Week 1 HW: Principles and Practices

1. First, describe a biological engineering application or tool you want to develop and why.
Effective wastewater treatment is essential for clean waterways, environmental health, and safe drinking water. However, rapid urbanization and population growth have overloaded our capacity to manage aquatic waste, jeopardizing clean water access and biodiversity as pathogens, heavy metals, and algal bloom-inducing nutrients get flushed into waterways. Additionally, current water treatment strategies do not effectively remove a number of harmful compounds, including some drugs and dyes (Renganathan et al., 2025).
Engineered microbial consortia, harnessing and enhancing bacterial communities’ innate capacity to degrade harmful compounds, may offer a promising way to strengthen our current wastewater management approaches and expand the variety of pollutants that we can effectively remove and remediate. I am especially interested in using cyanobacterial species for this purpose—I worked briefly with Nostoc and Oscillatoria species, both of which have promising nitrogen-fixation and biodegradation capabilities (Atoku et al., 2021), over the summer, and I am also studying Acinetobacter baylyi (not a cyanobacterium), which has strong catabolic capabilities and can degrade harmful toxins and aromatic compounds (Baugh et al., 2025; Li et al., 2021). I’m curious about how these species might interact as a microbial community and how we could potentially harness their interactions for better wastewater treatment.
A wastewater remediation project would both expand upon my research as a member of William & Mary’s 2025 iGEM team, which focused on developing design principles to apply SynBio to water-related problems, and is relevant to my local community in Williamsburg: Surrounding cities in Virginia and the broader Chesapeake Bay Watershed suffer from frequent water quality and sewage issues.
2. Next, describe one or more governance/policy goals related to ensuring that this application or tool contributes to an “ethical” future.
Goal 1: Ensure feasibility and effectiveness of deploying engineered microbes in a wastewater context.
- Ensure that engineered microbes function effectively under complex wastewater conditions in addition to laboratory conditions
- Overcome cost and accessibility barriers to deployment
- Promote public trust
Goal 2: Ensure effective containment of engineered microbes and prevent off-target ecological harm.
- Ensure responsible testing and monitoring of engineered solutions
- Develop targeted containment and safety strategies based on experimental results
3. Next, describe at least three different potential governance “actions” by considering the four aspects below (Purpose, Design, Assumptions, Risks of Failure & “Success”).
Proposed governance actions:
- Assess performance and safety of engineered solutions in simulated wastewater environments prior to deployment; adapt engineering approach based on findings.
- Facilitate discussion (e.g., scheduled meetings and/or townhalls) between scientists, wastewater plants, and the public about current needs and potential solutions.
- Require that scientists and wastewater treatment plants develop a detailed risk-mitigation and containment plan prior to deployment, outlining procedures for using engineered microbes in a water treatment context and strategies to maintain and monitor bacterial containment.
- Require approval from relevant local governmental health and environmental agencies (e.g., the Virginia Department of Health and Virginia Department of Environmental Quality) prior to deployment.
4. Next, score (from 1-3 with, 1 as the best, or n/a) each of your governance actions against your rubric of policy goals.
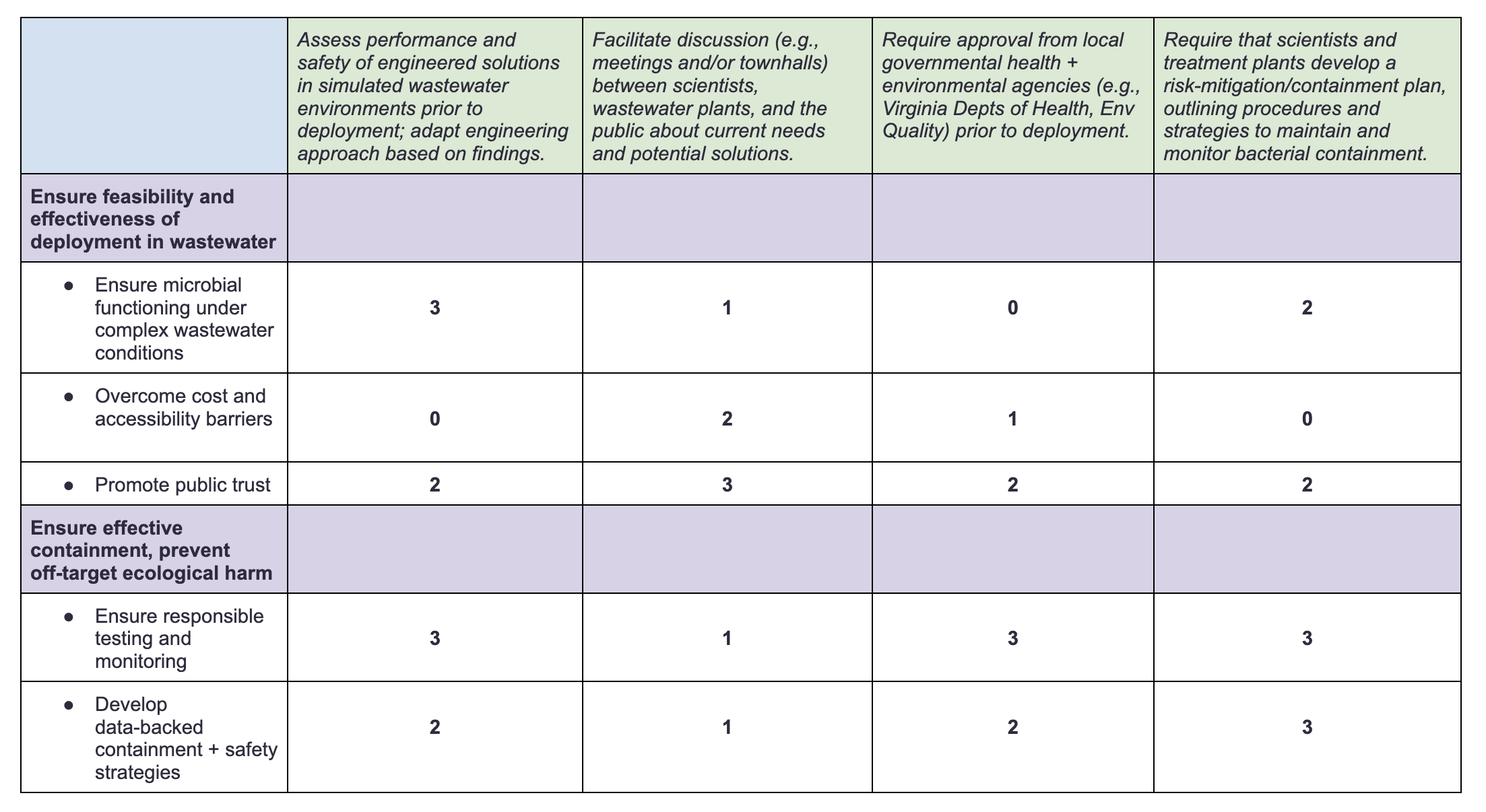
5. Last, drawing upon this scoring, describe which governance option, or combination of options, you would prioritize, and why. Outline any trade-offs you considered as well as assumptions and uncertainties.
