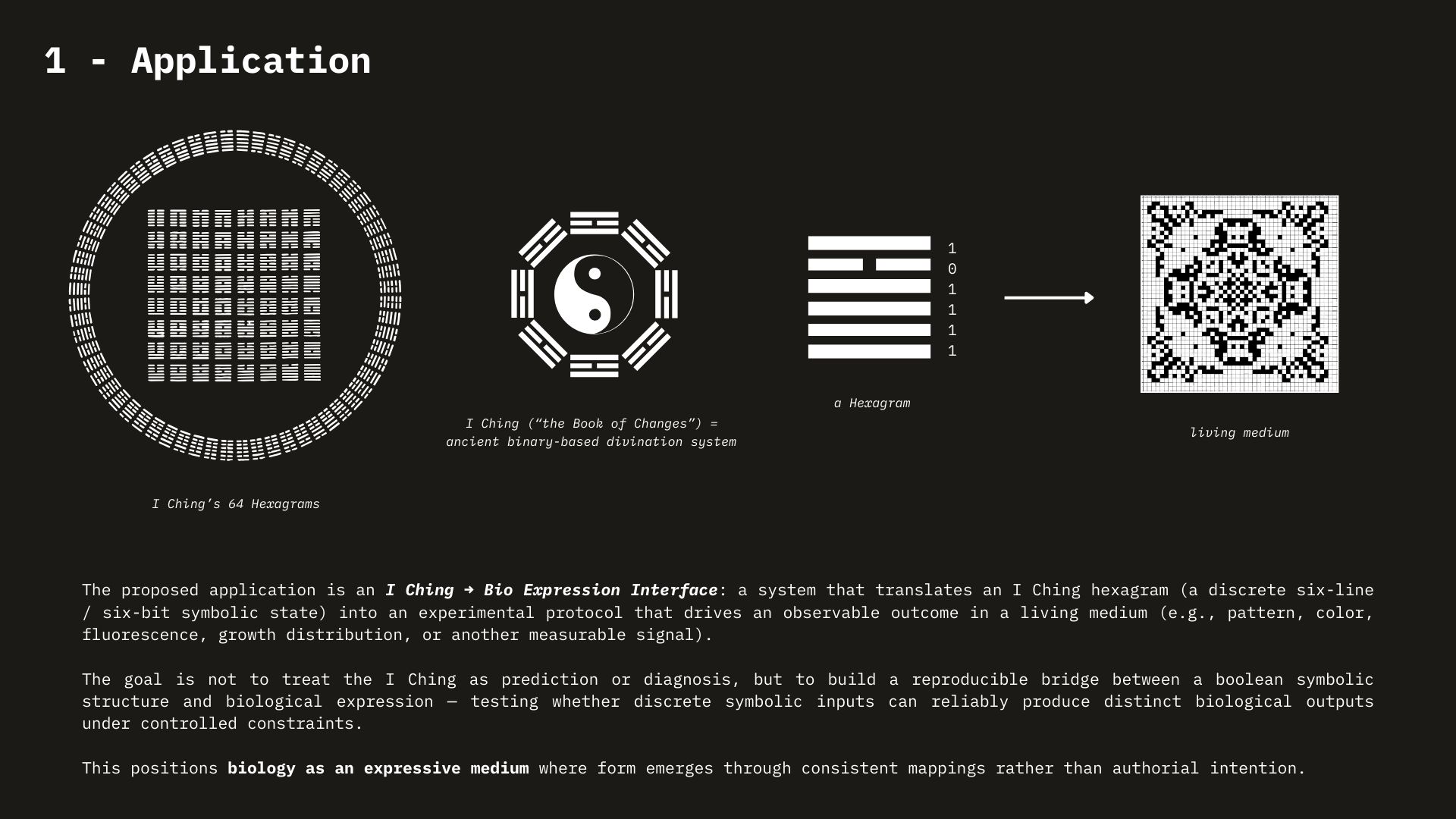
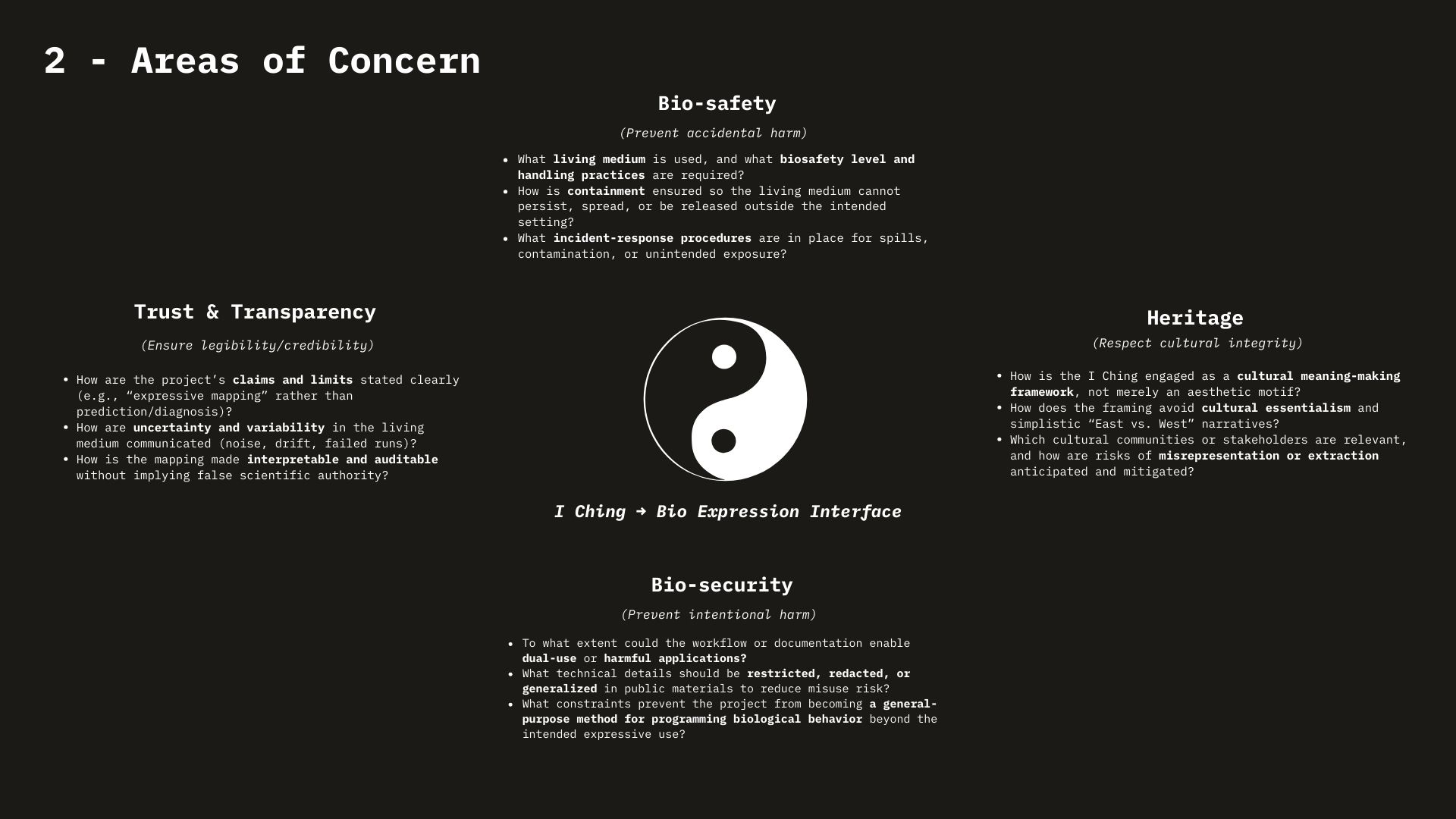
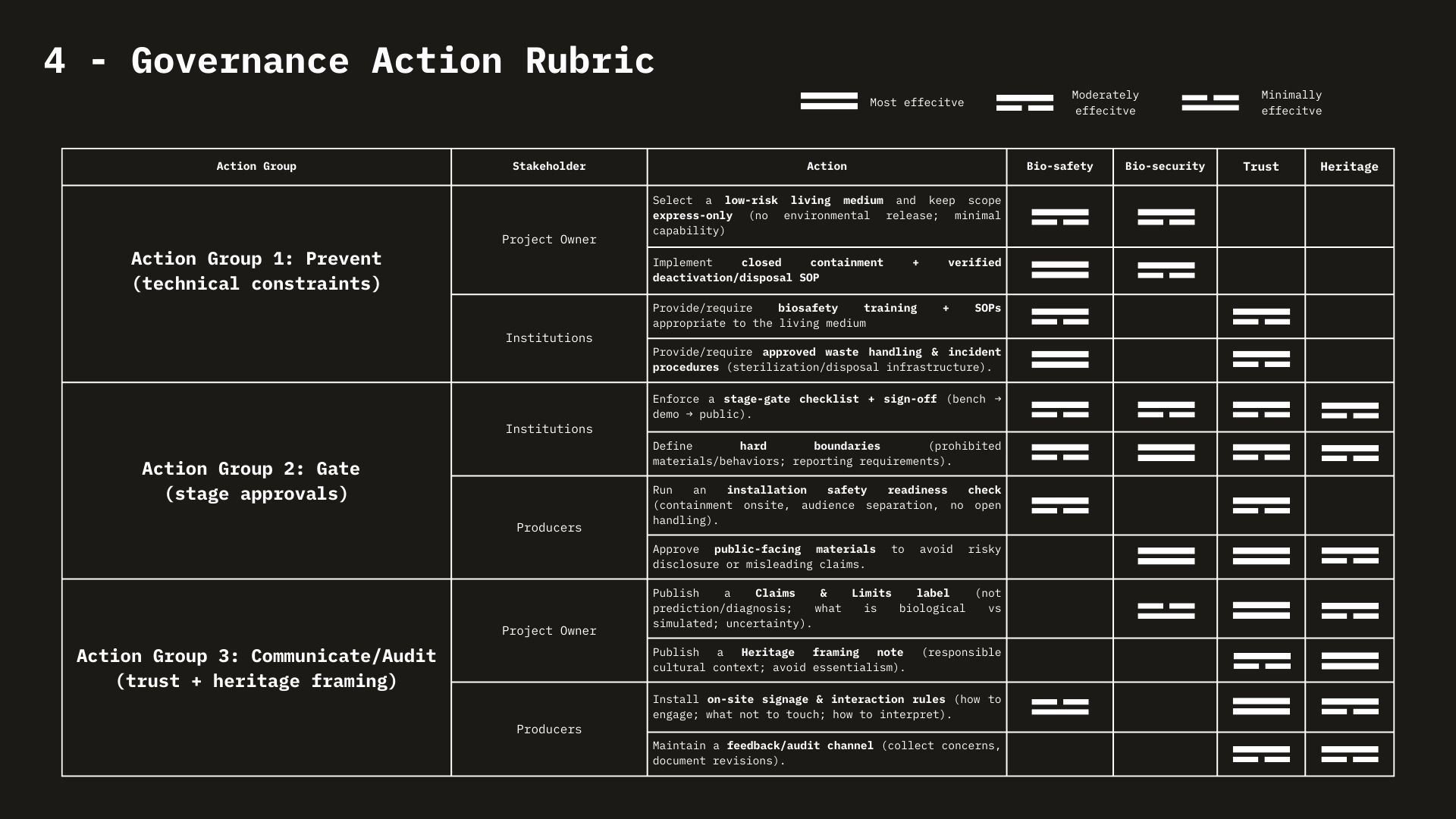
Week 1 HW: Principles and Practices
WEEKLY ASSIGNMENT
Ethical concerns that came up especially new to me
- “Halfpipe of Doom” / dual-use is not an edge case — it’s the default. Many tools in synthetic biology are structurally dual-use: the same capability that enables public good (vaccines, diagnostics, remediation) can also enable harm (accidents, misuse, weaponization). This shifts ethics from “don’t do bad things” to “assume capabilities will be repurposed.”
- Harm can be created while trying to prevent harm. The pandemic example (engineering SARS-CoV-2) made this concrete: building countermeasure capability can also expand the risk surface (accidental release, intentional misuse, normalization of high-risk methods). The ethical concern here is not only intent, but second-order effects and indirect harms.
