Week 2 HW: DNA Read, Write, & Edit
HW 2: DNA Read, Write, & Edit
Part 1: Benchling & In-silico Gel Art
Own experience on:
Make a free account at benchling.com Import the Lambda DNA.
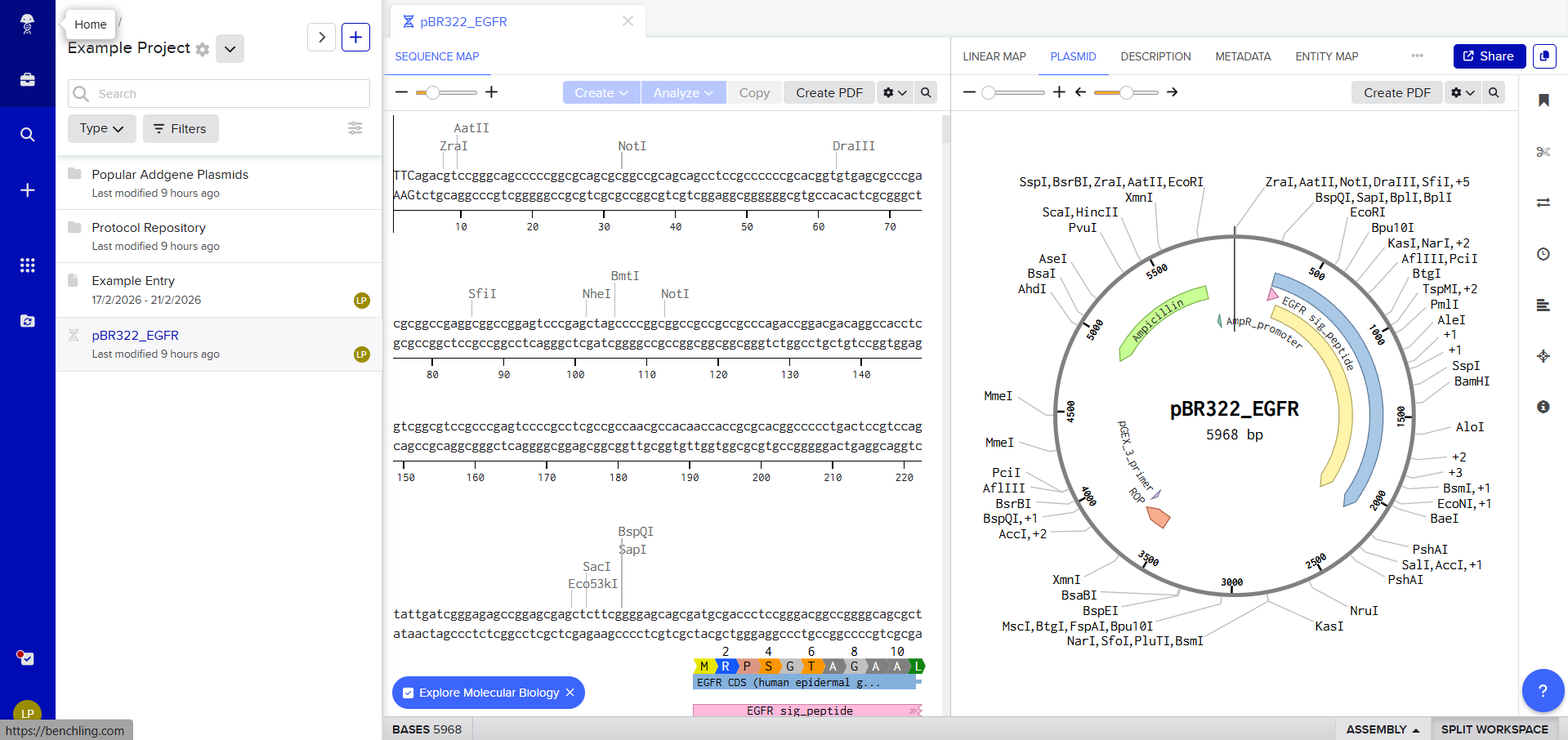
The first view of Benchling workspace
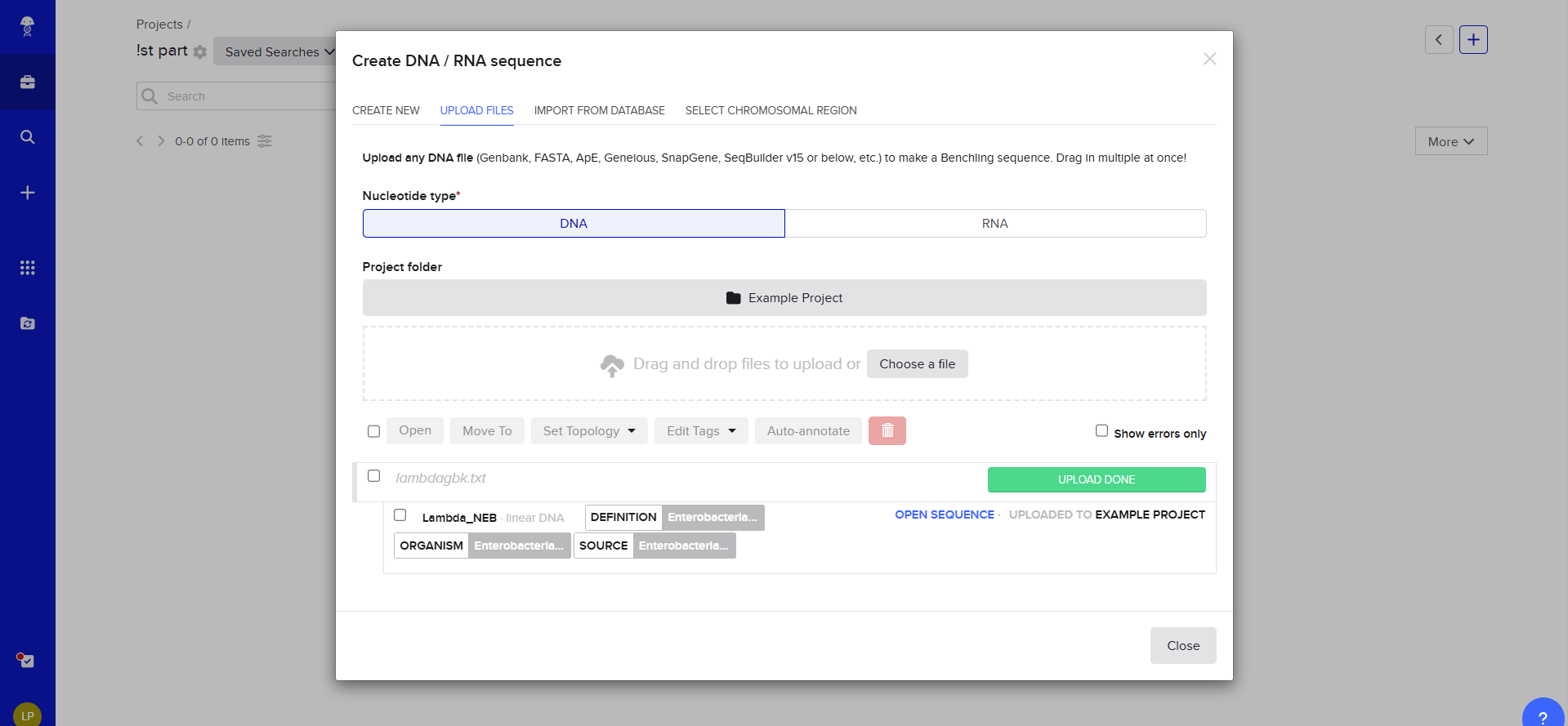
 Then i transfered de Lambda DNA, here we have two option for DNA document type, either we can use Genbank or, but Benchlink does have more options.
