Week 3 Lab: Opentrons Artwork


Overview
Pre-Lab Process
The idea of having to write my own code to create this art sounded terrifying at first given that I probably barely passed 6.100A. Then I found out a lot of it was written in Google Colab and felt relieved, until I kept running into issues so I decided to give up and just use Ronan’s code from his website: https://opentrons-art.rcdonovan.com/
The code can be found at the bottom.
Coordinates:
mscarlet_i_points = [(-8.8, 17.6),(0, 17.6),(-11, 15.4),(-6.6, 15.4),(-2.2, 15.4),(2.2, 15.4),(-11, 13.2),(-6.6, 13.2),(-2.2, 13.2),(2.2, 13.2),(-11, 11),(-6.6, 11),(-2.2, 11),(2.2, 11),(13.2, 11),(15.4, 11),(19.8, 11),(22, 11),(24.2, 11),(26.4, 11),(28.6, 11),(-11, 8.8),(-6.6, 8.8),(-2.2, 8.8),(2.2, 8.8),(13.2, 8.8),(15.4, 8.8),(19.8, 8.8),(22, 8.8),(24.2, 8.8),(26.4, 8.8),(28.6, 8.8),(-11, 6.6),(-6.6, 6.6),(-2.2, 6.6),(2.2, 6.6),(13.2, 6.6),(15.4, 6.6),(26.4, 6.6),(28.6, 6.6),(-11, 4.4),(-6.6, 4.4),(-2.2, 4.4),(2.2, 4.4),(13.2, 4.4),(15.4, 4.4),(26.4, 4.4),(28.6, 4.4),(-11, 2.2),(-6.6, 2.2),(-2.2, 2.2),(2.2, 2.2),(13.2, 2.2),(15.4, 2.2),(22, 2.2),(24.2, 2.2),(26.4, 2.2),(28.6, 2.2),(-11, 0),(-6.6, 0),(-4.4, 0),(-2.2, 0),(2.2, 0),(13.2, 0),(15.4, 0),(22, 0),(24.2, 0),(26.4, 0),(28.6, 0),(-13.2, -2.2),(4.4, -2.2),(13.2, -2.2),(15.4, -2.2),(22, -2.2),(24.2, -2.2),(-15.4, -4.4),(6.6, -4.4),(13.2, -4.4),(15.4, -4.4),(22, -4.4),(24.2, -4.4),(-15.4, -6.6),(-8.8, -6.6),(0, -6.6),(6.6, -6.6),(-15.4, -8.8),(6.6, -8.8),(13.2, -8.8),(15.4, -8.8),(22, -8.8),(24.2, -8.8),(-15.4, -11),(6.6, -11),(13.2, -11),(15.4, -11),(22, -11),(24.2, -11),(-13.2, -13.2),(4.4, -13.2),(-11, -15.4),(-8.8, -15.4),(-6.6, -15.4),(-4.4, -15.4),(-2.2, -15.4),(0, -15.4),(2.2, -15.4)]
mko2_points = [(15.4, 33),(17.6, 33),(13.2, 30.8),(17.6, 30.8),(19.8, 30.8),(17.6, 28.6),(19.8, 28.6),(17.6, 26.4),(19.8, 26.4),(13.2, 24.2),(17.6, 24.2),(19.8, 24.2),(15.4, 22),(17.6, 22),(-28.6, 19.8),(-28.6, 17.6),(-26.4, 17.6),(-28.6, 15.4),(-26.4, 15.4),(-24.2, 15.4),(-22, 15.4),(-19.8, 15.4),(-17.6, 15.4),(-35.2, 13.2),(-33, 13.2),(-30.8, 13.2),(-28.6, 13.2),(-26.4, 13.2),(-24.2, 13.2),(-22, 13.2),(-33, 11),(-30.8, 11),(-28.6, 11),(-26.4, 11),(-24.2, 11),(-22, 11),(-19.8, 11),(-28.6, 8.8),(-26.4, 8.8),(-24.2, 8.8),(-22, 8.8),(-19.8, 8.8),(-17.6, 8.8),(-28.6, 6.6),(-26.4, 6.6),(-28.6, 4.4),(-28.6, 2.2),(19.8, -17.6),(19.8, -19.8),(19.8, -22),(17.6, -24.2),(19.8, -24.2),(22, -24.2),(13.2, -26.4),(15.4, -26.4),(17.6, -26.4),(19.8, -26.4),(22, -26.4),(24.2, -26.4),(26.4, -26.4),(17.6, -28.6),(19.8, -28.6),(22, -28.6),(19.8, -30.8),(19.8, -33)]
During the Lab
This was a pretty chill session. To be honest, I don’t know the details of how running the program works, but I was allowed to hit the start button! It was fascinating watching the robot do it’s thing. Because people came before me, if there was any troubleshooting that had to go on, it was done by the time I arrived.