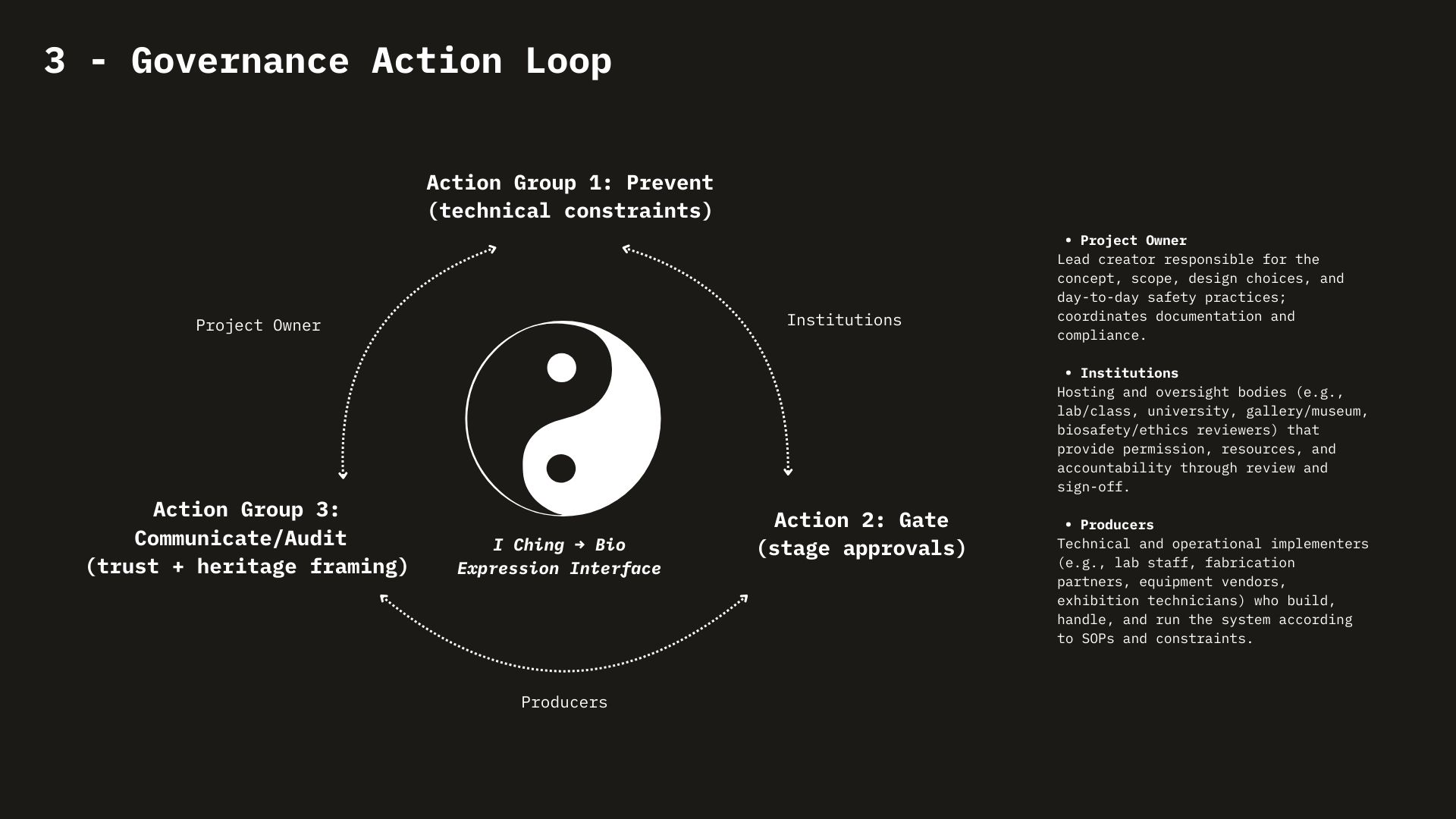
- Responsibility is diffuse across an ecosystem, not located in one lab. The governance framework (goals × actors × actions) highlighted a new point: risk isn’t just about a “bad actor.” It’s distributed across supply chains and institutions (gene firms, oligo manufacturers, synthesizer makers, end users). Ethical questions become: who has leverage to prevent incidents? who bears cost? who is accountable when the system fails?
- Trust is a technical constraint, not a PR layer. The Asilomar discussion reframed “public acceptance” as part of the engineering problem. Even a scientifically sound technology can fail socially if institutions are not trusted, if governance is opaque, or if the public feels excluded from deciding what futures are preferred.