Then i transfered de Lambda DNA, here we have two option for DNA document type, either we can use Genbank or, but Benchlink does have more options.
 Upload to Benchling
Upload to Benchling
 So we can now view it on Benchling and start exploring the different options available
So we can now view it on Benchling and start exploring the different options available
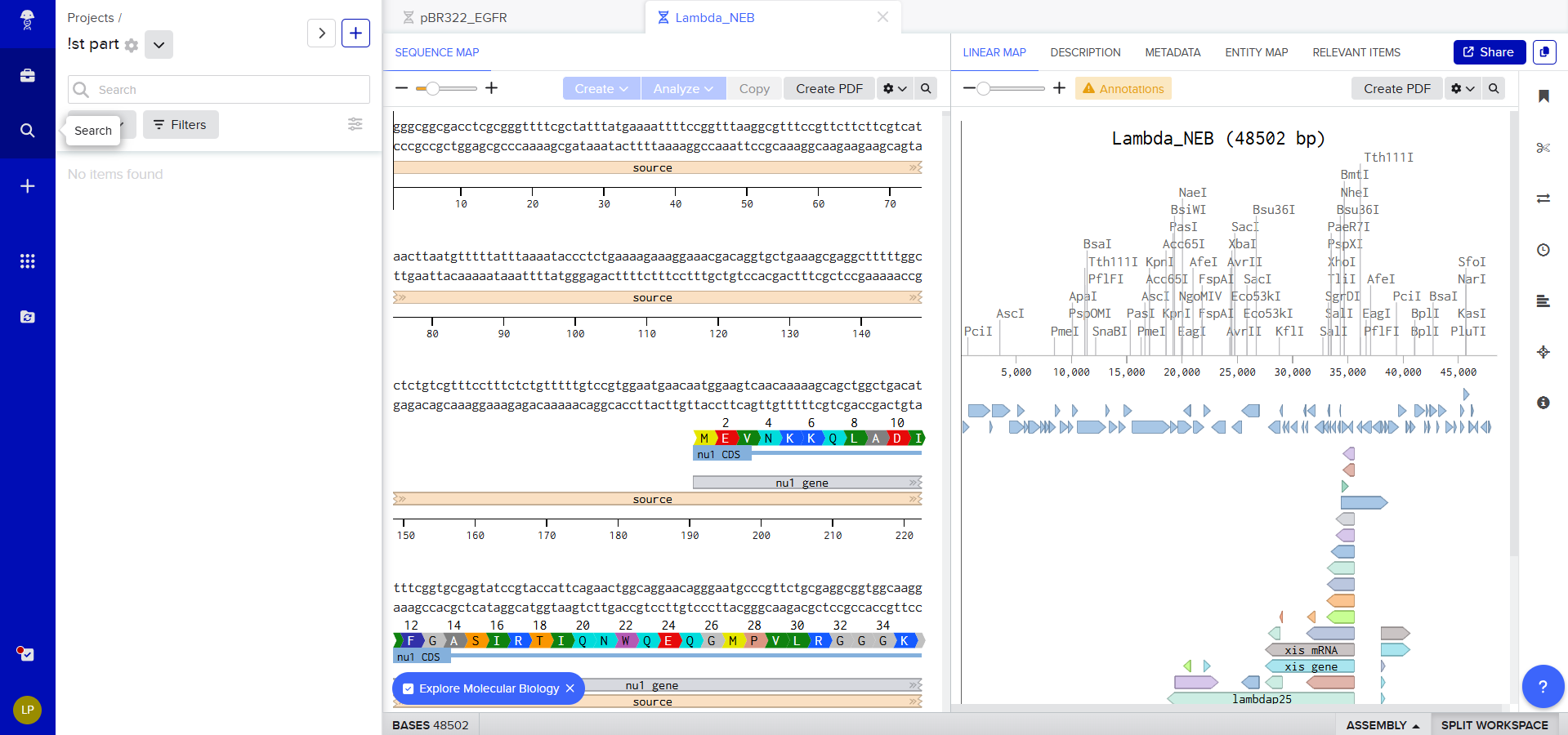

Journey on:
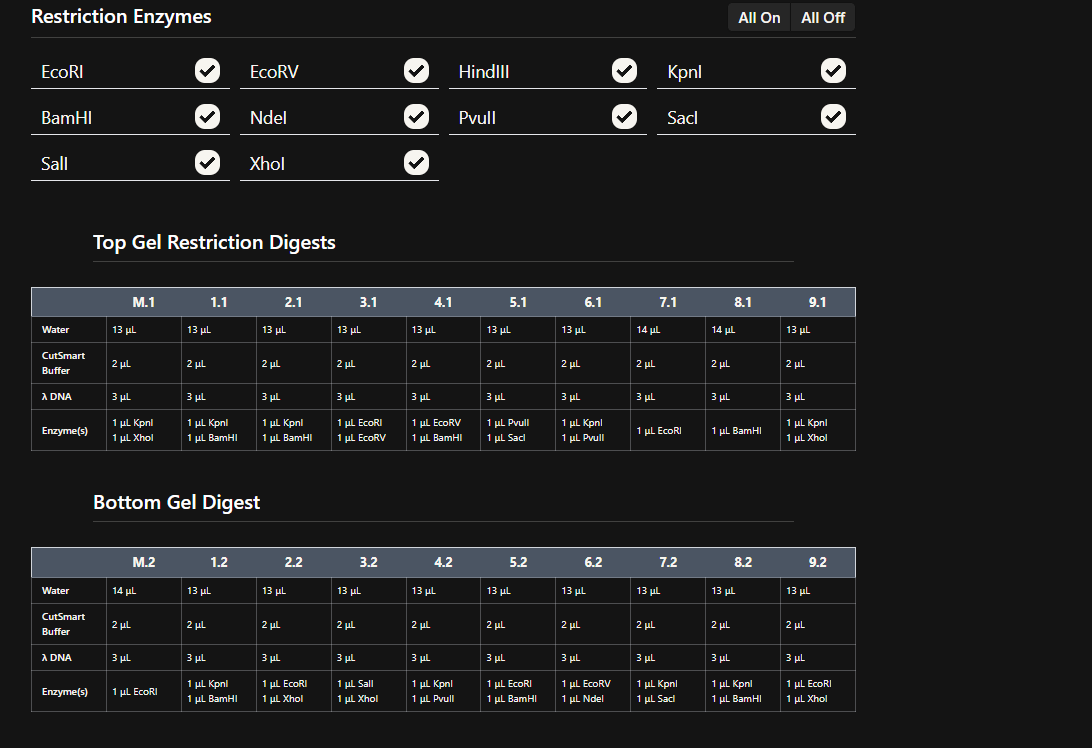
