Week 6 HW: Genetic Circuits Part I: Assembly Technologies
A. DNA Assembly
1. What are some components in the Phusion High-Fidelity PCR Master Mix and what is their purpose?
The Phusion High-Fidelity PCR Master Mix contains a DNA polymerase enzyme that is a modified version of one found in Pyrococcus furiosus – to create the second strand during the extension step; deoxynucleotides so that the polymerase has nucleotides to use for extension; and reaction buffer so that the pH and other conditions are optimal for PCR.
2. What are some factors that determine primer annealing temperature during PCR?
GC-content, primer length, and primer concentration all affect annealing temperature.
3. There are two methods from this class that create linear fragments of DNA: PCR, and restriction enzyme digests. Compare and contrast these two methods, both in terms of protocol as well as when one may be preferable to use over the other.
Restriction enzyme digests cut DNA at specific cut site sequences using enzymes of your choice. You obtain whatever fragments result from cutting the existing DNA at that site, and there is no amplification process. For PCR, you design oligo primers for a longer sequence and then selectively amplify it through repeated denaturation, annealing, and extension. PCR is useful for quickly determining whether a specific sequence is present in your sample, and for extending DNA fragments in such a way that they overlap and can be assembled via downstream reactions (e.g. creating sticky ends with exonucleases in Gibson Assembly). Restriction digests allow you to cut DNA at sites of interest so that you can stick fragments together in the way that you want.
