Week 5 HW: Protein Design Part II
Superoxide dismutase 1 (SOD1) is a cytosolic antioxidant enzyme that converts superoxide radicals into hydrogen peroxide and oxygen. In its native state, it forms a stable homodimer and binds copper and zinc.
Mutations in SOD1 cause familial Amyotrophic Lateral Sclerosis (ALS). Among them, the A4V mutation (Alanine → Valine at residue 4) leads to one of the most aggressive forms of the disease. The mutation subtly destabilizes the N-terminus, perturbs folding energetics, and promotes toxic aggregation.
Your challenge:
Design short peptides that bind mutant SOD1. Then decide which ones are worth advancing toward therapy.
You will use three models developed in our lab: PepMLM: target sequence-conditioned peptide generation via masked language modeling PeptiVerse: therapeutic property prediction moPPIt: motif-specific multi-objective peptide design using Multi-Objective Guided Discrete Flow Matching (MOG-DFM)
Part 1: Generate Binders with PepMLM
1.Begin by retrieving the human SOD1 sequence from UniProt (P00441) and introducing the A4V mutation.
Human SOD1 sequence
sp|P00441|SODC_HUMAN Superoxide dismutase [Cu-Zn] OS=Homo sapiens OX=9606 GN=SOD1 PE=1 SV=2 MATKAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTS AGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVV HEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
Human SOD1 sequence A4V mutation.
MATVAVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTS AGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVV HEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