Simulate Restriction Enzyme Digestion with the following Enzymes:
EcoRI
Finding the enzyme is the first step, the description displayed on the left shows improtant information regarding the enzyme and on the top the place where a cut is going to be made is also shown.
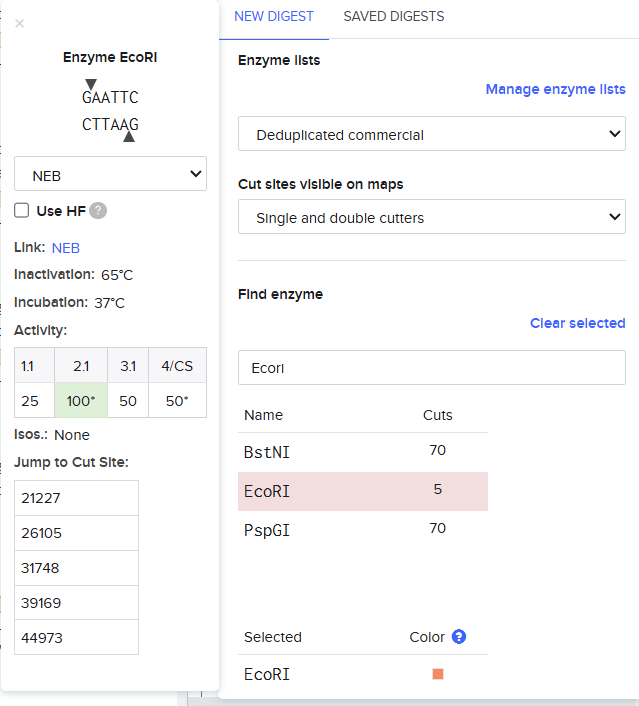
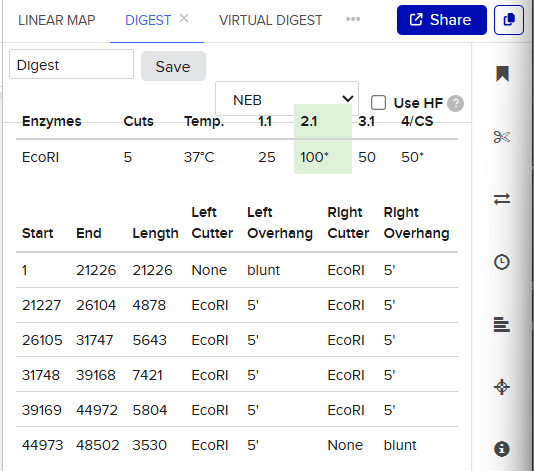
 So when we procede to run the digestion we get Digest and the virtual digest, both simulated ways of the final product view.