4. How can you ensure that the DNA sequences that you have digested and PCR-ed will be appropriate for Gibson cloning?
Prior to finalizing your selection of target sequences and primer design, you should use online calculators to ensure that relevant sequences are compatible with each other and include all sequences necessary (e.g., promoter, RBS, etc) for the expression of the gene(s) of interest, have a GC content that isn’t too high or too low, and that the insert-vector molar ratio will be 2:1.
5. How does the plasmid DNA enter the E. coli cells during transformation?
By diffusing through pores (generated through the electrical shock) in the cell membrane.
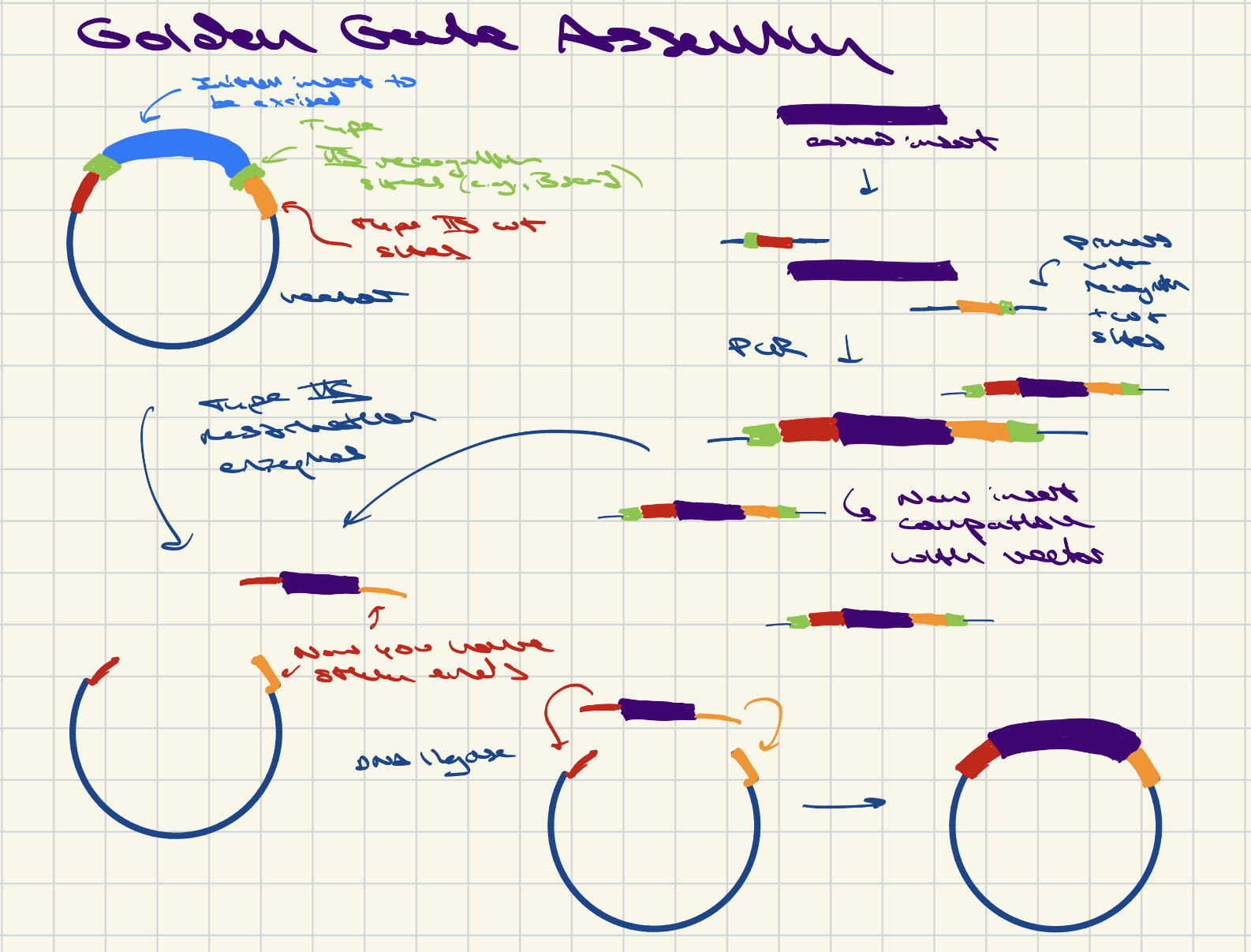
6. Describe another assembly method in detail (such as Golden Gate Assembly) Explain the other method in 5 - 7 sentences plus diagrams (either handmade or online). Model this assembly method with Benchling or Asimov Kernel!
Golden Gate Assembly uses TypeIIS recognition enzymes, which cleave DNA outside their recognition site, to create sticky overhangs in a destination vector/plasmid and corresponding fragment. Usually, the original plasmid contains an easily-screenable sequence so that you can determine whether you’ve effectively removed it and replaced it with your fragment of interest. To add your insert, you design PCR primers that