Week 7 HW: Genetic Circuits Part II: Neuromorphic Circuits
Part 1: IANNs
1. What advantages do IANNs have over traditional genetic circuits, whose input/output behaviors are Boolean functions?
IANNs are more complex and flexible than traditional genetic circuits and can involve a greater degree of abstraction (i.e., the inputs aren’t digital). Specifically, IANNs allow non-digital dosages for better spatial and temporal control of gene expression in ways that more effectively address the complex and systems-based natural of relevant deployment settings such as the human immune system.
2. Describe a useful application for an IANN; include a detailed description of input/output behavior, as well as any limitations an IANN might face to achieve your goal.
IANNs provide a non-digital behavior that could potentially be useful for developing sensitive engineered systems that control microbial behavior (the outputs) at the population or community level in response to specific combinations external/environmental factors (inputs) at given levels. For example, certain specific combinations of quroum sensing molecules at a given level are known to cause certain behavioral responses, some of which are relevant to human health and the environment – e.g., toxin production and pathogenicity-associated responses. Using IANNs, we could create systems that detect (and respond to) those combinations with high sensitivity and specificity, with applications in biosensing (e.g., producing reporters that indicate the dangerous conditions) and developing methods to target and reduce pathogenicity.
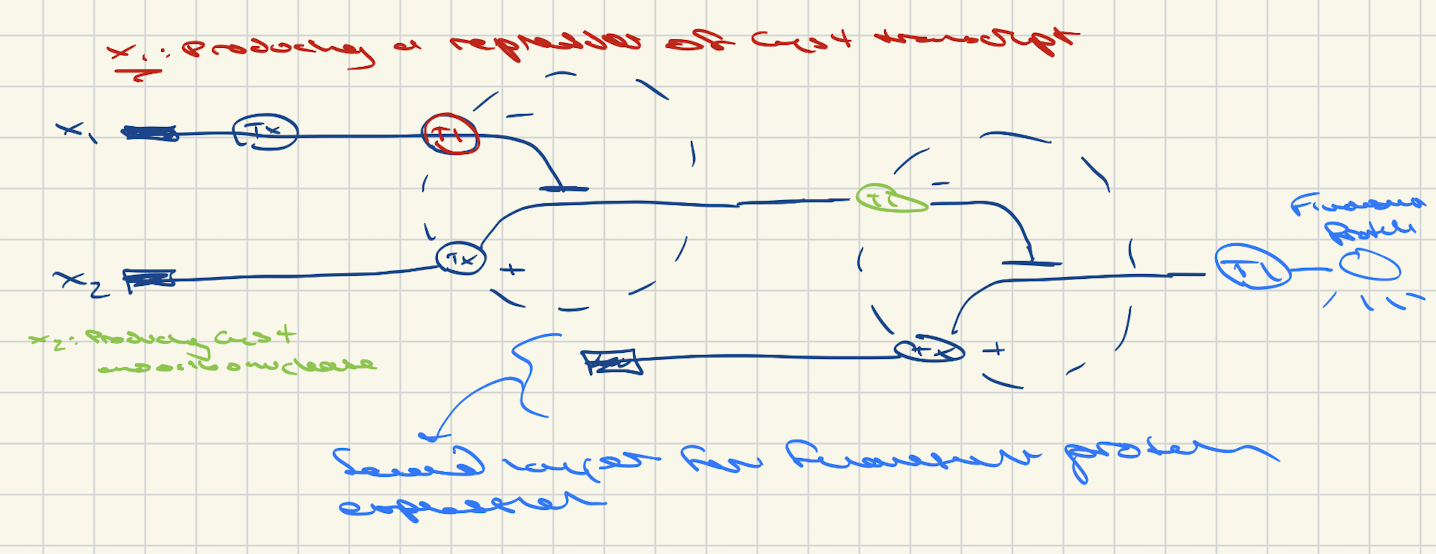
3. Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
Diagram below: The first layer regulates endoribonuclease production via a similar repression mechanism to that of the endoribonuclease – acting at the post-transcriptional level on the endoribonuclease transcript. When repressor acting upon the endoribonuclease is low enough, the endoribonuclease is expressed and can turn off the fluorescent protein’s expression (in the second layer) by binding to its transcripts before they can be translated. Enough expression of the repressor of the endoibonuclease results in the expression of the fluorescent protein.
Part 2: Fungal Materials
1. What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Fungal materials are usually mycelium-based, meaning they use the root part of the fungus. Mycelium-based materials have been proposed as a more sustainable and biodegradable alternative to traditional packaging components, building materials, and textiles. However, they tend to have less structural integrity than traditional alternatives, and we currently do not have large-scale mycelium material growth/production systems in place for manufacturers to use.
2. What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
Fungi could be engineered to self-assemble into useful biomaterials via modification of signaling systems and pathways invovled in poulation pattern formation. These systems could be engineered to respond to specific, experimentor-deterimed inducers, which manufacturors could use for fine-tuned control of fungal manufacturing at a larger scale.
Fungi are eukaryotes, so are more complex and likely can be engineered to produce a wider variety of compunds and perform a wider variety of functions than we can engineer into bacteria. Certain fungi, such as the budding yeast Saccharomyces cerevisiae are good model organsisms for studying eukaryotic cell behavior and are good platforms for using SynBio to conduct basic science research (e.g., understanding the effects of genetic variation and protein-protein interactions of interest by measuring and observing the effects of specific engineering changes, potentially involving complex circuits).
Part 3: First DNA Twist Order
Project Title: Developing an Engineerable Photogranule System for Wastewater Treatment with Cyanobacterium Oscillatoria sp. and Chassis Bacterium Acinetobacter baylyi ADP1
Brief Project Description: Oxygenic photogranules, glob-like consortia of filamentous photosynthetic cyanobacteria and non-photosynthetic bacteria, naturally absorb and break down harmful chemicals and offer several sustainability advantages over current wastewater treatment techniques. My long-term goal is to develop an engineerable model photogranule system, using filamentous cyanobacterium Oscillatoria and the heterotrophic chassis Acinetobacter baylyi,that could be easily modified to enhance versatility and resilience to diverse microbial conditions, target specific chemical products and pollutants of-interest for removal and/or detoxification, and enhance the viability of the bacterial-waste sludge for renewable downstream applications.
Draft Aim 1: My first immediate experimental aim is to enhance A. baylyi’s biofilm formation and/or extracellular polysaccharide production in such a way that it associates more effectively and reliably with Oscillatoria. Currently, I am considering engineering A baylyi to express Ag43, a well-characterized cell membrane protein that, when upregulated/expressed, promotes cellular aggregation and biofilm formation in E. coli. By forming aggregates with itself, I hypothesize that the engineered A. baylyi will be better suited (than the wild type) to form photogranules with Oscillatoria in a wastewater setting. My immediate experimental steps are to 1) design/order a plasmid that expresses Ag43 for use in A. baylyi or order genetic parts that I could assemble via Gibson Assembly, 2) assess wild type A. baylyi’s level of natural association with Oscillatoria in the lab, and 3) transform the Ag43 plasmid into A. baylyi (after any assembly steps, if necessary) and assess its effect on A. baylyi biofilm formation and association with Oscillatoria via visual observation and microscopy.