So when we procede to run the digestion we get Digest and the virtual digest, both simulated ways of the final product view.
Digest generals

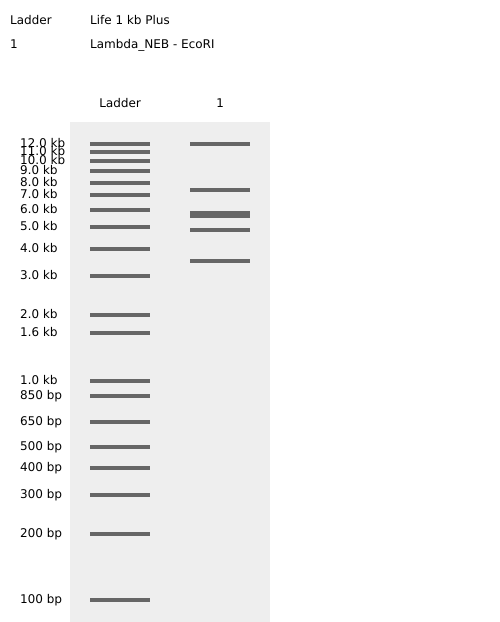
Virtual Digest

HindIII
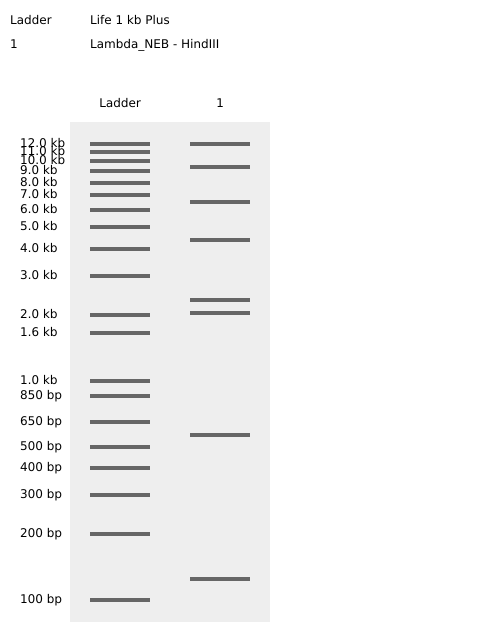

BamHI
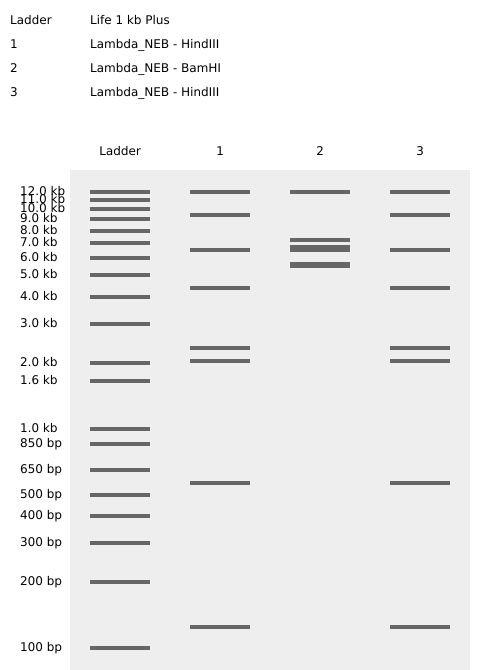


KpnI
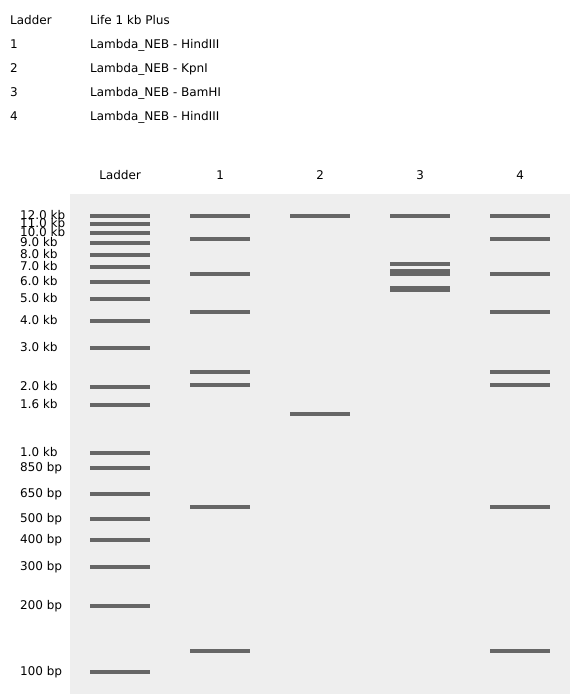