target the sequence of interest and that also extend it with TypeIIS recognition sequences and cut sites that you’ve designed to match those in the target vector. After PCR, the resulting amplicon should be compatible with the vector – you can use restriction enzymes to create sticky ends and ligate the insert and vector in a “one pot” reaction involving the fragments, vector, restriction enzyme, and DNA ligase. At the end of the reaction, you add more TypeIIS enzyme to ensure that any remaining uncleaved vectors are removed, and you inactivate the enzymes by increasing the reaction temperature.
B. Asimov Kernel
Create a blank Notebook entry to document the homework and save it to that Repository. Explore the devices in the Bacterial Demos Repo to understand how the parts work together. Recreate the Repressilator in that empty Construct by using parts from the Characterized Bacterial Parts repository.
Circuit schematic:
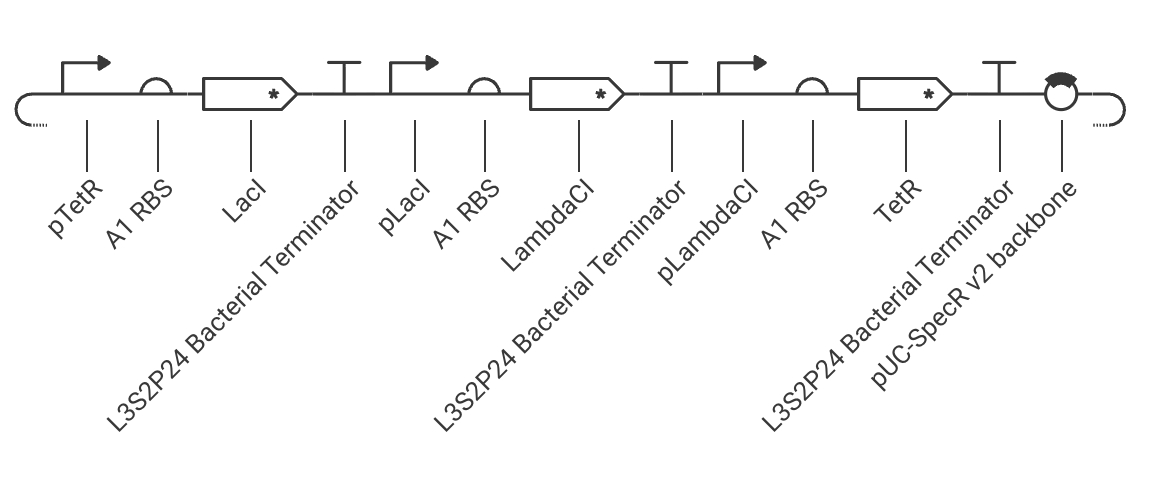 Oscillating protein expression after addition of aTc inducer:
Oscillating protein expression after addition of aTc inducer:
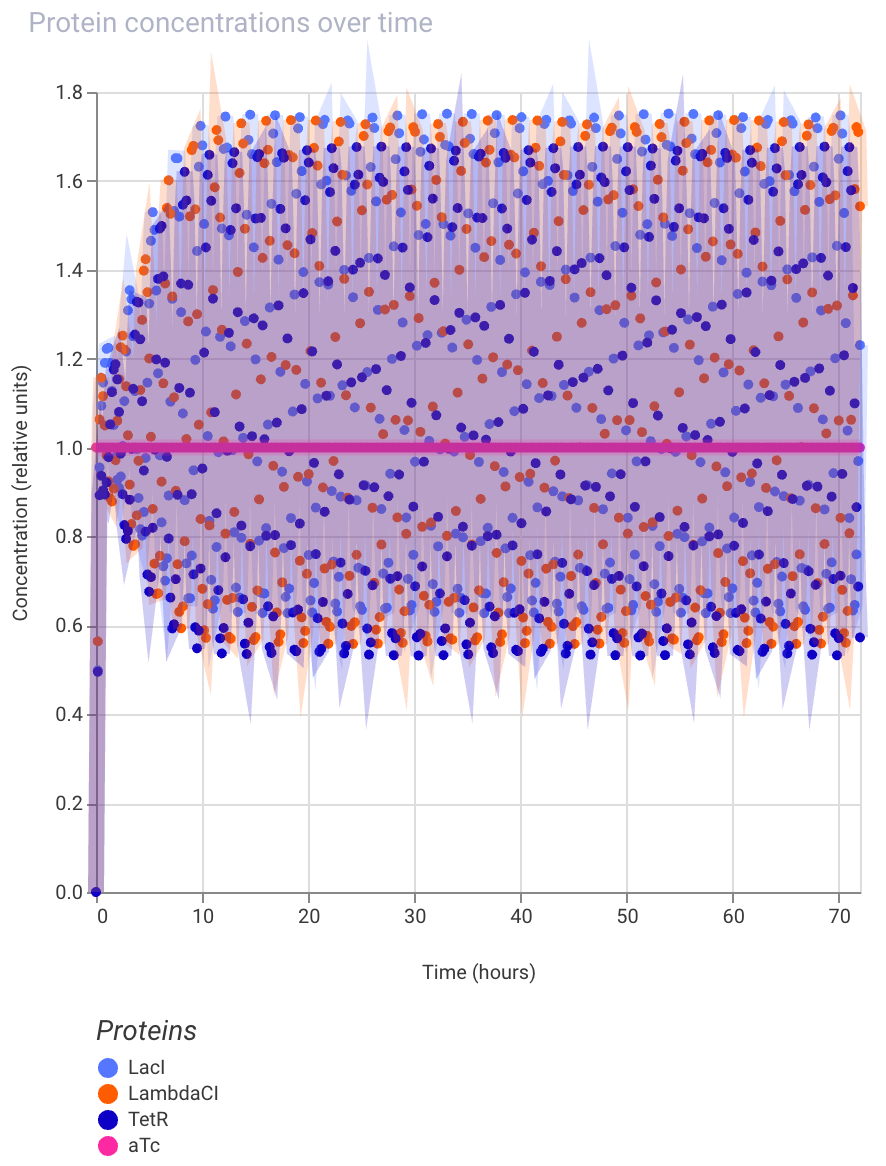
Build three of your own Constructs using the parts in the Characterized Bacterials Parts Repo. Explain in the Notebook Entry how you think each of the Constructs should function. Run the simulator and share your results in the Notebook Entry. If the results don’t match your expectations, speculate on why and see if you can adjust the simulator settings to get the expected outcome.
