Week 6 HW: Genetic Circuits Part I


Part 1: Homework Questions
- Phusion High-Fidelity PCR Master Mix contains several components required for DNA amplification. The Phusion DNA polymerase is the enzyme that synthesizes new DNA strands; it has proofreading (3’→5’ exonuclease) activity, which greatly reduces mutation rates compared with standard Taq polymerase. The mix also includes dNTPs (deoxynucleotide triphosphates), which are the nucleotide building blocks incorporated into the newly synthesized DNA. A reaction buffer provides the correct chemical environment (pH, salts, stabilizers) to maximize enzyme activity and fidelity. Mg²⁺ ions (usually from MgCl₂) act as essential cofactors for polymerase function and influence enzyme efficiency.
- Primer annealing temperature is mainly determined by the melting temperature (Tm) of the primers, which depends on their sequence composition. Primers with higher GC content generally have higher Tm because G–C pairs form three hydrogen bonds compared with two in A–T pairs. Primer length also affects Tm, as longer primers form more stable duplexes with the template. Salt concentration and Mg²⁺ levels in the reaction buffer influence DNA duplex stability and therefore the optimal annealing temperature. Additionally, primer secondary structure or mismatches can reduce binding stability and may require lower annealing temperatures.
- PCR and restriction enzyme digestion both produce linear DNA fragments but operate through different mechanisms. PCR uses primers and a DNA polymerase to amplify a specific DNA region from a template through thermal cycling. This method is highly flexible because primers can introduce mutations, overhangs, or homologous regions, making PCR useful when generating fragments for cloning or modifying sequences. In contrast, restriction enzyme digestion uses enzymes that recognize specific short DNA sequences