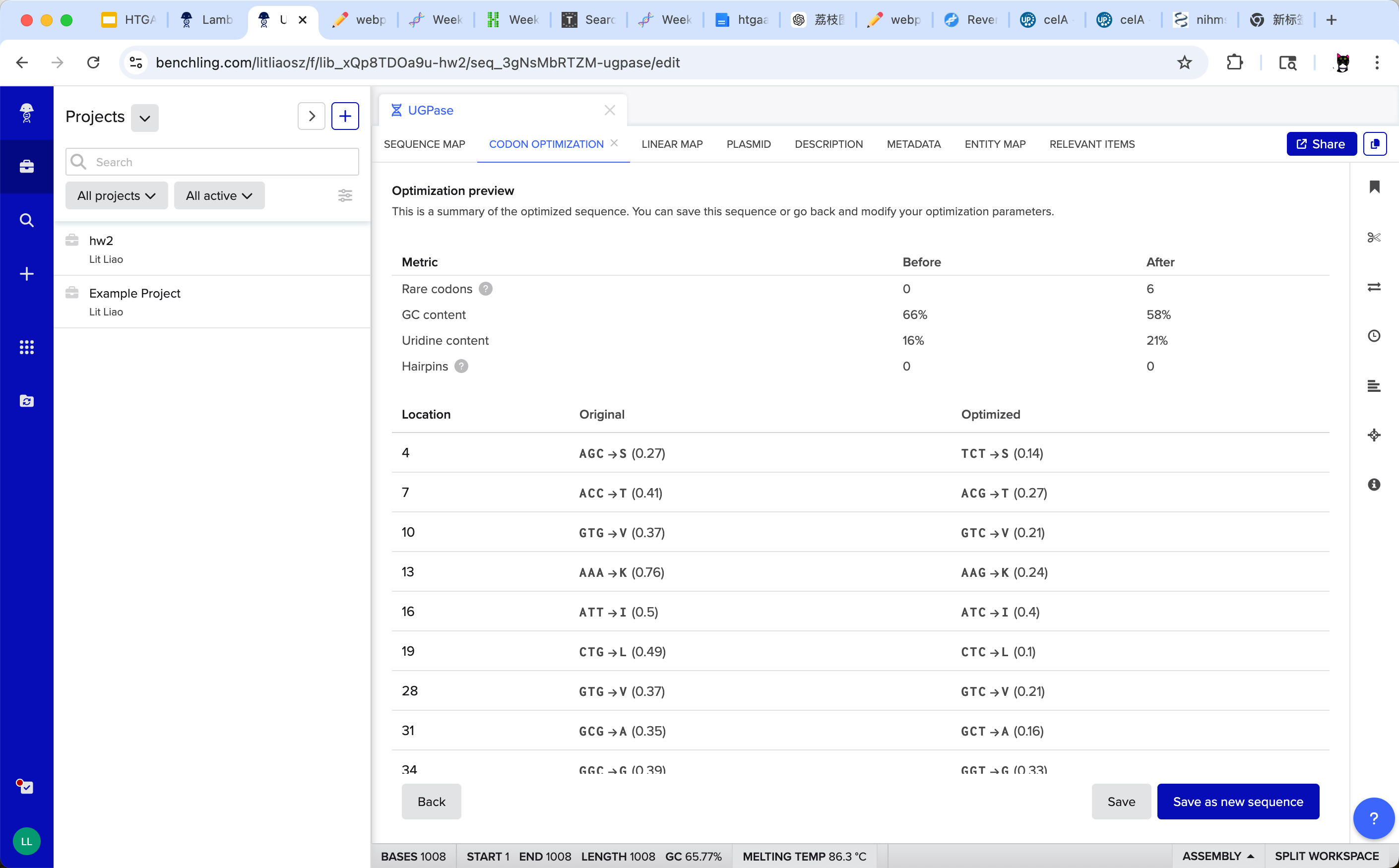
The optimization process replaced multiple synonymous codons without altering the protein sequence. Notably, the GC content decreased from 66% to 58%, bringing the sequence closer to typical bacterial expression ranges. No additional RNA hairpin structures were introduced during optimization. These changes may improve translation efficiency and overall protein expression in the selected host.