Practice Construct 1: System expressing Ag43 (biofilm formation) in E. coli. Ag43 can be repressed by the addition of an inducer, which triggers LacI expression tied to Ag43’s repressible promoter.
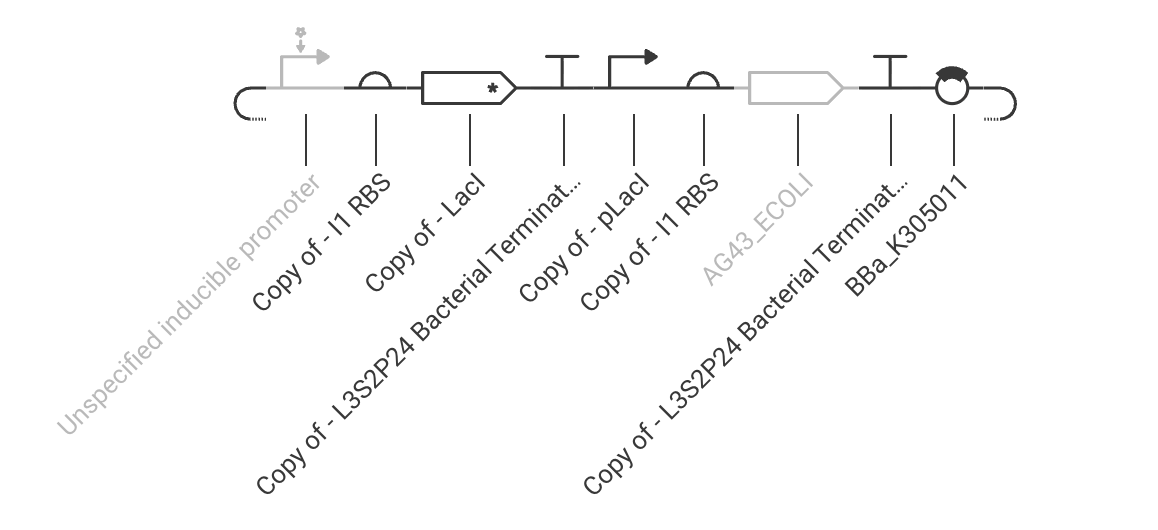
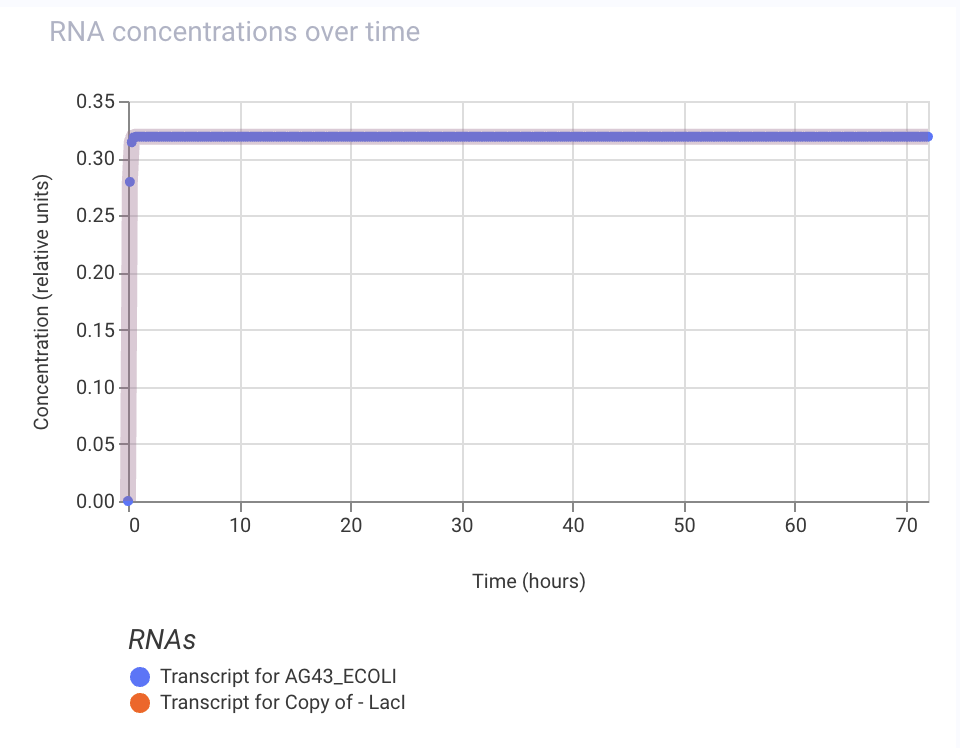
