week-02-hw-dna-read-write-and-edit
Part 1
For this part of the assignment I started playing with Ronan´s website by making random patterns with the enzymes given. Som of the electrophoresis digestion patterns of the DNA started to look like letters and I decided to make my own word.
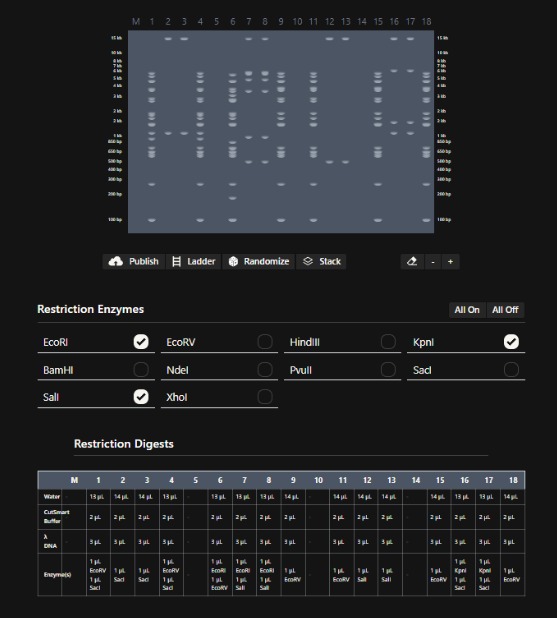

Then, i tried to design that same pattern in benchling digestion tool, but i think it look better in Ronan´s website, because there are sapces between the gel lanes.
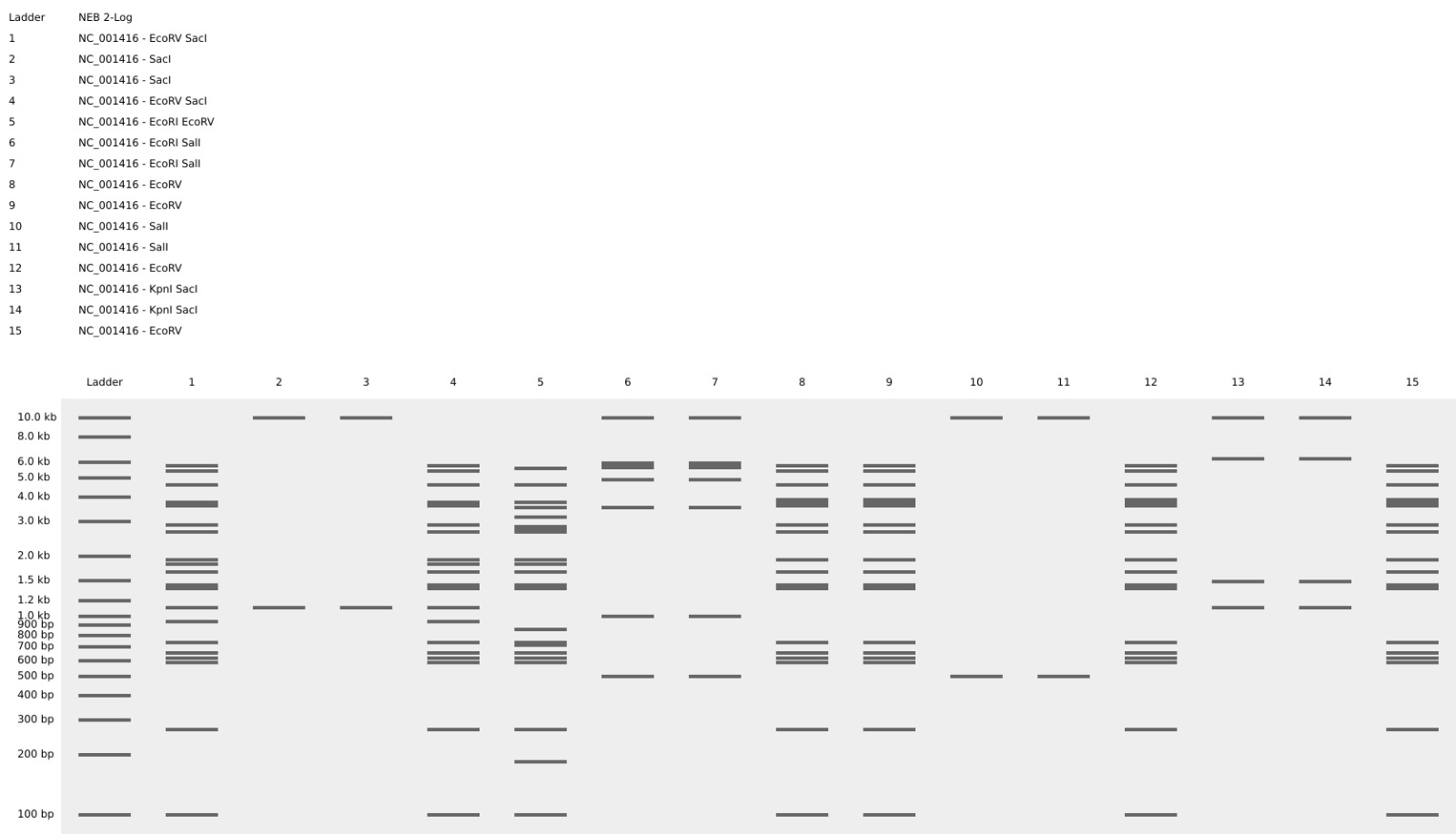