Optimized sequence is here:
atgtctacggtcaagatcctcgcgaacgtcgctggtgtcaaagcagacggcgttgtggtcccaacgggcgatctagcgaaggctggttgggtaattgttggtggcgatggctcgctctcggagactgtccgggtgggcaagttactggaggaagcgcagctgcgcgcctcacgcgaccctgctgaagttatcgttgccctgacaccggaaggccatatcctgggagatgcccagaccgttgttattggcgcggggggtacgggtaaatccggatatgaaggcctggcgcgcattcttcccgatgatagcctgtctgtgccacttggcatcactgtggaaaaagcccgagacgcttttctgcgcaacccgatcatcgatgctctgggtaaagtgatggggaaagatgcggttaatttggttgatcagggcgaactcgttgatctggatgctctgggcgtgagtcgtgcagcactgatcgacgcgggtggaggtacccgtggccacaccctggccgtggccgccgcgggagcgaacgcgagaggtttagaccgcctgaaagcgggtgcagacaaagcgaaactcggcggcgtagagattcttgataaaagcgtgggggcagctcatggattgcaggcgcttcgtggcctgggcattgacagtgacggtgcagccgtgattctgtcccgcaagctaggctcatacgagaaactgggggcaggaaccattgtggcgccgctggccttattagcggaagccgtaggtgcaaaagggatggtctacggggaagcccgtttgatcaccaatggcgagggccaaacgattgtagtagctggtgcaggcaacctggtcggggcggacactatagtggtgaccgaaggttatgatcgggggattttgagcggttatgaaggtgcccacgggctgcgtgtgggtattgaaggcgttgtgcaaccgataggagtgaatgccggtgaagcaacagatttaggcgtactgggtgtcgatttg
3.4. You have a sequence! Now what?
To produce the UDP-glucose pyrophosphorylase (UGPase) protein from the optimized DNA sequence, a cell-dependent expression system can be used.
In this approach, the optimized coding sequence is inserted into a plasmid vector. The plasmid contains regulatory elements necessary for gene expression, such as a promoter to initiate transcription, a ribosome binding site (RBS) to initiate translation, and a transcription terminator. The plasmid also includes an origin of replication and a selectable marker to maintain the plasmid within the host cells.
The recombinant plasmid is then introduced into a bacterial host, such as Komagataeibacter (the intended production organism). Once inside the cell, the DNA sequence follows the central dogma of molecular biology:
DNA → RNA → Protein
First, the host cell’s RNA polymerase binds to the promoter and transcribes the DNA sequence into messenger RNA (mRNA). The mRNA is then recognized by ribosomes, which translate the nucleotide sequence into the corresponding amino acid sequence according to the genetic code. Transfer RNAs (tRNAs) deliver the appropriate amino acids, and the ribosome assembles them into the UGPase protein.
Through this process, the host cell uses its natural transcription and translation machinery to produce the target enzyme. In the context of this project, increased expression of UGPase may enhance the production of UDP-glucose, potentially contributing to improved cellulose biosynthesis.
Part 5: DNA Read/Write/Edit
5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why? I am interested in DNA as a medium for long-term archival data storage. Specifically, I would want to sequence synthetic DNA molecules that encode digital information converted into nucleotide sequences.DNA has extremely high storage density and long-term stability compared to traditional silicon-based data storage systems. Many forms of data—such as historical records, scientific archives, or cultural information—do not require frequent editing or real-time access. These “cold data” could potentially be stored in DNA to significantly reduce energy consumption, freeing silicon-based servers for high-demand or “hot” data.
Conceptually, I imagine a future system where biological organisms, such as trees, could serve as long-term data carriers. Through photosynthesis and natural growth, such systems could theoretically maintain archived data with minimal external energy input. While this remains speculative, it reflects the idea of integrating biological processes into sustainable information storage infrastructures.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
To retrieve information encoded in DNA, sequencing technologies would be required to accurately read the nucleotide sequences.