EcoRV
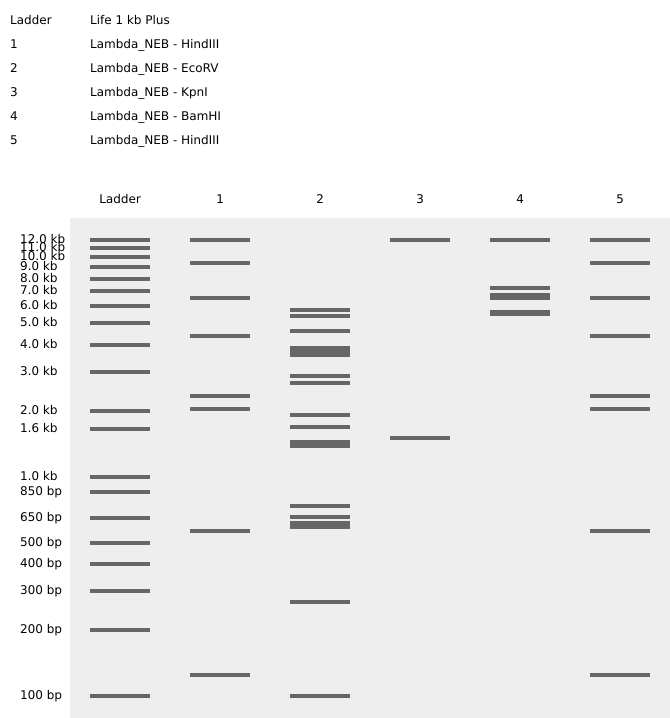


SacI
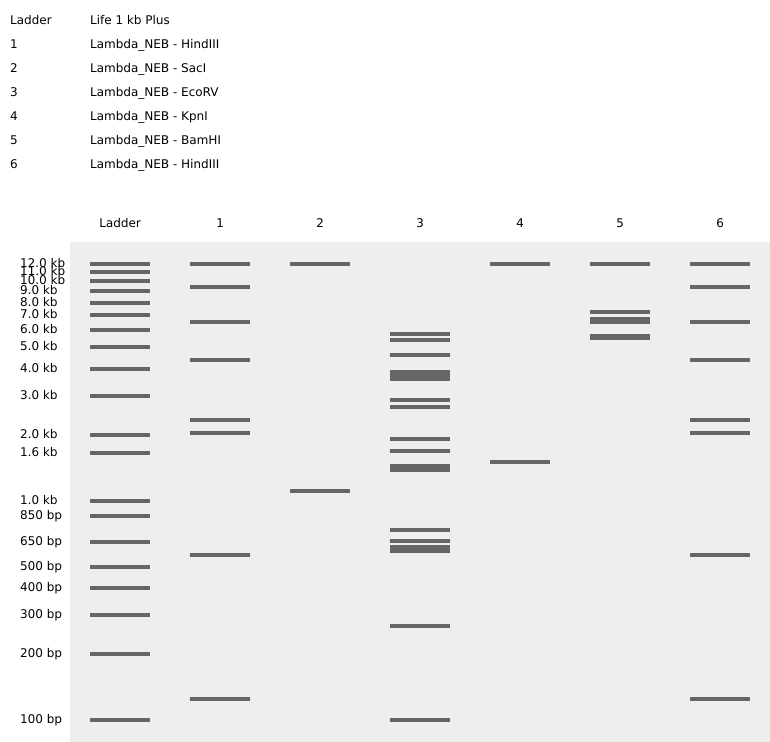


SalI
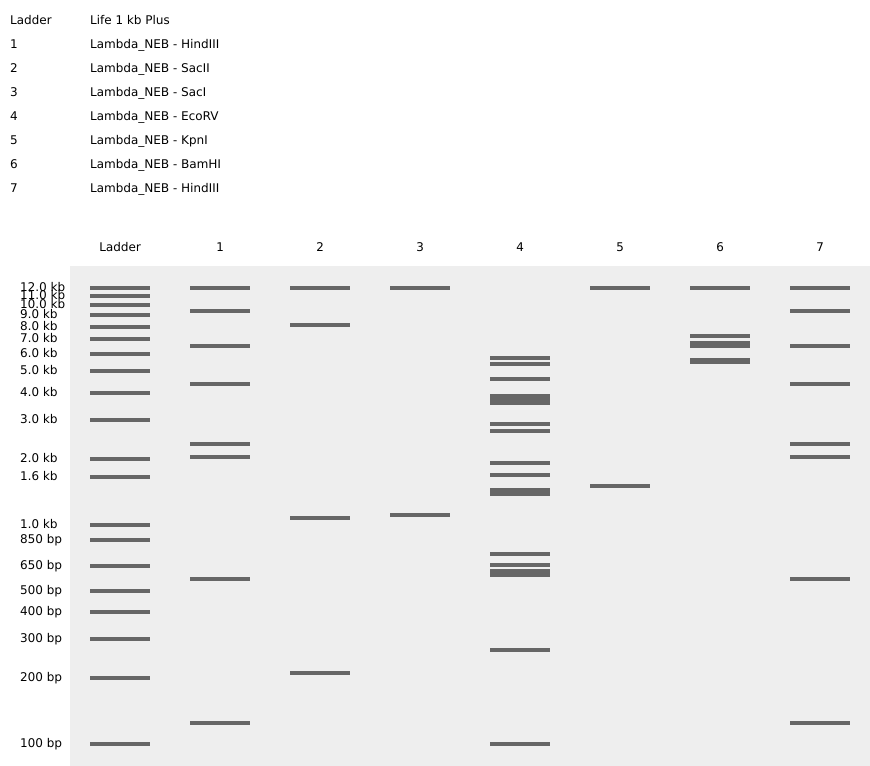


Journey on:
Create a pattern/image in the style of Paul Vanouse’s Latent Figure Protocol artworks. You might find Ronan’s website a helpful tool for quickly iterating on designs!