Part 3
For this part I chosed a protein that I am very familiarized with. It is the transcriptional repressor PhnF from the phn operon responsible for the phosphonate degradation in bacteria. It repress the transcription of the phng gen in the absence of phosphonates, a core catalitic enzyme of the C-P lyase complex. This promoter is also regulted by the PhoB activator, so in the presence of phosphonates and the deplition of inorganic phosphorous this degradative path activates and provides the bacteria of an alternate source of phosphate
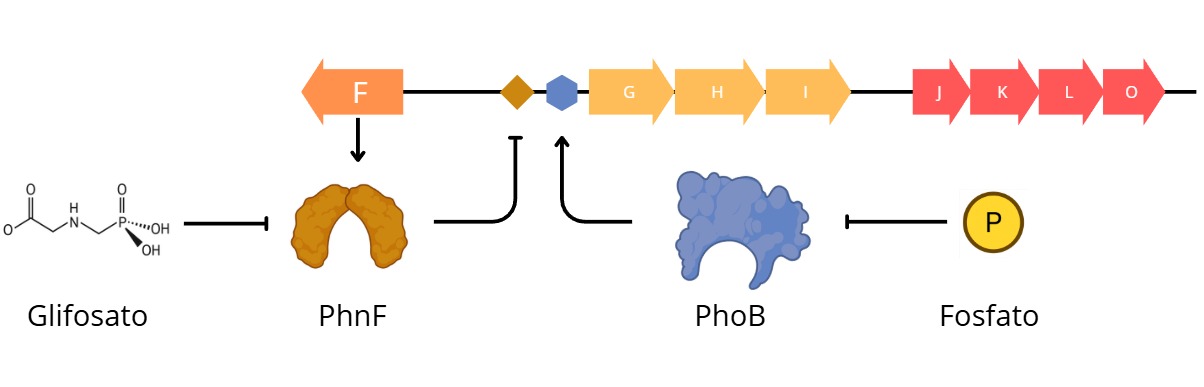

During my undergraduate thesis project, working as a research assistant at the Institute of Molecular and Celullar Biology (IBR-CONICET), I used this regulatory mechanism as a transcriptional phusion for the expression of the fluorescent protein GFP. In depletion of inorganic phosphorous and the prescent of a source of phosphonates, the promoter has an activity proportional to the amount of phosphonate in the media, then, by a non-linear regression the system can quantify the concentration or mass of gliphosate (or other phosphonates) in the media
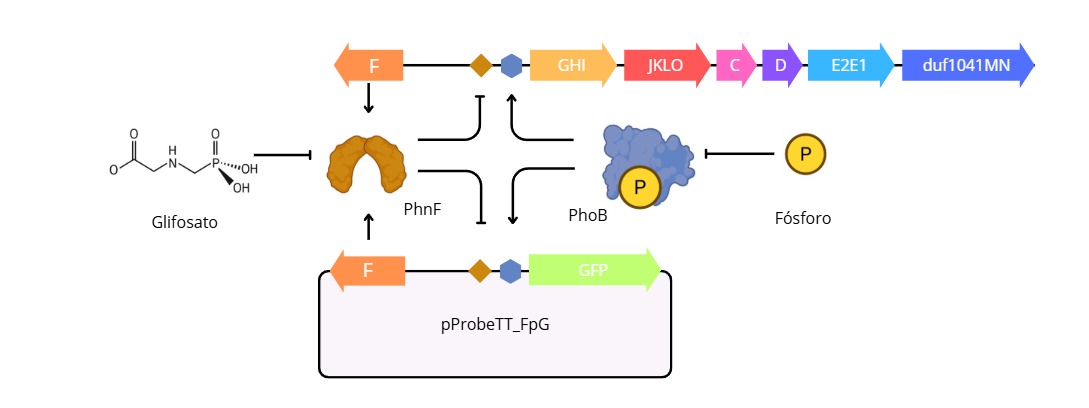