Based on the above rubric, I would prioritize a governance framework that integrates rigorous scientific testing and safety assessment with discussion between stakeholders, developing regulations that take into account both scientific data and the perspectives of regulators and the public. This framework would create a network of accountability between groups—e.g., ensuring that regulatory decisions are scientifically backed but also that deployment of scientific solutions doesn’t outpace the development of mechanisms to ensure their safety and sustainability. The system involves a tradeoff between testing/safety and the speed of development, but is necessary to ensure that solutions are effective and that we do not deploy them in haste.
References
Atoku, D. I., Ojekunle, O. Z., Taiwo, A. M., & Shittu, O. B. (2021). Evaluating the efficiency of Nostoc commune, Oscillatoria Limosa and chlorella vulgaris in a phycoremediation of heavy metals contaminated industrial wastewater. Scientific African, 12. https://doi.org/10.1016/j.sciaf.2021.e00817
Baugh, A. C., Tumen-Velasquez, M. P., Zempel, I. R., Duscent-Maitland, C. V., Slarks, L. E., Defalco, J. B., Johnson, C. W., Beckham, G. T., & Neidle, E. L. (2025). Rewiring aromatic compound consumption: Chromosomal amplification and evolution of a foreign pathway in acinetobacter baylyi ADP1. ACS Synthetic Biology, 14(9), 3543–3556. https://doi.org/10.1021/acssynbio.5c00341
Li, H., Yang, Y., Zhang, D., Li, Y., Zhang, H., Luo, J., & Jones, K. C. (2021). Evaluating the simulated toxicities of metal mixtures and hydrocarbons using the alkane degrading bioreporter Acinetobacter Baylyi adpwh_reca. Journal of Hazardous Materials, 419, 126471. https://doi.org/10.1016/j.jhazmat.2021.126471
Renganathan, P., Gaysina, L. A., García Gutiérrez, C., Rueda Puente, E. O., & Sainz-Hernández, J. C. (2025). Harnessing engineered Microbial Consortia for xenobiotic bioremediation: Integrating multi-omics and AI for next-generation wastewater treatment. Journal of Xenobiotics, 15(4), 133. https://doi.org/10.3390/jox15040133
—————————————————————————————————————————–
Week 2 Lecture Prep
Homework Questions from Professor Jacobson:
1. Nature’s machinery for copying DNA is called polymerase. What is the error rate of polymerase? How does this compare to the length of the human genome. How does biology deal with that discrepancy?
DNA polymerase’s error rate is 1:10^6 bp. The human genome is 3 billion bp, so cells have evolved DNA repair mechanisms to correct errors when they occur. For example, the protein MutS identifies incorrect DNA base pairings and starts the repair processes alongside other proteins.
2. How many different ways are there to code (DNA nucleotide code) for an average human protein? In practice what are some of the reasons that all of these different codes don’t work to code for the protein of interest?
There are 20 amino acids and, because of codon redundancy, about two to four different ways to encode each of them. If the average protein is about 300 amino acids long, this would mean about 3^300 different possible ways to encode the protein. In reality, these codes don’t all work because the specific codons are translated at different rates—and translation speed may affect protein folding.
Homework Questions from Dr. LeProust:
1. What’s the most commonly used method for oligo synthesis currently?
The Phosphoramidite DNA Synthesis method
2. Why is it difficult to make oligos longer than 200nt via direct synthesis?
It takes a long time and the yield is low due to the long coupling time and capping process.
3. Why can’t you make a 2000bp gene via direct oligo synthesis?
The yield and time constraints make this impossible.
Homework Question from George Church:
1. What are the 10 essential amino acids in all animals and how does this affect your view of the “Lysine Contingency”?
The 10 essential amino acids are arginine, histidine, isoleucine, leucine, lysine, methionine, phenylalanine, threonine, tryptophan, and valine. Animals must obtain the “essential” amino acids through their diet. The “Lysine Contingency” does not make sense because it involves a scientist removing dinosaurs’ ability to produce lysine, which they could not produce in the first place.