First attempt was a dog:
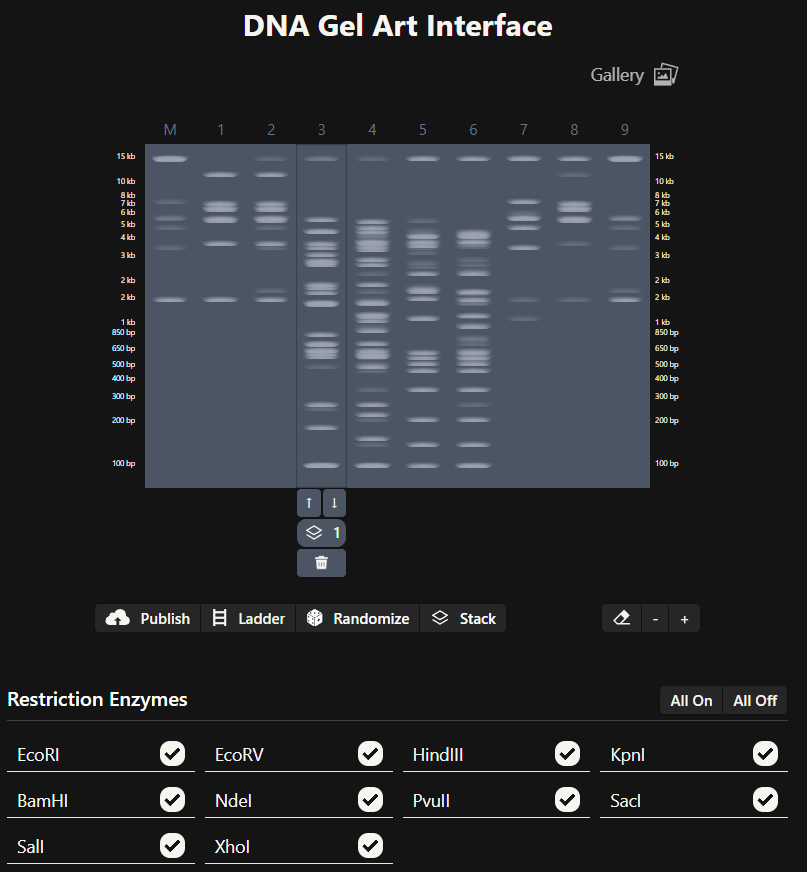

Since i am not doing a lab i have plenty of freedom to choose what enzymes to use on the DNA Gel Art Interface
Part 2: Gel Art - Restriction Digests and Gel Electrophoresis
Lab only
Part 3: DNA Design Challenge
3.1. Choose your protein.
The protein I chose is the Histo-blood group ABO system transferase from Homo sapiens. This protein is responsible for determining human blood types (A, B, AB, or O). It is a glycosyltransferase enzyme that modifies carbohydrate antigens on the surface of red blood cells by adding specific sugar residues. The differences in this enzyme’s activity result in the different ABO blood groups.
The sequence was obtained from UniProt: P16442
The protein:
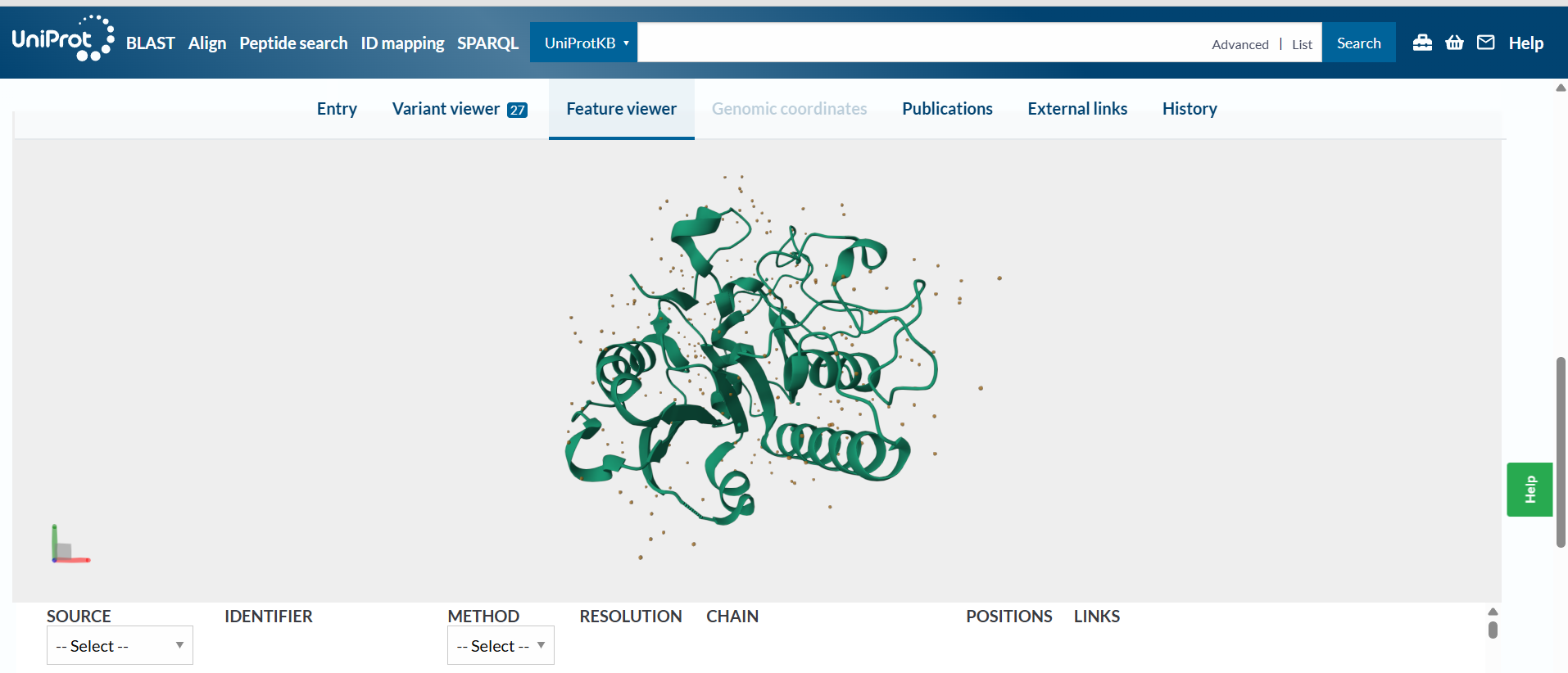
 sp|P16442|BGAT_HUMAN Histo-blood group ABO system transferase OS=Homo sapiens OX=9606
MAEVLRTLAGKPKCHALRPMILFLIMLVLVLFGYGVLSPRSLMPGSLERGFCMAVREPDH
LQRVSLPRMVYPQPKVLTPCRKDVLVVTPWLAPIVWEGTFNIDILNEQFRLQNTTIGLTV
FAIKKYVAFLKLFLETAEKHFMVGHRVHYYVFTDQPAAVPRVTLGTGRQLSVLEVRAYKR
WQDVSMRRMEMISDFCERRFLSEVDYLVCVDVDMEFRDHVGVEILTPLFGTLHPGFYGSS
REAFTYERRPQSQAYIPKDEGDFYYLGGFFGGSVQEVQRLTRACHQAMMVDQANGIEAVW
HDESHLNKYLLRHKPTKVLSPEYLWDQQLLGWPAVLRKLRFTAVPKNHQAVRNP
sp|P16442|BGAT_HUMAN Histo-blood group ABO system transferase OS=Homo sapiens OX=9606
MAEVLRTLAGKPKCHALRPMILFLIMLVLVLFGYGVLSPRSLMPGSLERGFCMAVREPDH
LQRVSLPRMVYPQPKVLTPCRKDVLVVTPWLAPIVWEGTFNIDILNEQFRLQNTTIGLTV
FAIKKYVAFLKLFLETAEKHFMVGHRVHYYVFTDQPAAVPRVTLGTGRQLSVLEVRAYKR
WQDVSMRRMEMISDFCERRFLSEVDYLVCVDVDMEFRDHVGVEILTPLFGTLHPGFYGSS
REAFTYERRPQSQAYIPKDEGDFYYLGGFFGGSVQEVQRLTRACHQAMMVDQANGIEAVW
HDESHLNKYLLRHKPTKVLSPEYLWDQQLLGWPAVLRKLRFTAVPKNHQAVRNP