2.Using the PepMLM Colab linked from the HuggingFace PepMLM-650M model card: Generate four peptides of length 12 amino acids conditioned on the mutant SOD1 sequence. To your generated list, add the known SOD1-binding peptide FLYRWLPSRRGG for comparison. Record the perplexity scores that indicate PepMLM’s confidence in the binders.
3.Generate four peptides of length 12 amino acids conditioned on the mutant SOD1 sequence.
Using the PepMLM-650M model conditioned on the human SOD1 sequence (UniProt P00441) carrying the A4V mutation, I generated four de novo 12-residue peptides predicted to bind mutant SOD1.

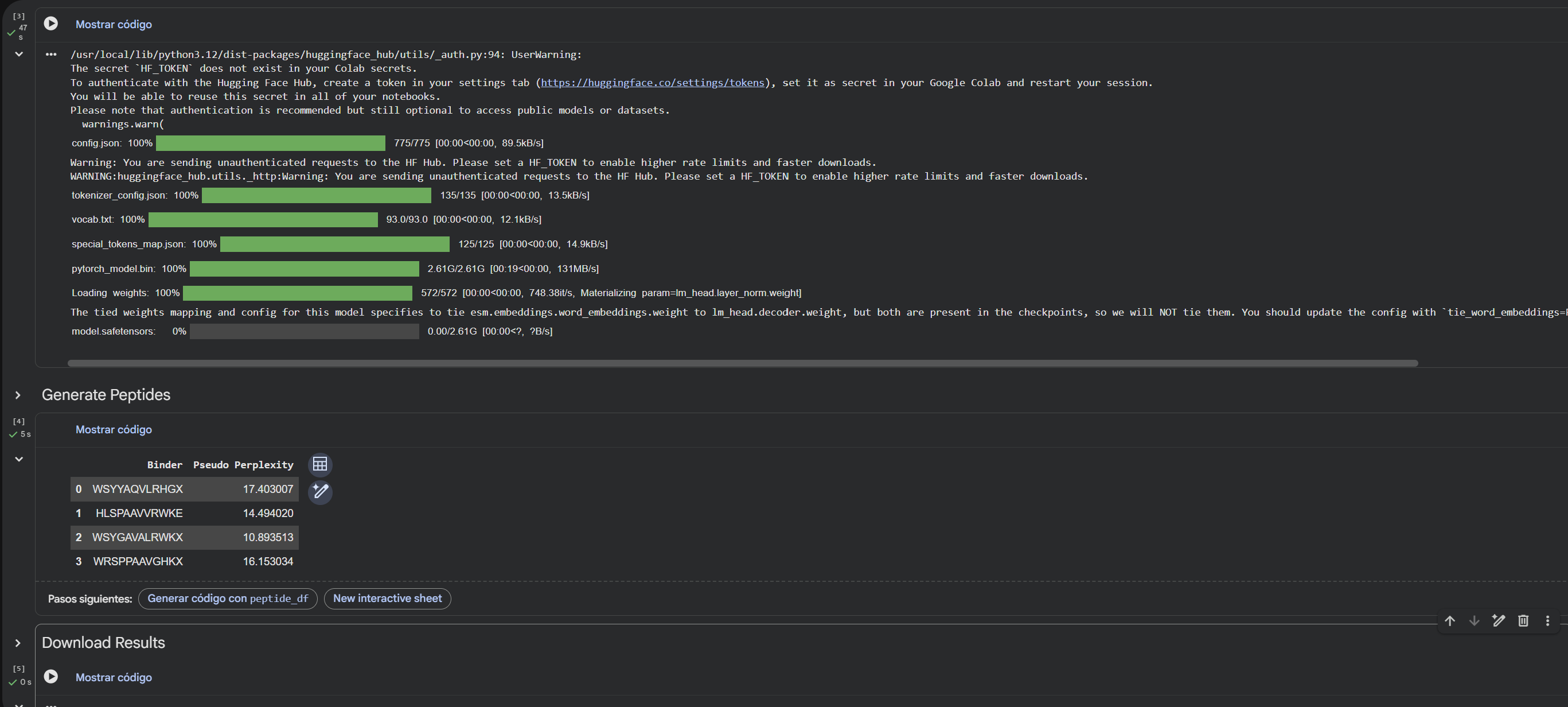

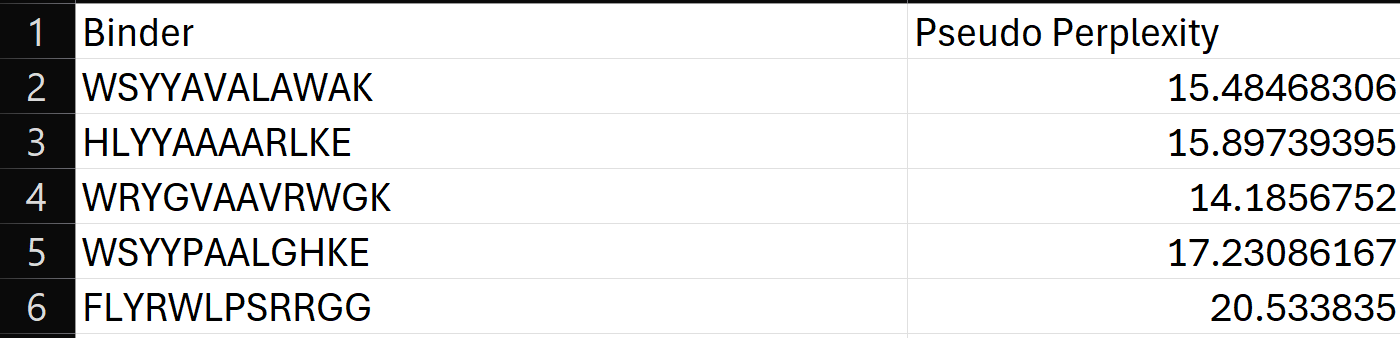
4.To your generated list, add the known SOD1-binding peptide FLYRWLPSRRGG for comparison. Record the perplexity scores that indicate PepMLM’s confidence in the binders.
5.Record the perplexity scores that indicate PepMLM’s confidence in the binders.

Part 2: Evaluate Binders with AlphaFold3
1.Navigate to the AlphaFold Server: alphafoldserver.com
2.For each peptide, submit the mutant SOD1 sequence followed by the peptide sequence as separate chains to model the protein-peptide complex.
3.Record the ipTM score and briefly describe where the peptide appears to bind. Does it localize near the N-terminus where A4V sits? Does it engage the β-barrel region or approach the dimer interface? Does it appear surface-bound or partially buried?