and cut at those sites, producing predictable fragments with defined ends (often sticky or blunt). The digest protocol is simpler and faster if the required restriction sites already exist in the DNA. PCR is preferable when amplifying small regions, adding sequences, or working from low DNA amounts, while restriction digests are preferable when cutting large plasmids or isolating fragments with existing restriction sites without introducing polymerase errors.
- For Gibson Assembly, DNA fragments must have overlapping homologous sequences at their ends so they can anneal during the assembly reaction. These overlaps are usually designed into PCR primers, ensuring that adjacent fragments share complementary sequences. After PCR amplification or digestion, the fragments should be checked by gel electrophoresis to confirm the correct size and purity. It is also important to verify that the overlaps match the intended assembly order and that no incompatible restriction sites remain within the overlaps.
- Golden Gate Assembly is a DNA cloning method that uses Type IIS restriction enzymes and DNA ligase in a single reaction to assemble multiple DNA fragments in a defined order. Unlike standard restriction enzymes, Type IIS enzymes cut outside their recognition site, producing custom overhangs that can be designed to be unique for each fragment. During the reaction, the restriction enzyme cuts the DNA to create compatible overhangs, and DNA ligase simultaneously joins the fragments together. Because the recognition sites are removed after ligation, the final assembled product cannot be cut again, allowing the reaction to proceed efficiently toward the correct construct.
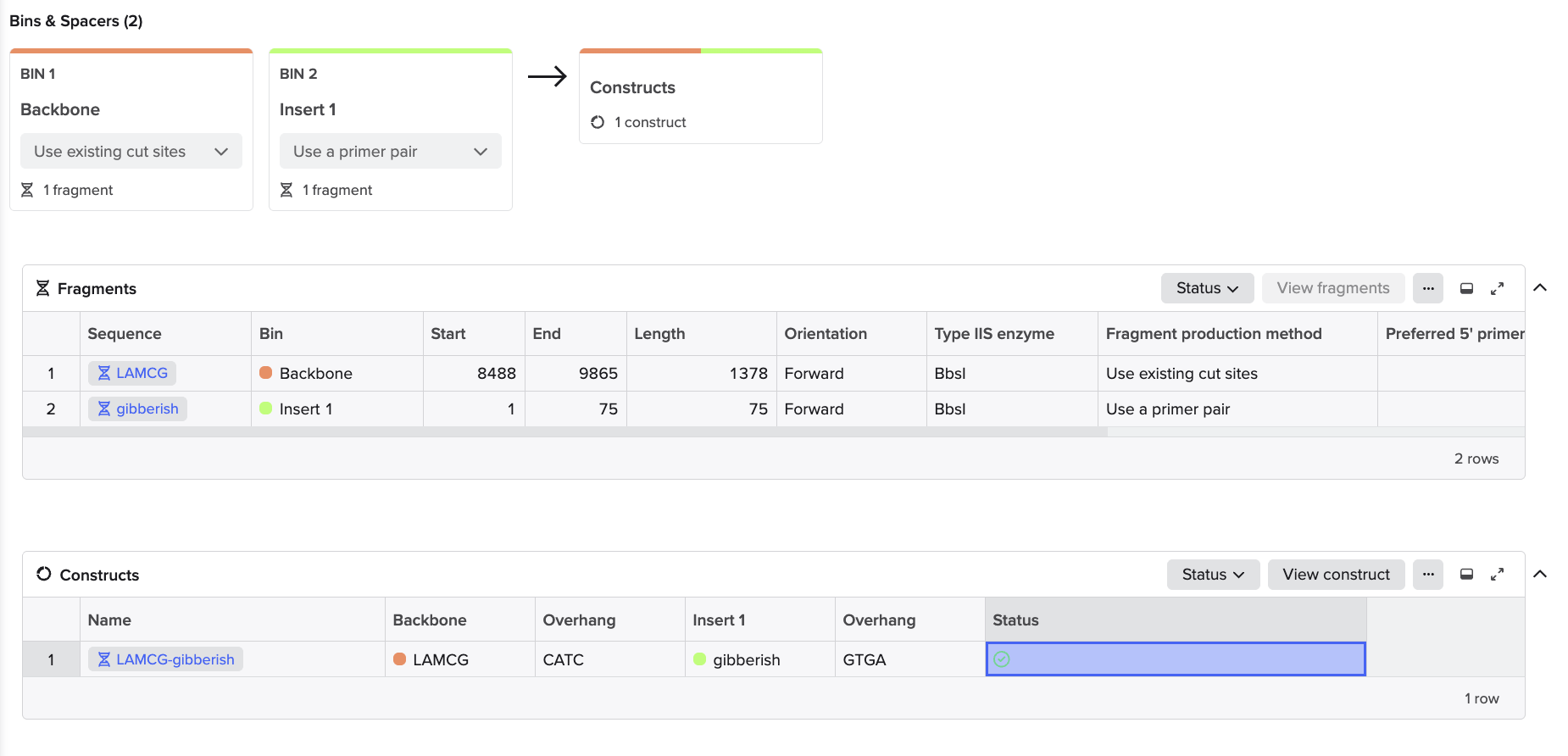
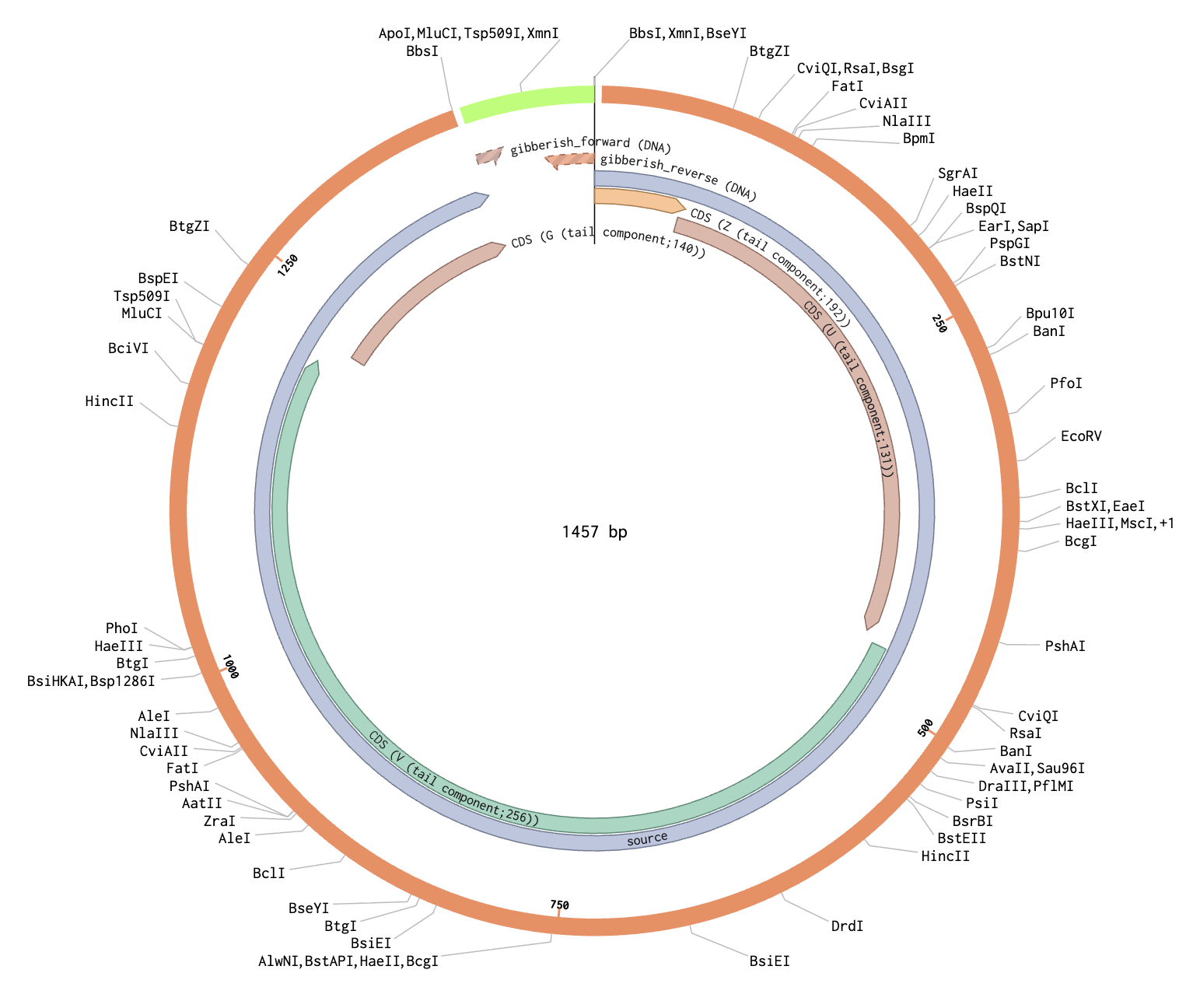
To model this, I just chose two random sequences (LACMG that we worked with once, and a gibberish one that I have for some reason). I went to Benchling to model the assembly, and this is what came out of it:
Hopefully the steps I followed were right because I just followed the template that I Was provided through Benchling.
Part 2: Kernel
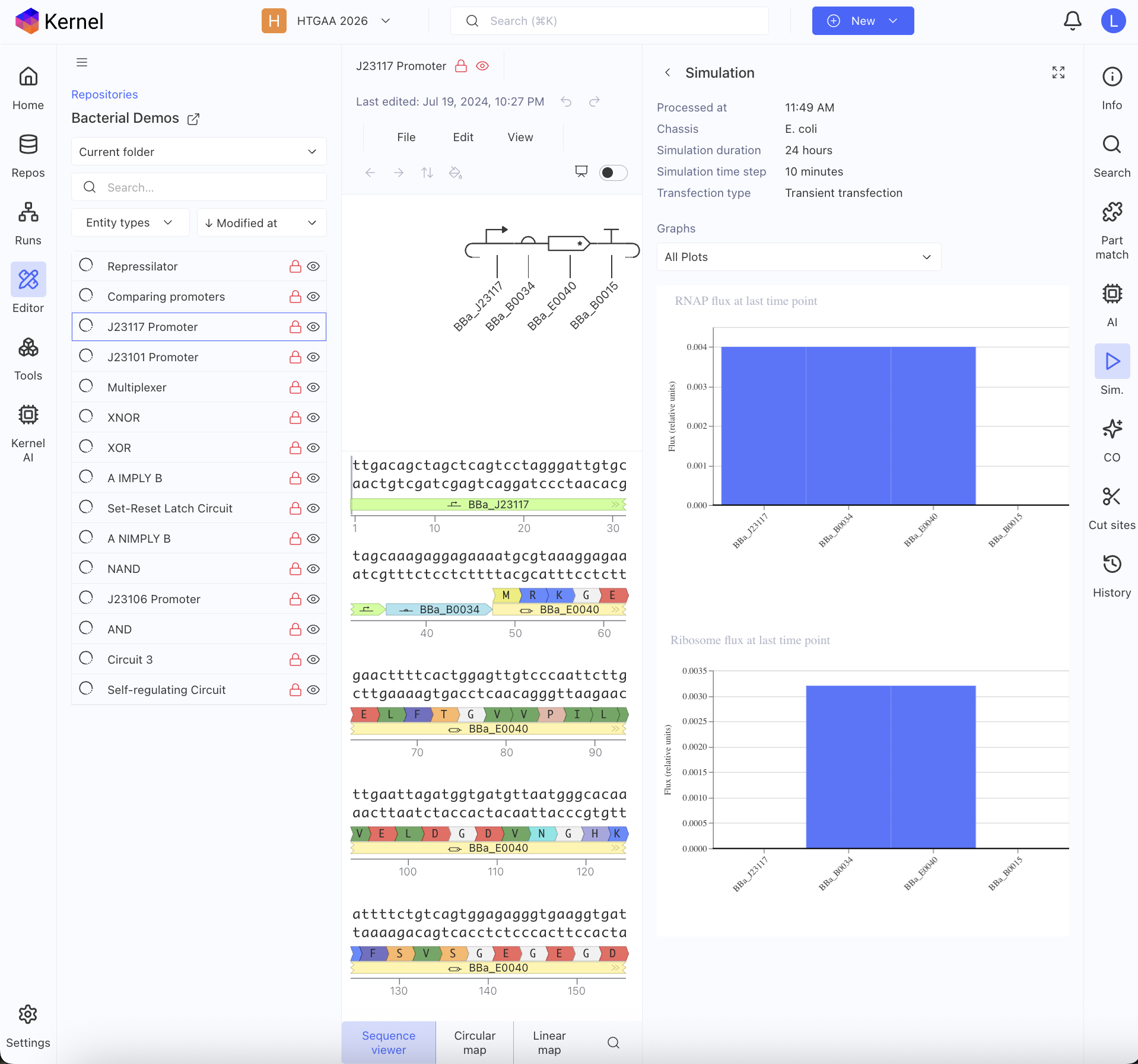
First, I just messed around with the Bacterial Demos Repository, and clicked o random parts to see what the platform felt like to me. i then went to the J23117 Promoter and viewed what was going on there. I went to the simulate button to simulate the process at a random timestep and duration of my choosing, with E. Coli as the chassis. Here is a screenshot of my results:

Then, I created a repressilator, with the comparison below of the one located in bacterial demos:
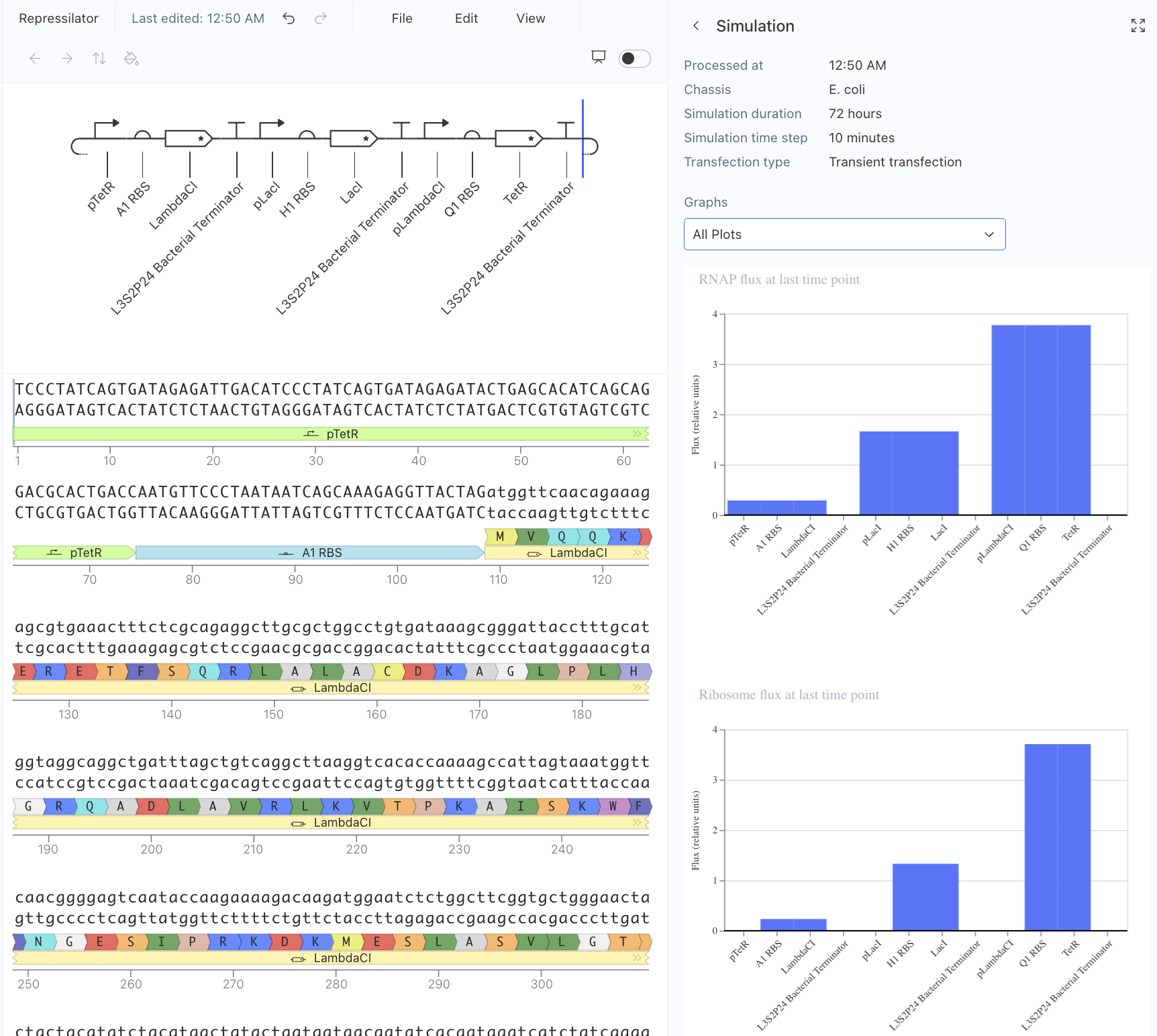

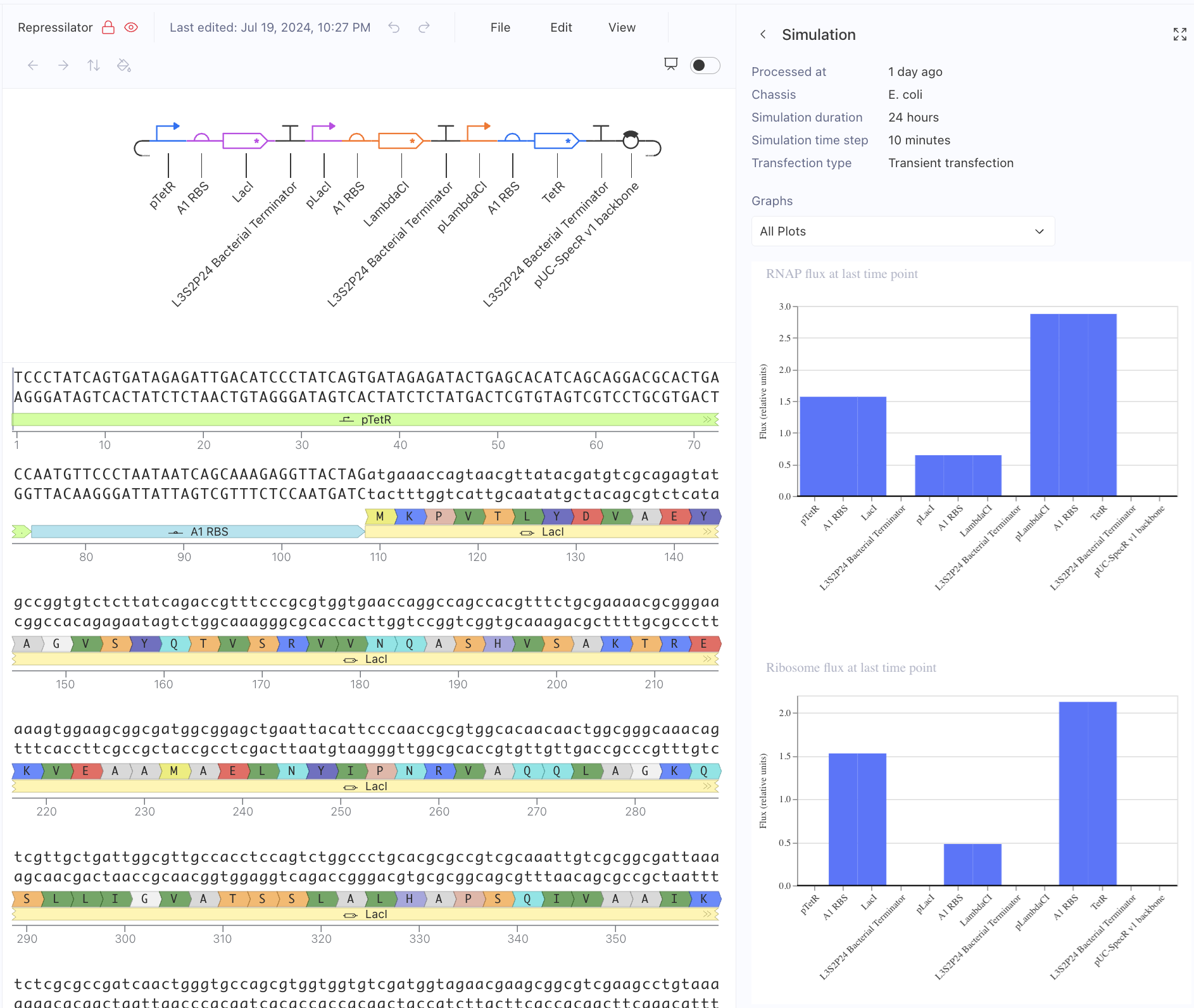
 I’m fairly confident that I did mine correctly as the simulations look quite similar, and the overall set-up contains similar components.