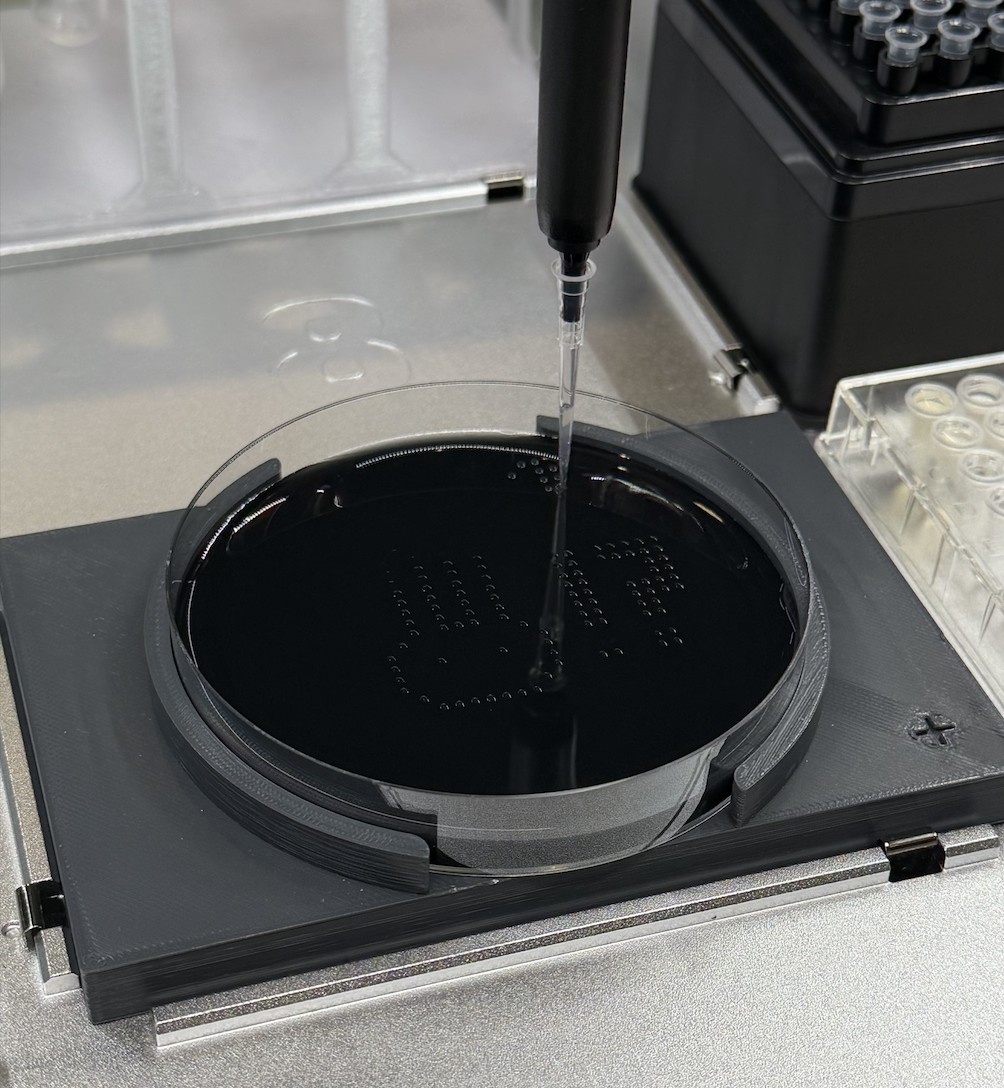


Post Lab
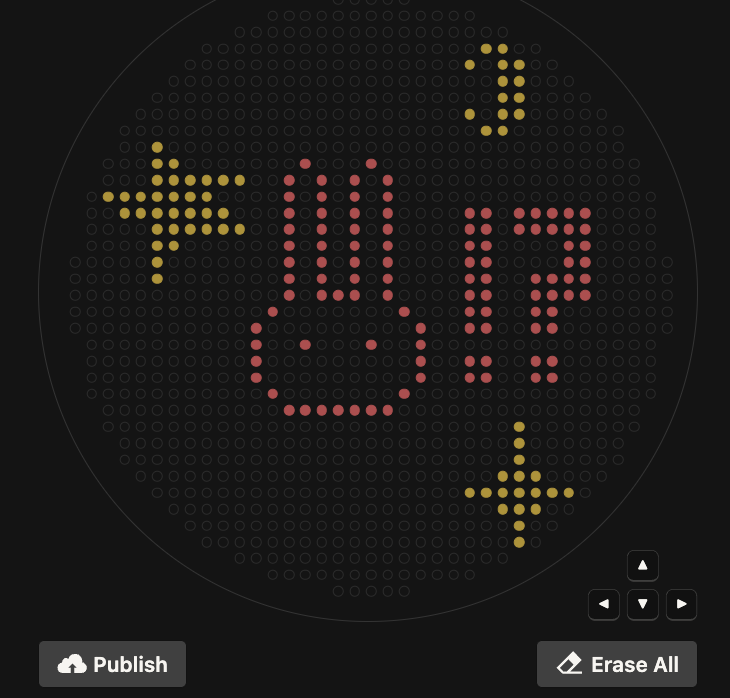
My design turned out super cute! I’m not super sure why there are tiny dots that appeared which don’t pertain to my design, perhaps this is contamination?


Post-Lab Reflection
I think I’m a materials major for a reason.
Code:
from opentrons import types
import string
metadata = {
'protocolName': '{YOUR NAME} - Opentrons Art - HTGAA',
'author': 'HTGAA',
'source': 'HTGAA 2026',
'apiLevel': '2.20'
}
Z_VALUE_AGAR = 2.0
POINT_SIZE = 0.75
mscarlet_i_points = [(-8.8,17.6), (0,17.6), (-11,15.4), (-6.6,15.4), (-2.2,15.4), (2.2,15.4), (-11,13.2), (-6.6,13.2), (-2.2,13.2), (2.2,13.2), (-11,11), (-6.6,11), (-2.2,11), (2.2,11), (13.2,11), (15.4,11), (19.8,11), (22,11), (24.2,11), (26.4,11), (28.6,11), (-11,8.8), (-6.6,8.8), (-2.2,8.8), (2.2,8.8), (13.2,8.8), (15.4,8.8), (19.8,8.8), (22,8.8), (24.2,8.8), (26.4,8.8), (28.6,8.8), (-11,6.6), (-6.6,6.6), (-2.2,6.6), (2.2,6.6), (13.2,6.6), (15.4,6.6), (26.4,6.6), (28.6,6.6), (-11,4.4), (-6.6,4.4), (-2.2,4.4), (2.2,4.4), (13.2,4.4), (15.4,4.4), (26.4,4.4), (28.6,4.4), (-11,2.2), (-6.6,2.2), (-2.2,2.2), (2.2,2.2), (13.2,2.2), (15.4,2.2), (22,2.2), (24.2,2.2), (26.4,2.2), (28.6,2.2), (-11,0), (-6.6,0), (-4.4,0), (-2.2,0), (2.2,0), (13.2,0), (15.4,0), (22,0), (24.2,0), (26.4,0), (28.6,0), (-13.2,-2.2), (4.4,-2.2), (13.2,-2.2), (15.4,-2.2), (22,-2.2), (24.2,-2.2), (-15.4,-4.4), (6.6,-4.4), (13.2,-4.4), (15.4,-4.4), (22,-4.4), (24.2,-4.4), (-15.4,-6.6), (-8.8,-6.6), (0,-6.6), (6.6,-6.6), (-15.4,-8.8), (6.6,-8.8), (13.2,-8.8), (15.4,-8.8), (22,-8.8), (24.2,-8.8), (-15.4,-11), (6.6,-11), (13.2,-11), (15.4,-11), (22,-11), (24.2,-11), (-13.2,-13.2), (4.4,-13.2), (-11,-15.4), (-8.8,-15.4), (-6.6,-15.4), (-4.4,-15.4), (-2.2,-15.4), (0,-15.4), (2.2,-15.4)]