Length: 354 amino acids Organism: Homo sapiens
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
ATGGCTGAAGTTCTGCGTACTCTGGCTGGCAAGCCCAAGTGCCATGCTCTGCGTCCCATG ATCCTGTTTCTGATCATGCTGGTGCTGGTGCTGTTTGGCTATGGAGTTCTGTCTCCGCGC
This in NCBI
3.3. Codon optimization.
I chose the ABO blood group transferase because I find it biologically interesting and medically relevant. It determines human blood type, which is important for transfusions and healthcare. I also selected it because there is a lot of available information about it, making it a good protein to start practicing sequence analysis and better understand how genes relate to protein function.
ATGGCGGAAGTGCTGCGTACGCTGGCGGGCAAACCGAAATGCCACGCGCTGCGTCCGATG ATCCTGTTTCTGATCATGCTGGTGCTGGTGCTGTTTGGCTATGGCGTGCTGAGCCCGCGC AGCCTGATGCCGGGCTCGCTGGAACGCGGTTTCTGCATGGCGGTGCGCGAGCCGGATCAC CTGCAGCGCGTGAGCCTGCCGCGCATGGTGTACCCGCAGCCGAAAGTGCTGACGCCGTGC CGCAAAGATGTGCTGGTGACGCCGTGGCTGGCGCCGATCGTGGTGGAGGGCACGTTCAAC ATCGACATCCTGAACGAGCAGTTCAGGCTGCAGAACACGACCATCGGCCTGACGGTGTTC GCGATCAAGAAGTACGTGGCGTTCCTGAAGCTGTTTGAGACGGCGGAGAAGCACTTCATG GTGGGCCACCGTGTGCATTATTACGTGTTCACCGACCAGCCGGCGGCGGTGCCGCGCGTG ACGCTGGGCACCGGCCGTCAGCTGTCGGTGCTGGAGGTGCGCGCGTACAAGCGCTGGCAA GACGTGAGCATGCGCCGCATGGAGATGATCAGCGACTTCTGCGAGCGCCGCTTCCTGAGC GAGGTGGACTACCTGGTGTGCGACGTGGACATGGAGTTCCGCGACCACGTGGGCGTG GAGATCCTGACGCCGCTGTTCGGCACGCTGCACCCGGGCTTCTACGGCAGCAGCCGCGAG GCGTTCACGTACGAGCGCCGCCAGAGCCAGGCCTACATCCCGAAAGACGAGGGCGACTTC TACTACCTGGGCGGCTTCTTCGGCGGCAGCGTGCAGGAGGTGCAGCGCCTGACGCGCGCG TGCCACCAGGCGATGATGGTGGACCAGGCCAACGGCATCGAGGCGGTGTGG CACGACGAG AGCCACCTGAACAAGTACCTGCTGCGCCACAAGCCGACGAAGGTGCTGAGCCCGGAGTAC CTGTGGGACCAGCAGCTGCTGGGCTGGCCGGCGGTGCTGCGCAAGCTGCGCTTCACGGCG GTGCCGAAGAACCACCAGGCGGTGCGCAACCCGTAA