For DNA-based data storage, high-accuracy sequencing platforms such as Illumina sequencing would be suitable due to their low error rates and scalability. Illumina systems are widely used for short-read, high-fidelity sequencing, which aligns well with synthetic DNA data storage formats.
Alternatively, long-read sequencing technologies such as Oxford Nanopore could be considered, especially if larger DNA constructs are used. Nanopore sequencing offers portability and real-time readout, which may be advantageous for decentralized biological storage systems.
While genome editing technologies like CRISPR can insert DNA sequences into living organisms, they are not sequencing technologies. If biological hosts were used as storage media, genome sequencing would still rely on established sequencing platforms rather than gene-editing tools.
In the long term, bridging silicon-based storage systems and carbon-based biological storage systems may require new bioelectronic interfaces. Such interfaces could enable digital-to-biological encoding and biological-to-digital decoding, potentially integrating molecular biology with computational systems.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why?For this project, I would like to synthesize DNA constructs aimed at enhancing the production of sustainable biomaterials that could potentially replace petroleum-based plastics.
Specifically, I am interested in engineering bacterial cellulose-producing systems, such as Komagataeibacter, to increase the yield and functional properties of cellulose-based biofilms. Bacterial cellulose is a promising biodegradable material with high mechanical strength, purity, and biocompatibility. It has potential applications in food packaging, medical materials, and sustainable textiles.
Conceptual DNA Constructs
Rather than synthesizing an entirely new genome, I would focus on designing modular genetic constructs that include:
Genes involved in UDP-glucose biosynthesis
Since UDP-glucose is the direct precursor for cellulose synthesis, enhancing enzymes in this pathway (e.g., UDP-glucose pyrophosphorylase, UGPase) may increase substrate availability and improve cellulose yield.
Regulatory elements for enhanced expression
These would include promoters and regulatory sequences that enable controlled, elevated expression of cellulose-related enzymes without imposing excessive metabolic burden.
Material-functionalization modules (optional future direction)
Additional genetic modules could encode proteins that modify cellulose properties—for example:
The overall goal would not be to fundamentally redesign the organism, but rather to create a tunable genetic system that enhances production of biodegradable cellulose-based biomaterials.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
To synthesize the DNA constructs, I would use a commercial DNA synthesis platform, such as those provided by biotechnology companies. These platforms chemically synthesize DNA sequences based on a digital design. This allows precise writing of custom genetic sequences for biomaterial production.
Commercial synthesis is suitable because it is reliable, scalable, and widely used in synthetic biology research.
To confirm the DNA sequence, I would use a next-generation sequencing (NGS) technology.
In general, sequencing involves:
This ensures that the synthesized DNA matches the intended design.
5.3 DNA Edit
(i) What DNA would you want to edit and why?I am interested in exploring DNA editing as a way to create a biological–digital interface rather than traditional therapeutic or enhancement applications.
Specifically, I am curious about editing genetic pathways that influence how biological systems sense and respond to external signals. Instead of modifying core human traits, I would focus on engineering cells capable of detecting and translating molecular or electrical signals into readable outputs. Such systems could potentially serve as an interface between biological organisms and external data storage systems, whether silicon-based digital storage or DNA-based archival storage.
The goal would not be human augmentation in terms of physical or cognitive enhancement, but rather the creation of a communication bridge between biological systems and computational systems. This concept resembles brain–computer interfaces, but extended toward a broader biological–informational integration.
(ii) What technology or technologies would you use to perform these DNA edits and why?
For conceptual DNA editing, I would consider programmable genome editing tools such as CRISPR-based systems, which allow targeted modification of specific DNA regions.
These systems function by using a guide RNA to direct a nuclease to a specific DNA sequence. The nuclease creates a break at the targeted location, and the cell’s natural repair mechanisms then introduce changes or insert new genetic material.
In principle, this allows precise editing of genes involved in sensing, signal processing, or molecular response pathways.
Essential steps (conceptual overview)
Preparation and Inputs (conceptual)
Limitations