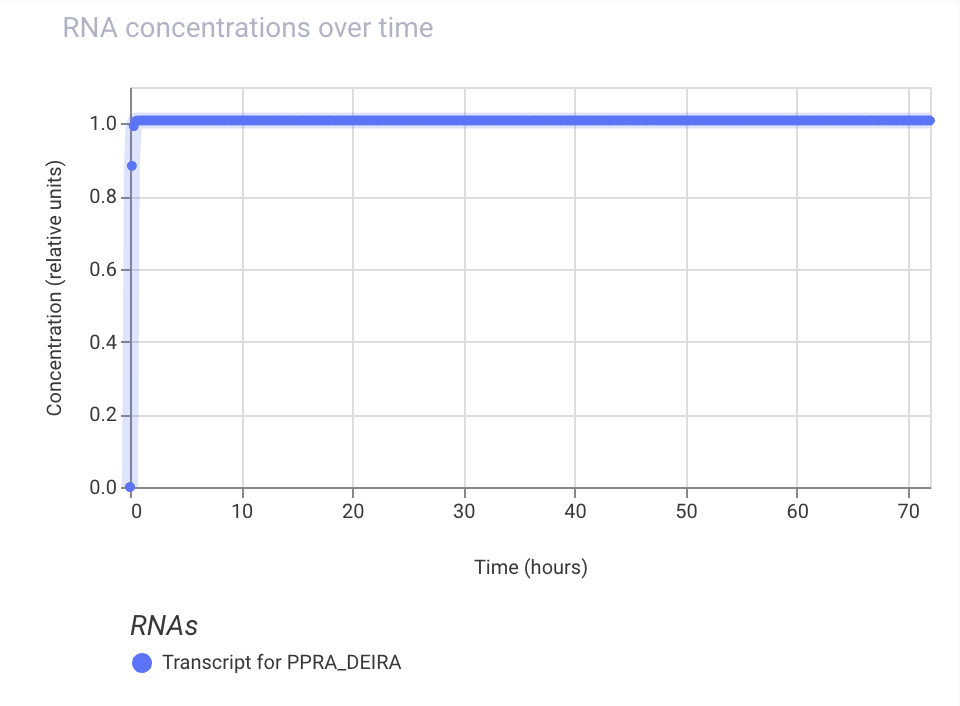
Practice Construct 2: Simple constitutive expression system for Ppra, a radiation resistance protein found in D. radiodurans
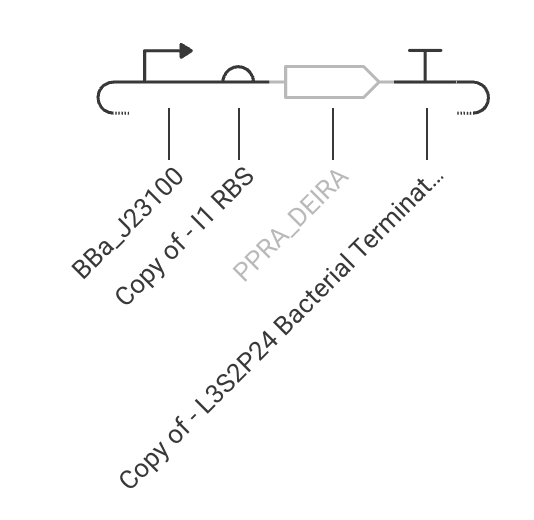
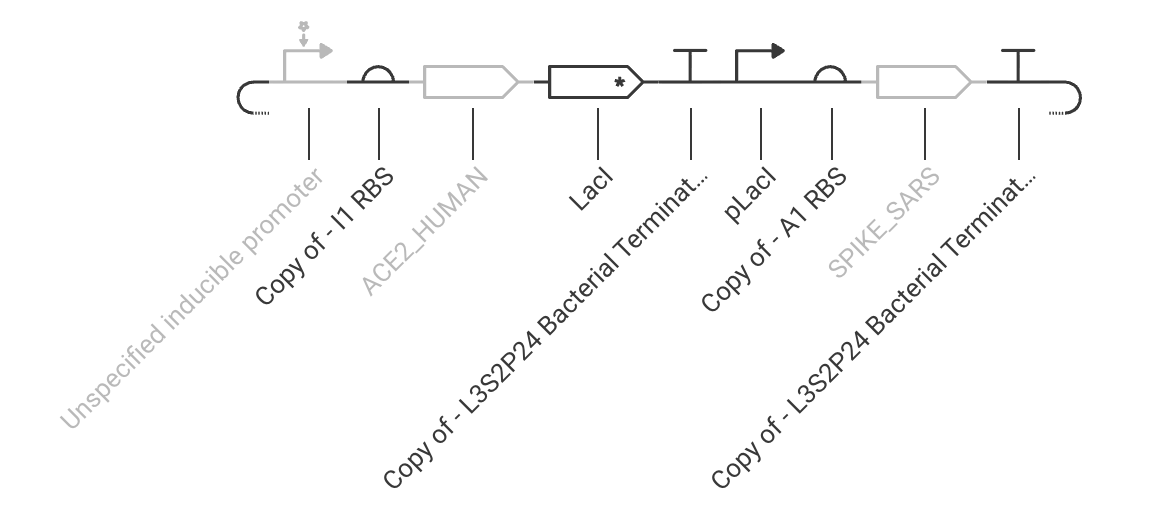