3.4. You have a sequence! Now what?
The optimized gene can be chemically synthesized and inserted into a plasmid designed with regulatory elements such as a strong promoter, a ribosome binding site, and a transcription terminator. These components function like biological switches, controlling when and how much protein is produced. The engineered plasmid is then introduced into E. coli, where the synthetic gene is transcribed into mRNA and translated by ribosomes into the ABO transferase protein. Because the sequence was codon-optimized, the bacteria can produce the protein more efficiently.
In synthetic biology, DNA is treated as modular and programmable, allowing us to design regulatory elements and optimize sequences to precisely control protein production and convert a natural human gene into an engineered system for research or biotechnology applications.
Part 4: Prepare a Twist DNA Synthesis Order
Due to my location (Mexico) i was not able to perform this point.

Part 5: DNA Read/Write/Edit
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
I would sequence the human ABO gene from genomic DNA samples. My main interest would be investigating how specific genetic variants in this gene are associated with disease risk, particularly cardiovascular disease, clotting disorders, and susceptibility to certain infections. By sequencing the ABO gene across different individuals, researchers could identify mutations or patterns that correlate with clinical outcomes and better understand how genetic variation contributes to disease mechanisms.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
Also answer the following questions: Is your method first-, second- or third-generation or other? How so? What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps. What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)? What is the output of your chosen sequencing technology?
I would use Illumina sequencing technology because it is highly accurate and well suited for detecting small mutations such as single nucleotide changes. This method is considered second-generation sequencing because it sequences millions of short DNA fragments in parallel rather than reading single long molecules in real time. The input would be extracted human genomic DNA. The DNA would first be fragmented into short pieces, and synthetic adapter sequences would be attached to their ends. These fragments would then be amplified on a flow cell to create clusters. The essential step of the sequencing technology is sequencing-by-synthesis. During this process, fluorescently labeled nucleotides are incorporated one base at a time into a growing DNA strand. Each incorporated base emits a specific signal that is detected by the instrument. The machine converts these signals into a digital sequence through base calling. The output consists of short DNA sequence reads with associated quality scores, which can then be analyzed computationally to identify variants in the ABO gene and examine their relationship to disease.
5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
See some famous examples of DNA design
I would synthesize specific variants of the ABO gene that have been associated with altered disease risk. For example, I would design versions containing mutations suspected to influence clotting or inflammatory pathways. By synthesizing these variants and expressing them in a controlled system, researchers could directly compare how different DNA sequences affect protein function and potentially contribute to disease-related phenotypes.
(ii) What technology or technologies would you use to perform this DNA synthesis and why?
Also answer the following questions: What are the essential steps of your chosen sequencing methods? What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability?
I would use commercial gene synthesis technology, where short DNA fragments are chemically synthesized and assembled into a full-length gene. The essential steps involve synthesizing short oligonucleotides, assembling them into the complete coding sequence, inserting the gene into a plasmid, and verifying the sequence through quality control sequencing. The input is a digitally designed DNA sequence file. The output is a physical DNA construct, typically delivered in a plasmid vector. Limitations include cost for longer or complex sequences, potential synthesis errors that require verification, and turnaround time for manufacturing and validation.
5.3 DNA Edit
(i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
I would edit the ABO gene directly in cultured human cells to introduce specific disease-associated mutations. Editing the gene in its natural genomic context would allow researchers to study how those precise genetic changes influence cellular processes related to clotting, vascular biology, or immune responses. This approach moves beyond correlation and allows functional testing of suspected disease-related variants. But of course this part has its limitations.
(ii) What technology or technologies would you use to perform these DNA edits and why?
Also answer the following questions: How does your technology of choice edit DNA? What are the essential steps? What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing? What are the limitations of your editing methods (if any) in terms of efficiency or precision?
I would probably use CRISPR-Cas9 genome editing technology because it allows precise targeting of specific DNA sequences within the genome.
This system works by designing a guide RNA that matches the target sequence in the ABO gene. The guide RNA directs the Cas9 enzyme to that location, where it creates a double-strand break in the DNA. The cell then repairs the break either by introducing small insertions or deletions or, if a repair template is provided, by incorporating a specific desired mutation.
The preparation involves designing the guide RNA sequence, preparing the Cas9 protein or plasmid, and optionally designing a repair template containing the intended mutation. These components are delivered into cultured cells as the input material.
Limitations include the possibility of off-target edits at similar DNA sequences, variable editing efficiency, and the fact that precise edits using repair templates may not occur in all cells.
References
GenBank overview: https://www.ncbi.nlm.nih.gov/genbank/ NCBI: https://www.ncbi.nlm.nih.gov/genome/ UCSC Genome Browser: https://genome.ucsc.edu/ UniProt: https://www.uniprot.org/uniprotkb/P16442/feature-viewer