mko2_points = [(15.4,33), (17.6,33), (13.2,30.8), (17.6,30.8), (19.8,30.8), (17.6,28.6), (19.8,28.6), (17.6,26.4), (19.8,26.4), (13.2,24.2), (17.6,24.2), (19.8,24.2), (15.4,22), (17.6,22), (-28.6,19.8), (-28.6,17.6), (-26.4,17.6), (-28.6,15.4), (-26.4,15.4), (-24.2,15.4), (-22,15.4), (-19.8,15.4), (-17.6,15.4), (-35.2,13.2), (-33,13.2), (-30.8,13.2), (-28.6,13.2), (-26.4,13.2), (-24.2,13.2), (-22,13.2), (-33,11), (-30.8,11), (-28.6,11), (-26.4,11), (-24.2,11), (-22,11), (-19.8,11), (-28.6,8.8), (-26.4,8.8), (-24.2,8.8), (-22,8.8), (-19.8,8.8), (-17.6,8.8), (-28.6,6.6), (-26.4,6.6), (-28.6,4.4), (-28.6,2.2), (19.8,-17.6), (19.8,-19.8), (19.8,-22), (17.6,-24.2), (19.8,-24.2), (22,-24.2), (13.2,-26.4), (15.4,-26.4), (17.6,-26.4), (19.8,-26.4), (22,-26.4), (24.2,-26.4), (26.4,-26.4), (17.6,-28.6), (19.8,-28.6), (22,-28.6), (19.8,-30.8), (19.8,-33)]
point_name_pairing = [("mscarlet_i", mscarlet_i_points),("mko2", mko2_points)]
# Robot deck setup constants
TIP_RACK_DECK_SLOT = 9
COLORS_DECK_SLOT = 6
AGAR_DECK_SLOT = 5
PIPETTE_STARTING_TIP_WELL = 'A1'
# Place the PCR tubes in this order
well_colors = {
'A1': 'sfGFP',
'A2': 'mRFP1',
'A3': 'mKO2',
'A4': 'Venus',
'A5': 'mKate2_TF',
'A6': 'Azurite',
'A7': 'mCerulean3',
'A8': 'mClover3',
'A9': 'mJuniper',
'A10': 'mTurquoise2',
'A11': 'mBanana',
'A12': 'mPlum',
'B1': 'Electra2',
'B2': 'mWasabi',
'B3': 'mScarlet_I',
'B4': 'mPapaya',
'B5': 'eqFP578',
'B6': 'tdTomato',
'B7': 'DsRed',
'B8': 'mKate2',
'B9': 'EGFP',
'B10': 'mRuby2',
'B11': 'TagBFP',
'B12': 'mChartreuse_TF',
'C1': 'mLychee_TF',