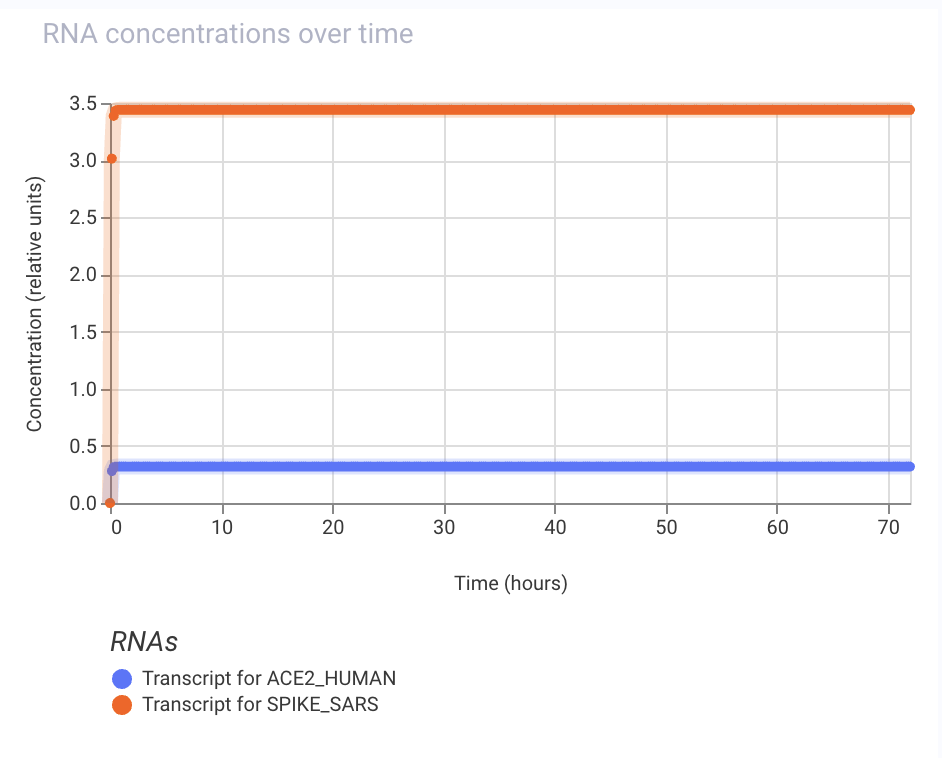
Practice Construct 3: System that expresses either the Covid spike protein or a human ACE2 receptor protein. ACE2 is expressed in response to inducer addition and is linked to the expression of LacI, which represses the spike protein in this circuit.