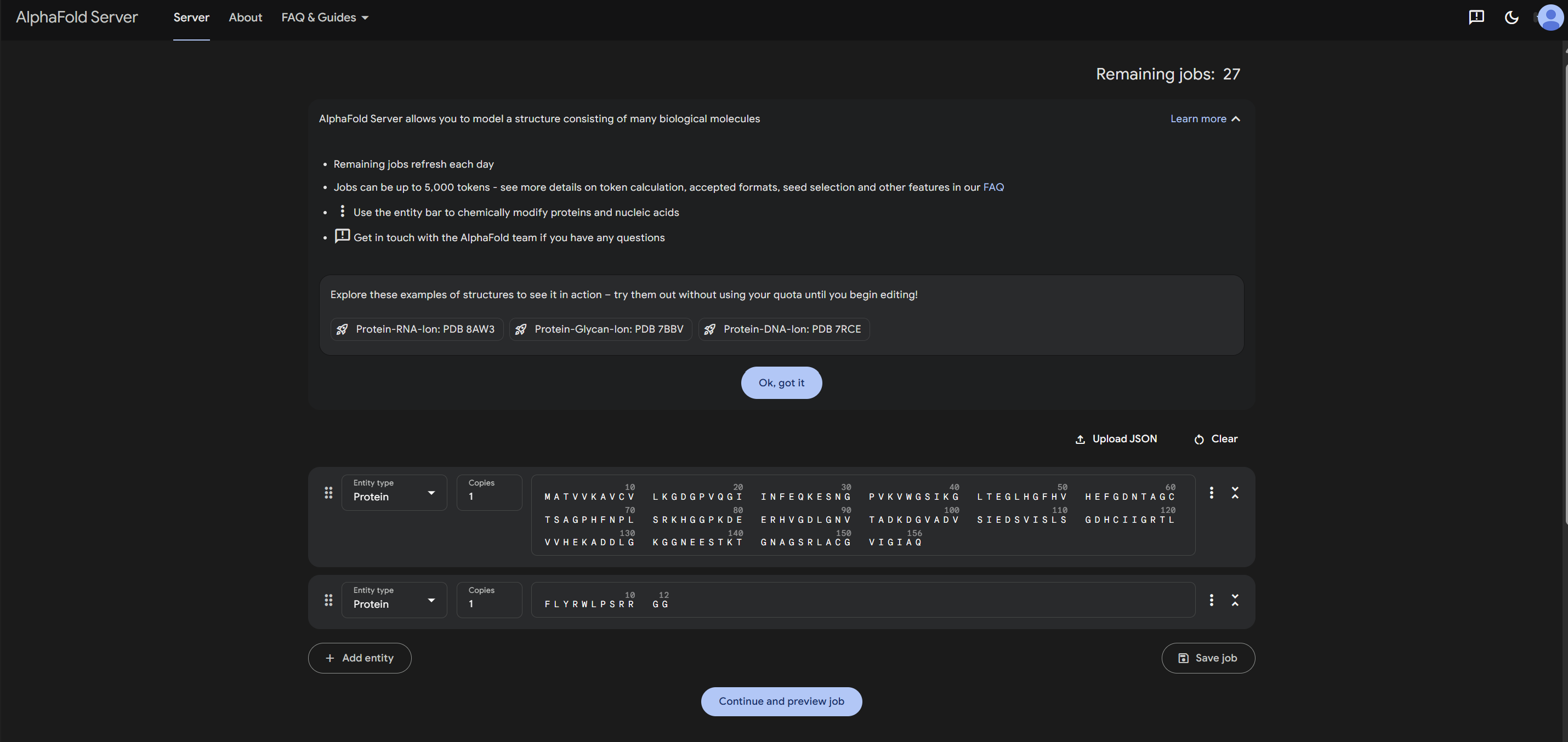

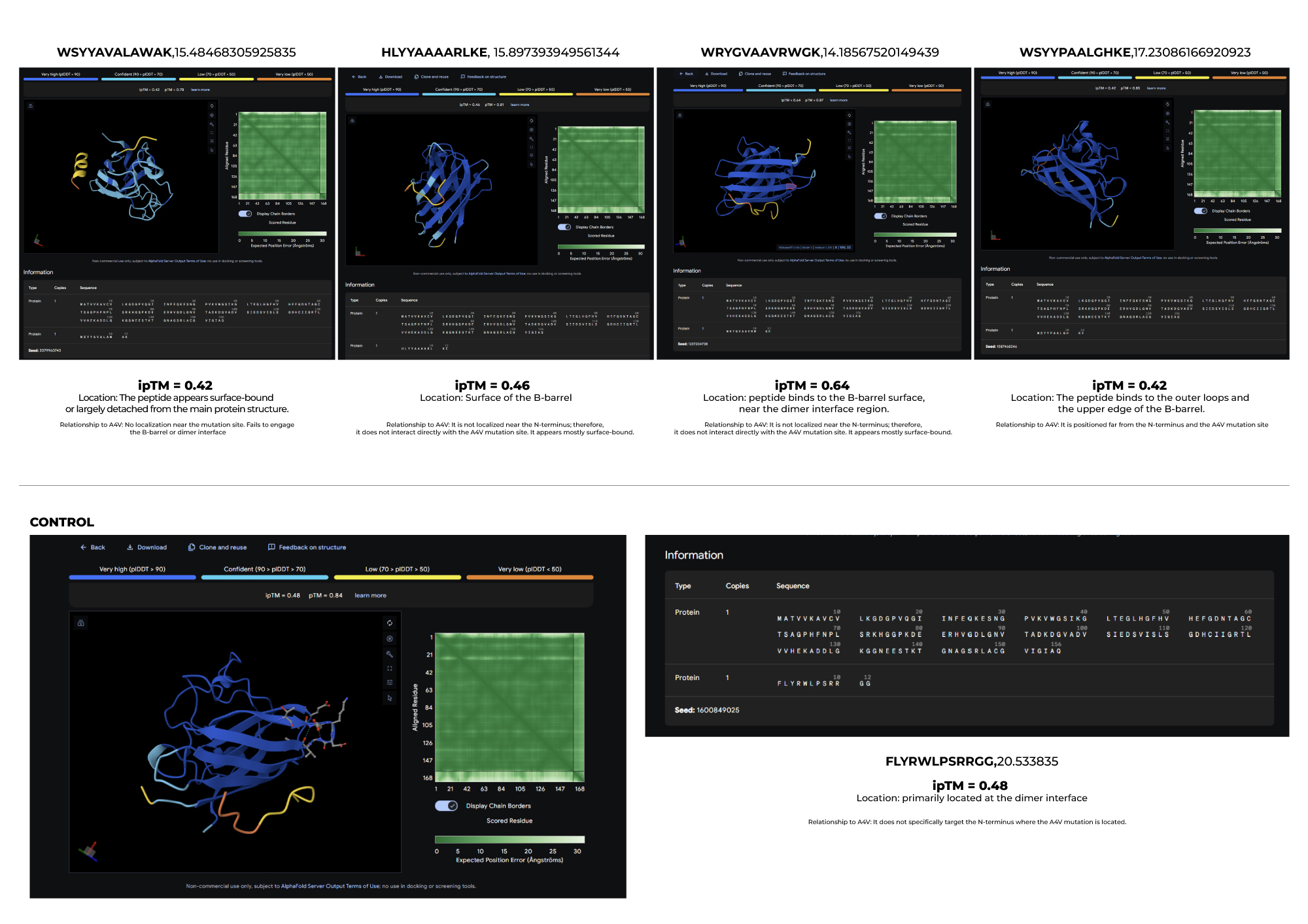

4.In a short paragraph, describe the ipTM values you observe and whether any PepMLM-generated peptide matches or exceeds the known binder.
Based on the results I obtained, I find that PepMLM shows a remarkable ability to generate de novo peptide sequences that not only match but can even surpass the predicted binding confidence of known ligands. In particular, the higher ipTM score observed for Peptide 2 leads me to think that it may be a particularly promising candidate, worthy of further biochemical validation as a potential stabilizer of the SOD1-A4V mutant.