I’m fairly confident that I did mine correctly as the simulations look quite similar, and the overall set-up contains similar components.
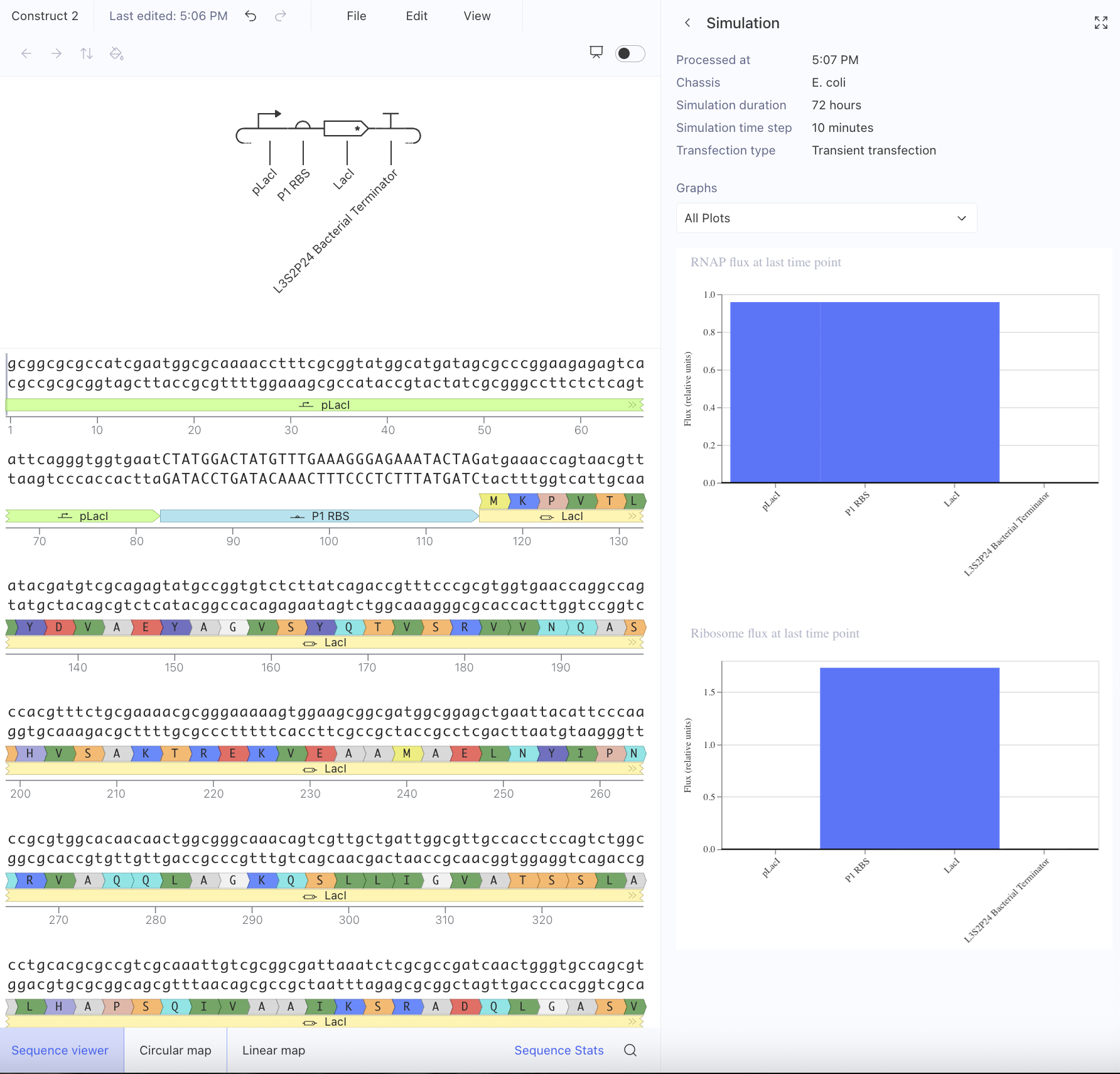