'C2': 'mTagBFP2',
'C3': 'mEGFP',
'C4': 'mNeonGreen',
'C5': 'mAzamiGreen',
'C6': 'mWatermelon',
'C7': 'avGFP',
'C8': 'mCitrine',
'C9': 'mVenus',
'C10': 'mCherry',
'C11': 'mHoneydew',
'C12': 'TagRFP',
'D1': 'mTFP1',
'D2': 'Ultramarine',
'D3': 'ZsGreen1',
'D4': 'mMiCy',
'D5': 'mStayGold2',
'D6': 'PA_GFP'
}
volume_used = {
'mscarlet_i': 0,
'mko2': 0
}
def update_volume_remaining(current_color, quantity_to_aspirate):
rows = string.ascii_uppercase
for well, color in list(well_colors.items()):
if color == current_color:
if (volume_used[current_color] + quantity_to_aspirate) > 250:
# Move to next well horizontally by advancing row letter, keeping column number
row = well[0]
col = well[1:]
# Find next row letter
next_row = rows[rows.index(row) + 1]
next_well = f"{next_row}{col}"
del well_colors[well]
well_colors[next_well] = current_color
volume_used[current_color] = quantity_to_aspirate
else:
volume_used[current_color] += quantity_to_aspirate
break
def run(protocol):
# Load labware, modules and pipettes
protocol.home()
# Tips
tips_20ul = protocol.load_labware('opentrons_96_tiprack_20ul', TIP_RACK_DECK_SLOT, 'Opentrons 20uL Tips')
# Pipettes
pipette_20ul = protocol.load_instrument("p20_single_gen2", "right", [tips_20ul])
# PCR Plate
temperature_plate = protocol.load_labware('opentrons_96_aluminumblock_generic_pcr_strip_200ul', 6)
# Agar Plate
agar_plate = protocol.load_labware('htgaa_agar_plate', AGAR_DECK_SLOT, 'Agar Plate')
agar_plate.set_offset(x=0.00, y=0.00, z=Z_VALUE_AGAR)
# Get the top-center of the plate, make sure the plate was calibrated before running this
center_location = agar_plate['A1'].top()
pipette_20ul.starting_tip = tips_20ul.well(PIPETTE_STARTING_TIP_WELL)