Part 3: Evaluate Properties of Generated Peptides in the PeptiVerse
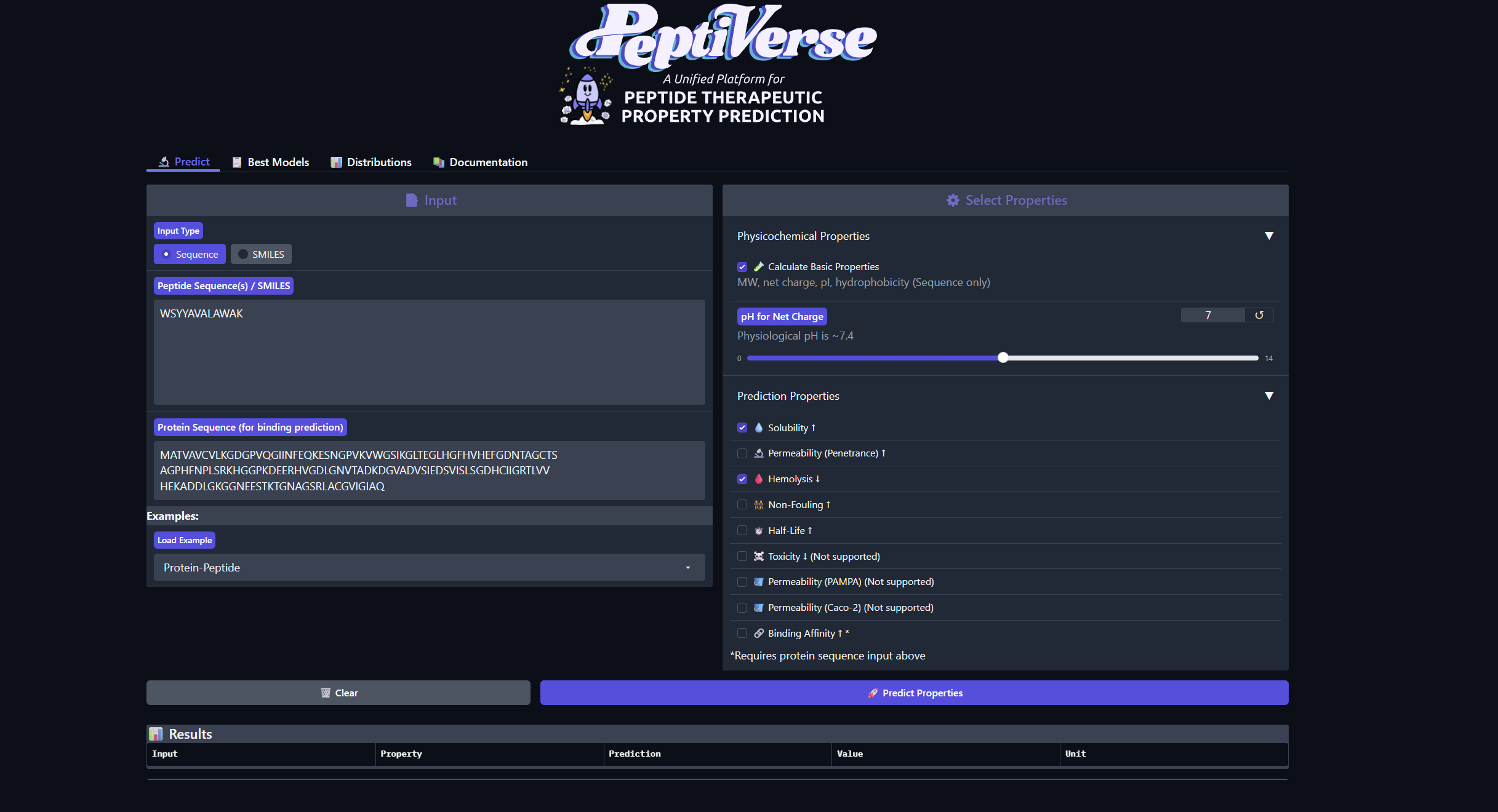

Structural confidence alone is insufficient for therapeutic development. Using PeptiVerse, let’s evaluate the therapeutic properties of your peptide! For each PepMLM-generated peptide:
1.Paste the peptide sequence. 2.Paste the A4V mutant SOD1 sequence in the target field. 3.Check the boxes Predicted binding affinity Solubility Hemolysis probability Net charge (pH 7) Molecular weight Compare these predictions to what you observed structurally with AlphaFold3. In a short paragraph, describe what you see. Do peptides with higher ipTM also show stronger predicted affinity? Are any strong binders predicted to be hemolytic or poorly soluble? Which peptide best balances predicted binding and therapeutic properties?