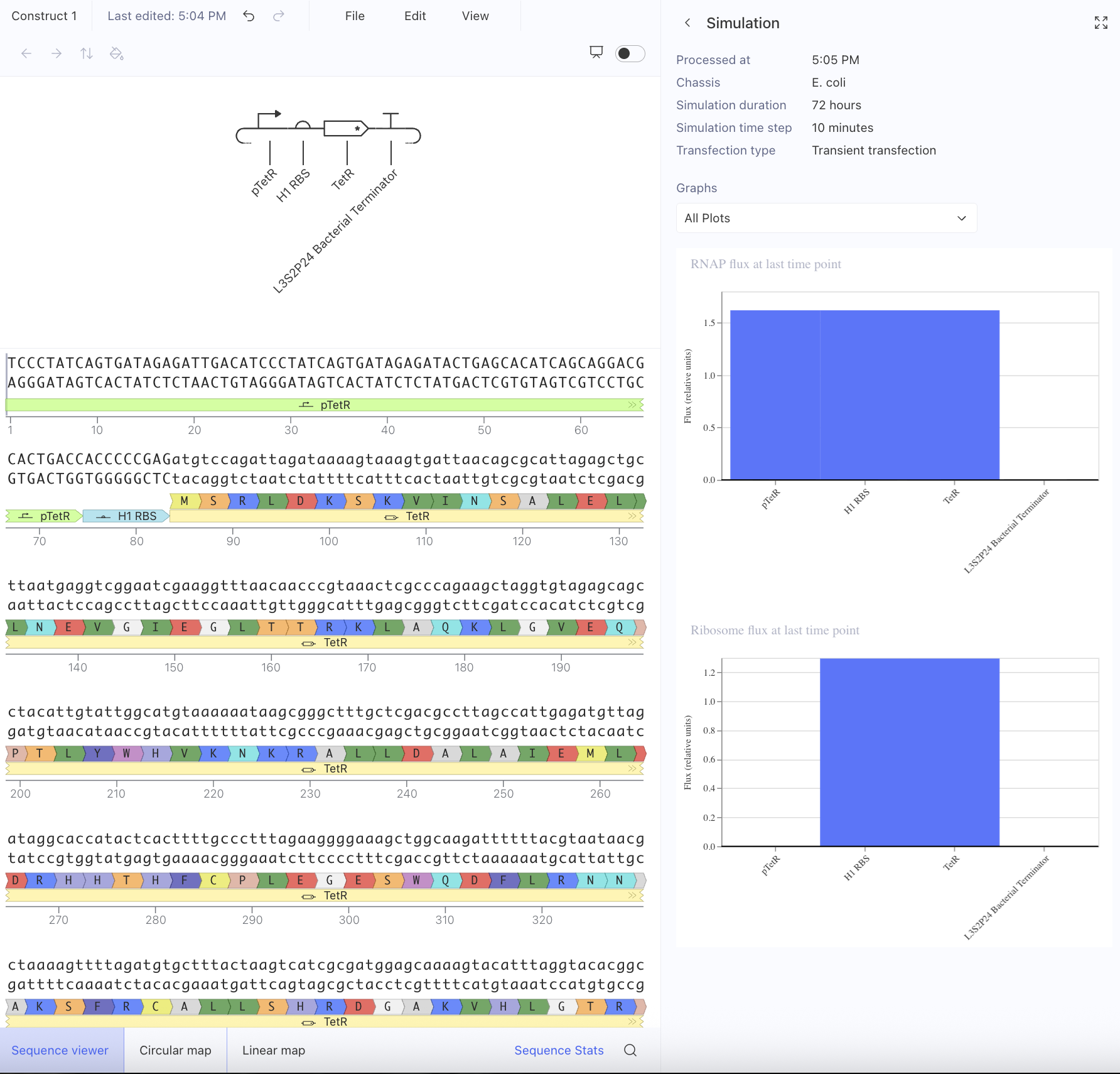
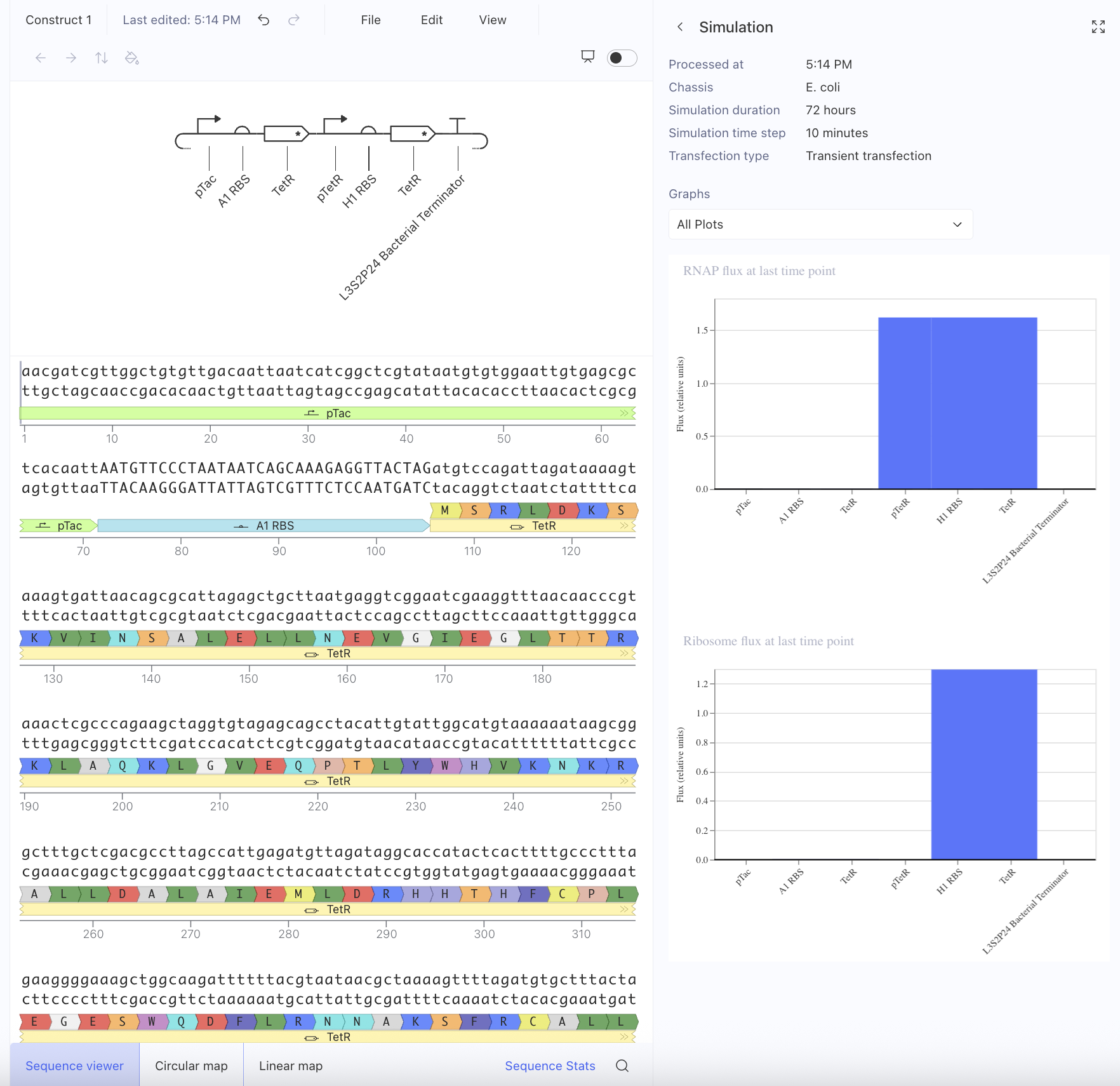
Afterwards, I just put together some random constructs after fumbling around Kernel forever. Not sure if there were guidelines but I thought they would be cool.
Predictions here:


I think overall, the simulations have similar outputs as to what my predictions are. Not very much to comment on either!