Week 7: HW genetic circuits part II
This week covers neuromorphic genetic circuits, showing how engineered gene networks can implement neural-network “perceptron”-like computation and learning.
Objective:
- Understand intracellular artificial neural networks (IANNs).
- Explore applications of neuromorphic genetic circuits.
- Learn about fungal materials and possible engineering applications in fungi.
- Make progress on the individual final project and first DNA design order.
Assignment Part 1: Intracellular Artificial Neural Networks (IANNs)
Assignees for the following sections
| MIT/Harvard students | Required |
| Committed Listeners | Required |
1. What advantages do IANNs have over traditional genetic circuits, whose input/output behaviours are Boolean functions?
Intracellular artificial neural networks (IANNs) are advantageous to Boolean genetic circuits because they can exhibit graded biological signals rather than absolute on-or-off states. In reality, cells do not have simple on and off outputs, making IANNs approximate the functionality of the cells with better accuracy. Many inputs can affect a single output and vary like a spectrum; biomolecules like miRNA, TFs, and plasmids drastically alter cellular states. As Anthony Genot explains, “Enzymes bring high nonlinearity, which we exploit to improve the sharpness of computation.” This nonlinearity is difficult to model via Boolean functions using logic gates. IANNs can also integrate many inputs with varying weights to create a more accurate representation of the biological system. This way, inhibitory and stimulatory signals are weighted rather than treated as binary. Compared with Boolean circuits, which become disproportionately complex with growing inputs, multilayer networks can approximate graded, weighted, and non-linear signals of a cell.
2. Describe a useful application for an IANN; include a detailed description of input/output behaviour, as well as any limitations an IANN might face to achieve your goal.
One potential application for IANNs could be to optimise biosynthesis pathways by controlling metabolic outputs of the microbial strain. Since IANNs allow for specific control of the cells, they could be used to mimic natural enzymatic kinetics to specify and optimise metabolic products. The IANN could help by sensing several continuous intracellular signals at the same time, such as precursor availability, intermediate accumulation, stress level, and growth state, and then using those signals to regulate key pathway enzymes in a graded way instead of a simple ON/OFF manner. For example, if harmful intermediates start to build, the system could reduce the specific pathway tied to it to relieve cellular stress and prevent cell death. Although these systems are useful, they can come with many costs. Presently, IANNs suffer from noisy intracellular signals, cell-to-cell variability, and changing cellular context, which makes it difficult to get precise and reproducible input/output behaviour. They can also suffer from cross-talk and resource competition, where different regulators interfere with each other or compete for limited cellular machinery. In biosynthesis, this is even harder because pathways need very precise balancing, and multilayer circuits respond slowly since each layer depends on transcription and translation.
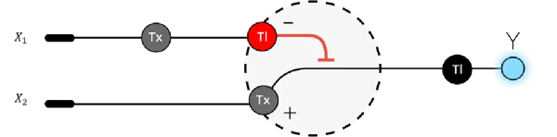
3. Below is a diagram depicting an intracellular single-layer perceptron where the X1 input is DNA encoding for the Csy4 endoribonuclease and the X2 input is DNA encoding for a fluorescent protein output whose mRNA is regulated by Csy4. Tx: transcription; Tl: translation.
Draw a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
I created a diagram for an intracellular multilayer perceptron where layer 1 outputs an endoribonuclease that regulates a fluorescent protein output in layer 2.
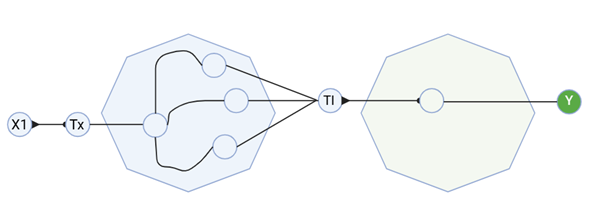
Assignment Part 2: Fungal Materials
Assignees for the following sections
| MIT/Harvard students | Required |
| Committed Listeners | Required |
1. What are some examples of existing fungal materials and what are they used for? What are their advantages and disadvantages over traditional counterparts?
Existing fungal materials are mostly mycelium-based materials, including packaging foams, insulation boards/panels, construction composites, and leather-like textiles made from fungal biomass grown on agricultural waste. These materials are used as substitutes for polystyrene packaging, synthetic foams, animal leather, and some lightweight building materials, because mycelium can act as a natural binder and be shaped into different forms during growth. The main advantage is that they are biobased, biodegradable, and can be produced from waste streams, which can lower environmental impact compared with plastics or animal-derived materials. Their disadvantages are that they still often have lower and more variable mechanical performance, limited water resistance and durability, and struggle to adapt to diverse environments.
2. What might you want to genetically engineer fungi to do and why? What are the advantages of doing synthetic biology in fungi as opposed to bacteria?
If I were to genetically engineer fungi, I would want to make them better microbial factories for complex pharmaceuticals such as paclitaxel. In that case, the goal would be to engineer fungi to express more of the paclitaxel pathway efficiently and improve precursor supply. Synthetic biology in fungi can be advantageous over bacteria for this type of problem because fungi are eukaryotes, so they are generally better at handling complex eukaryotic biosynthetic enzymes, performing post-translational modifications, secreting proteins, and supporting secondary metabolite pathways that are often harder to reconstruct in bacterial hosts. This makes fungi, especially yeasts and filamentous fungi, attractive hosts for producing plant- or fungus-derived pharmaceuticals, since many of these molecules depend on enzyme systems and intracellular organisation that are more similar to those of other eukaryotes than to bacteria. Fungi are also already widely used as industrial production organisms, which makes them a promising chassis for scaling the biosynthesis of valuable drugs. However, bacteria tend to grow faster and are usually easier to manipulate genetically, making it easier to adapt to a variety of situations and contexts.
Assignment Part 3: First DNA Twist Order
Assignees for the following sections
| MIT/Harvard students | Required |
| Committed Listeners | Required |
0. Review the Individual Final Project documentation guidelines.
I reviewed the Individual Final Project documentation guidelines.
1. Submit this Google Form with your draft Aim 1, final project summary, HTGAA industry council selections, and shared folder for DNA designs.
N/A, waiting on feedback for the final project, but I have made an explanation of my preject for Paxlitaxel on my project page
2. Review Part 3: DNA Design Challenge of the week 2 homework. Design at least 1 insert sequence and place it into the Benchling/Kernel/Other folder you shared in the Google Form above. Document the backbone vector it will be synthesized in on your website.
N/A, waiting on feedback for the final project.