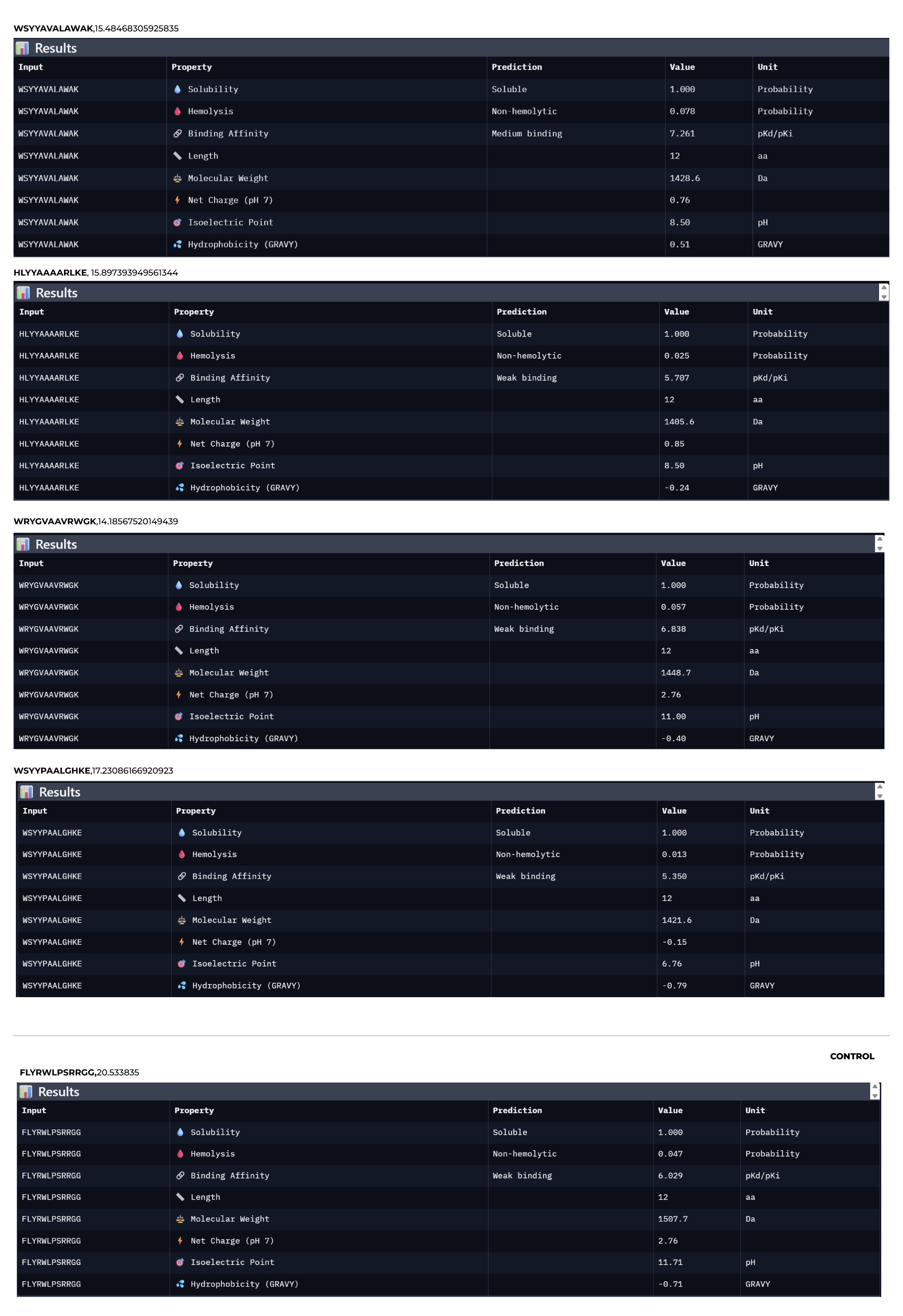

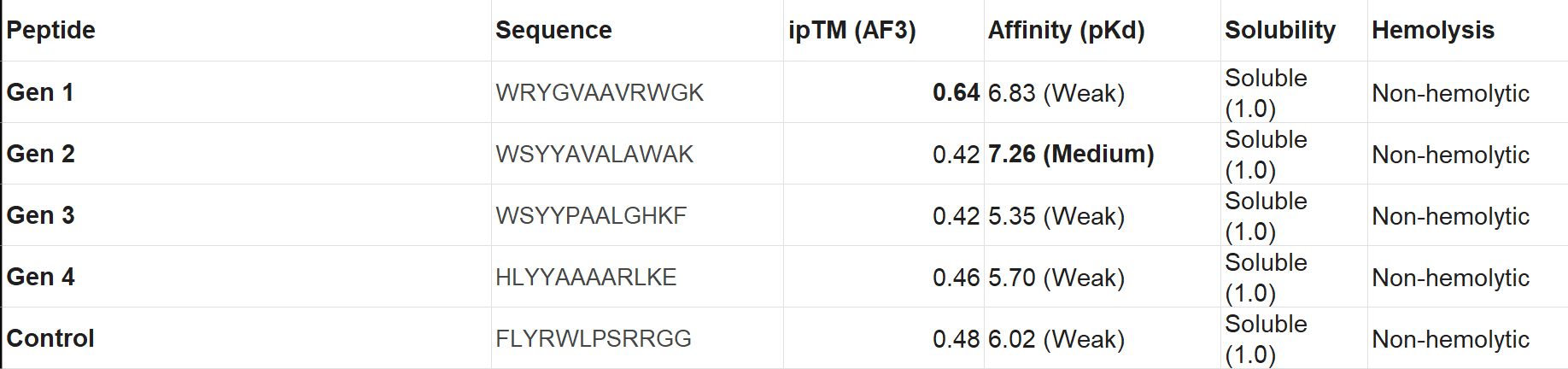

Choose one peptide you would advance and justify your decision briefly.
