Week 6 HW: Genetic Circuits Part I: Assembly Technologies
Assignment: DNA Assembly
1.What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
According to the protocol, the Phusion High-Fidelity PCR Master Mix is used to amplify specific DNA regions and prepare the necessary fragments for Gibson Assembly with high precision and minimal errors.
This pre-mixed solution is composed of four key elements. First, the high-fidelity DNA polymerase, which is responsible for copying the DNA and ensuring accurate replication with minimal errors. Second, dNTPs, which function as the building blocks to assemble the new DNA strands. Then, the reaction buffer, which maintains the chemical environment, such as pH, at optimal levels. And finally, magnesium an essential component that activates the polymerase so it can start joining the blocks together.
2.What are some factors that determine primer annealing temperature during PCR?
Following what was mentioned in class, several factors determine the primer annealing temperature, primarily the length and specific sequence of the primers. According to the protocol, primers should be 18–22 bp long with a GC content of 40–60%, including a GC-clamp to ensure specific binding. These elements define the melting temperature, and the is Ta is typically set 2–5°C below that value. In practice, this results in temperatures that typically range between 50°C and 65°C for a duration of 5 to 30 seconds, allowing the primers to bind accurately to the template DNA.
3.There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
PCR and restriction enzyme digests both generate linear DNA fragments, but they do so in different ways and at different points in the cloning workflow. In my view, one of the most important differences is that PCR can be carried out entirely in a small PCR tube, inside a thermocycler, as a cell‑free reaction that directly copies and amplifies the DNA. In contrast, restriction enzyme digestion only cuts the DNA at specific sites; it does not copy it. The actual replication of the fragment occurs only after the digested DNA is ligated into a vector and introduced into a living host (such as E. coli), which then replicates the plasmid during cell growth.
PCR: is a method to copy a specific piece of DNA many times. It is done in a small tube inside a machine called a thermocycler, where you add the original DNA (the template), two primers that are short DNA sequences marking the start and end of the region you want to copy, a polymerase enzyme that copies the DNA, and the nucleotides that are the “letters” of DNA needed to make the new strands.
This process is divided in 3 steps:
- Denaturation: it heats the DNA so that it opens up and separates into two strands.
- Annealing: it cools down a bit so that the primers can stick to their matching sites on the DNA.
- Extension: it heats up a little more so that the polymerase can copy the region between the primers.
RED: Restriction enzyme digestion is a method that cuts DNA at very specific places. It is done in a small tube, where the DNA is mixed with a restriction enzyme and a buffer that has the right conditions, such as pH and salt. The enzyme recognizes a short sequence of letters in the DNA and cuts it at that site, generating linear fragments. But the enzyme only cuts the DNA; it does not copy it. The actual copying happens later, when the cut fragment is introduced into a bacterium (like E. coli) and the cell replicates it as it divides.
Now that I understand it better, I think restriction enzyme digestion is “easier to replicate” in the sense that, once the DNA is inside a living cell like E. coli, the cell copies the plasmid by itself while it grows, and I don’t have to control temperature cycles like in PCR. However, the technique still needs many lab steps—cutting the DNA, ligating the insert into the vector, doing the transformation, and growing the bacteria—and uses expensive materials like restriction enzymes and ligase, so it is not always simpler or cheaper than it first seems. In terms of when to use each method, PCR is better when you want to amplify a specific region and introduce mutations. While restriction enzyme digestion is better when the insert and the vector already have compatible restriction sites and you want to do simple cut‑and‑ligate cloning.
4.How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
To ensure that the DNA sequences are appropriate for Gibson Assembly, three main requirements from the protocol must be met.
Mechanism: To ensure the assembly works, the DNA must be compatible with the four-step enzymatic process
1.Exonuclease 2.Annealing 3.Polymerase 4.LigaseOverlaps: The fragments must have 20–40 bp of sequence identity at their ends.
Molar Ratios: You must use a 2:1 (insert:vector) molar ratio
5.How does the plasmid DNA enter the E. coli cells during transformation?
During transformation, the plasmid DNA enters the E. coli cells thanks to a heat shock step. The cells are first made more able to take up DNA in the cold, then briefly heated to 42°C and cooled again. This creates temporary openings in the cell membrane so the plasmid can get inside by diffusion. After transformation, only the cells that have taken up the plasmid can survive on the antibiotic‑containing plates and start to grow.
6.Describe another assembly method in detail (such as Golden Gate Assembly)
I’m still trying to understand this, but I understood that Golden Gate Assembly is another method to join pieces of DNA into a plasmid, and it uses a special type of restriction enzyme (like BsaI) together with a ligase in the same tube. In this method, the BsaI enzyme cuts the DNA a little beyond its recognition site, leaving ends that can be custom‑designed. The thermocycler repeats cycles of cutting and ligating, and at the end you get a circular plasmid with all the fragments in the correct order, without the original restriction sites.
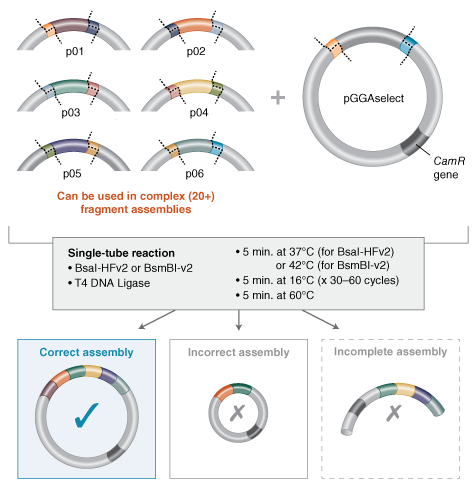

Source:https://www.neosynbio.com/golden-gate-assembly
7.Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online). Model this assembly method with Benchling or Asimov Kernel!
First, I design the DNA fragments and the vector so that each fragment has a unique end (overhang) that will only fit the next fragment, like puzzle pieces in a fixed order.
Prepare the DNA fragments and make sure each one has the BsaI cut site with the custom overhang on each side.
In a small tube, I mix all the fragments, the circular vector, the restriction enzyme BsaI, DNA ligase, ATP, and the reaction buffer.
I put the tube in a thermocycler and run many cycles where the enzyme cuts the DNA a little outside its site and the ligase rejoins the compatible ends.
After the reaction, the main product is a circular plasmid where all fragments are joined in the correct order, and the BsaI sites disappear from the final construct.
Then I transform this plasmid into competent E. coli cells, select the colonies on antibiotic plates, and check the correct assembly by PCR or digestion.