# Helper function (dispensing)
def dispense_and_jog(pipette, volume, location):
assert(isinstance(volume, (int, float)))
# Go above the location
above_location = location.move(types.Point(z=location.point.z + 2))
pipette.move_to(above_location)
# Go downwards and dispense
pipette.dispense(volume, location)
# Go upwards to avoid smearing
pipette.move_to(above_location)
# Helper function (color location)
def location_of_color(color_string):
for well,color in well_colors.items():
if color.lower() == color_string.lower():
return temperature_plate[well]
raise ValueError(f"No well found with color {color_string}")
# Print pattern by iterating over lists
for i, (current_color, point_list) in enumerate(point_name_pairing):
# Skip the rest of the loop if the list is empty
if not point_list:
continue
# Get the tip for this run, set the bacteria color, and the aspirate bacteria of choice
pipette_20ul.pick_up_tip()
max_aspirate = int(18 // POINT_SIZE) * POINT_SIZE
quantity_to_aspirate = min(len(point_list)*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Iterate over the current points list and dispense them, refilling along the way
for i in range(len(point_list)):
x, y = point_list[i]
adjusted_location = center_location.move(types.Point(x, y))
dispense_and_jog(pipette_20ul, POINT_SIZE, adjusted_location)
if pipette_20ul.current_volume == 0 and len(point_list[i+1:]) > 0:
quantity_to_aspirate = min(len(point_list[i:])*POINT_SIZE, max_aspirate)
update_volume_remaining(current_color, quantity_to_aspirate)
pipette_20ul.aspirate(quantity_to_aspirate, location_of_color(current_color))
# Drop tip between each color
pipette_20ul.drop_tip()