- Biotech is becoming economic and geopolitical infrastructure. Bio-apps → bio-economies → bio-power made it clear that biotech is not just science; it’s tied to national strategies, competition, and incentives. That raises ethical risks around: profit-driven priorities, uneven access, and “who benefits” vs “who is exposed.”
Governance actions that seem appropriate:
Rather than one solution, I would propose overlapping governance layers across the ecosystem:
- Upstream capability safeguards (prevent incidents)
- Screening + verification norms for sensitive DNA orders (where relevant), with clear escalation pathways when flags appear.
- Access friction for high-risk capabilities: tiered access, training requirements, or licensing for certain tools/processes—aimed at reducing accidental/low-skill misuse.
- Lab safety + containment as baseline (prevent + respond)
- Reinforce biosafety culture (not only compliance): training, incident reporting without punishment, and continuous review as techniques evolv
- Build explicit response readiness: what happens if something goes wrong—communication, coordination, and corrective action.
- Ecosystem accountability (shared responsibility)
- Make responsibility legible: map actors (firms, manufacturers, users) and define who is expected to do what under specific risk scenarios.
- Encourage governance actions that reduce “accountability gaps,” where risk is created upstream but harms land downstream.
- Trust-building as governance
- Treat transparency and participation as governance tools:
- clearer explanation of benefits/risks and why certain work is done
- channels for stakeholder input (not only experts)
- The goal isn’t consensus on everything, but earned legitimacy.
- Bioliteracy as a long-term governance strategy
- Expand bio-literacy as a capacity-building governance move (analogous to how computer literacy scaled).
- This isn’t about making everyone a scientist; it’s about enabling the public to meaningfully engage with choices about biotech futures.
WEEK 2 LECTURE PREP Homework Questions from Professor Jacobson:
- Polymerase’s error rate is 1: 10⁶ bases (10 ⁻⁶ per base). The human genome is ~3.1 × 10⁹ to 3.3 x 10⁹ bp, so at that rate we’d expect ~3.1-3.3 × 10³ (~3,000) errors per genome copy. Biology handles this with layered error correction: polymerase proofreading plus post-replication repair (especially mismatch repair) and other DNA repair/checkpoints, which together drive the effective error rate much lower.
- An average human protein is about 1036 bp (~345 codons), and because the genetic code is redundant (many amino acids have multiple codons), there are a huge number of different nucleotide sequences that can still encode the same protein sequence. However, many of these recoded sequences don’t work well in practice because changing codons changes the DNA/RNA’s GC% and base-pairing strength (A/T ~ −1.2 vs G/C ~ −2.0 kcal/mol), which can drastically change minimum free energy secondary structure; some sequences fold into structures that make expression or stability worse, so a lot of possible codes aren’t actually good choices for expressing the protein you want.
Homework Questions from Dr. LeProust:
- The most commonly used method is solid-phase oligonucleotide synthesis using phosphoramidite chemistry
- Direct synthesis struggles past ~200 nt because each base-addition step can fail a little, and over hundreds of steps those small failures pile up into lots of errors and many incomplete (“truncation”) products, so the amount of correct full-length oligo drops a lot. Additionally, the effective error situation gets much worse at that baseline scale (it explicitly contrasts 1:3,000 nt vs. 1:200 nt) and shows truncation products when pushing longer syntheses.
- You can’t make a 2000 bp gene by direct oligo synthesis because chemical synthesis is only dependable for much shorter lengths (on the order of a few hundred nucleotides), and beyond that the reaction produces lots of errors and truncated (incomplete) products, so the chance of getting a correct, full-length 2000 bp molecule becomes vanishingly small. That’s why longer DNA is made by assembling it from shorter pieces (gene fragments/oligos) rather than synthesizing the whole 2000 bp in one go.
Homework Questions from George Church:
- The way I connect these two is pretty simple: lysine being “essential” means animals already can’t make it and have to get it from outside—food, microbes, the whole ecosystem. So the “lysine contingency” idea (engineer something to need lysine so it can’t survive outside) sounds clever, but it’s kind of backwards in an animal world, because lysine isn’t some rare lab-only thing. It’s literally a normal dietary requirement that the environment is already structured to provide. That makes lysine-dependence a weak containment strategy, and it pushes me to take biocontainment more seriously—like making the organism dependent on something that doesn’t exist in nature, not a basic nutrient every animal already relies on.