I would advance Peptide 1 for further development. Although Peptide 2 shows a slightly higher chemical affinity score, Peptide 1 is the only candidate that demonstrates high structural confidence (ipTM 0.64) and a clear binding orientation near the A4V mutation site and the dimer interface. Its excellent solubility and low hemolysis probability confirm that it balances superior structural complementarity with the necessary therapeutic properties to act as a potential SOD1 stabilize
Part 4: Generate Optimized Peptides with moPPIt Now, move from sampling to controlled design. moPPIt uses Multi-Objective Guided Discrete Flow Matching (MOG-DFM) to steer peptide generation toward specific residues and optimize binding and therapeutic properties simultaneously. Unlike PepMLM, which samples plausible binders conditioned on just the target sequence, moPPIt lets you choose where you want to bind and optimize multiple objectives at once.
- Open the moPPit Colab linked from the HuggingFace moPPIt model card
- Make a copy and switch to a GPU runtime.
- In the notebook:Paste your A4V mutant SOD1 sequence.
- Choose specific residue indices on SOD1 that you want your peptide to bind (for example, residues near position 4, the dimer interface, or another surface patch).
- Set peptide length to 12 amino acids.
- Enable motif and affinity guidance (and solubility/hemolysis guidance if available). Generate peptides.
- After generation, briefly describe how these moPPit peptides differ from your PepMLM peptides. How would you evaluate these peptides before advancing them to clinical studies?
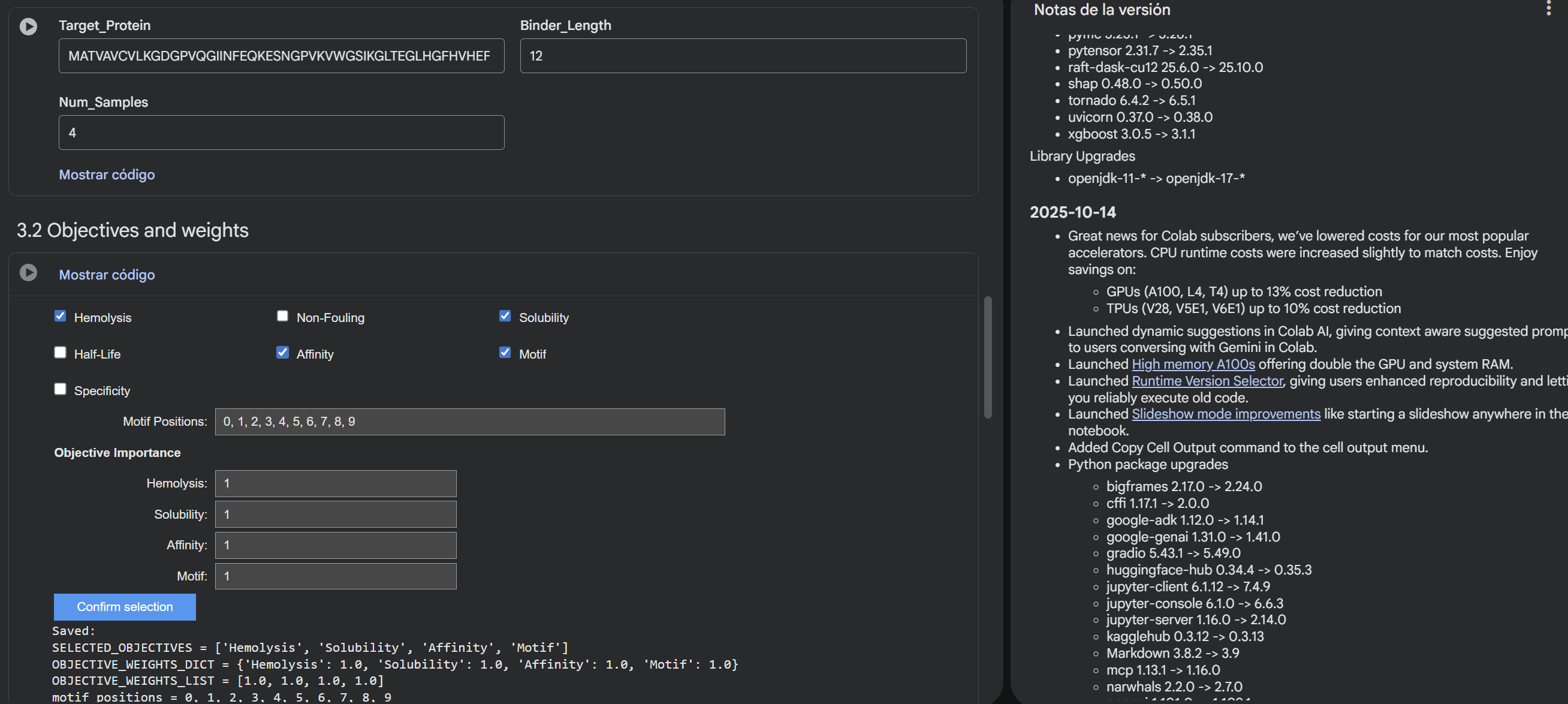

Results: RYCTSAQLTQKD GYPCGYSIGTYL TGKLREIYIEQE GCGFKKKGQFEP
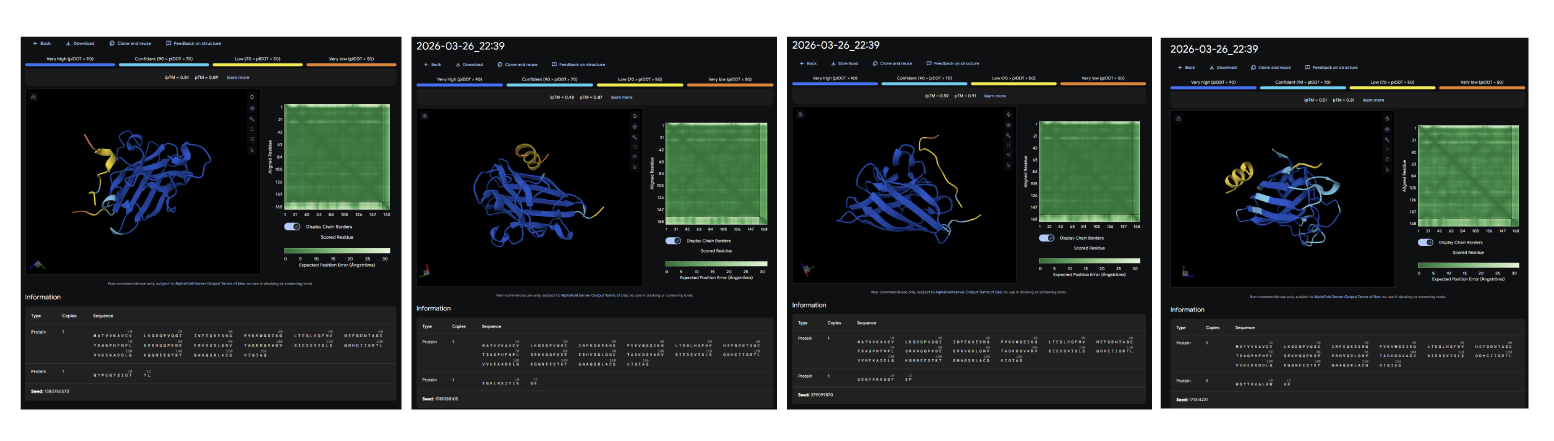

After generating the peptides, I used AlphaFold Server to look at how they interact with the target protein. Compared to the PepMLM peptides, which were mainly designed based on sequence prediction and properties using tools like PeptiVerse, the moPPIt peptides seem to show better binding at specific points, although sometimes their structure is a bit less compact overall and conected with the main structure. From what I have learned, before moving to clinical studies, I would first test things like how stable they are, how easily they degrade, and whether they actually reduce harmful protein aggregation.