phosphonate metabolism transcriptional regulator PhnF [Agrobacterium tumefaciens] Protein sequence XHP91991.1 “MSPSAAMKRKNGVALWRQIADRIRGEIAAGDHDETGMLPPEMTLAEKFGVNRHTVRSAIAALASEGILEARQGRGTMIARKERLSFPISLRTRFTAGISDQVKDMQALLLSHTTEPANADLAARLQLQPGAPLVRLETLRKADSRPVSRSTTWFPADRFSGIGDAYQKSGSITAAFEMLGVADYVRISTVISASHADTQDLADLELAPGAILLVTQALNADMDGIPVQYAISRFSADTVEFTVEN”
For the reverse translation i used benchling own tool. Here are the specifications I used
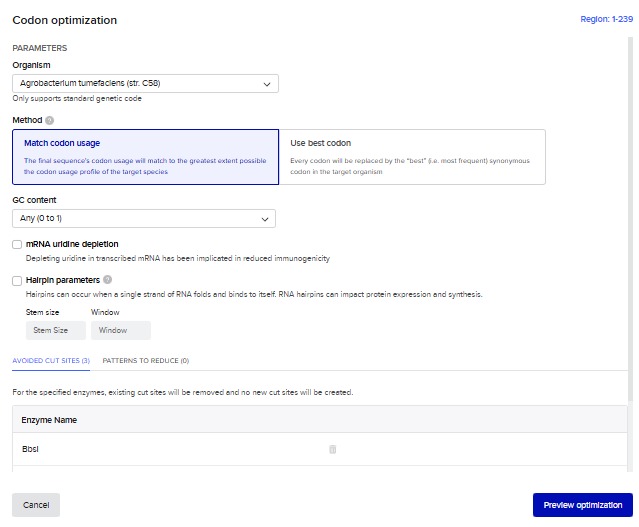

phosphonate metabolism transcriptional regulator PhnF [Agrobacterium tumefaciens] DNA sequence codon optimized “ATGAAGCGGAAGAATGGCGTAGCCCTCTGGCGCCAGATCGCGGATCGAATCCGTCAGGAAATCGCGGCCGGCGACCATGACGCCACCGGCATGCTTCCTCCAGAGATGACCCTGTCCGAAAAGTTCGGCGTCAACCGCCACACAGTGCGCTCCGCTATCGCCGCATTGGCGTCCGAGGGCATCGTCGAGCCGCTCCAGGGTCGCGGCACCATGATCGTGCGTAAGGGGCGCCTGAGCTTCCCGATCTCGAAGCGCACCCGGTTCACGGCCGGAATAGGCGAACAAGCCAAGGACATGGAGGCGCTGCTCCTGTCTCACAGCATCGAGAAGGCTACTGAGGATCTCGCCACGCGCCTTAAACTGGATGTCGGCGAACCGCTGATTAGAATGGAAACCCTTCGTAAAGCCGACCACCATCCGGTGTCGCGGTCCACGACCTGGTTCCCGGCGAAGCGCTTTTCAGGGATCGAGGAAGCCTATCGCGCGAGCGGCAGTATCACCGAGGCATTCGCCAAACTTGGCCTGAAAGACTATGTTAGGGATACGACGGTCATTAGCGCGACGCATGCGGACGCCGAAGATCTGTCGGACCTCGAACTGTCCCCCGGTGCCATCGTCTTGGTGACCTACGCACTCAACACCGATATTGATGGTGTCCCCGTGCAGTACGCTATCTCGCGGTTCTCGGCGGATACGGTTGAATTTACAATTGAAAAT”
For the expression of this protein, you could use a pET family vector in an E. coli JM109 starin, inducible by IPTG and translated by the T7 phage RNA polymerase. Then by cell lysis and protein extraction you could purificate (if you need) by adding an His x6 tag and in an Ni+2 column
Part 4
For this part I used the same sequence from part #3. I added individually every component as a DNA sequence to benchling (eg: promoter, rbs, terminator, His tag), then i edited the sequence of the codon optimized PhnF by adding this components in their respective place.

I actually couldn´t access the twist webpage, the site directed me to a local supplier in Argentina.

Even so, I found the pTwist ampR HighCopy cloning vector sequence and downloaded it into Benchling. With the unedited vector and the PhnF insert I simulated a Gibson Assembly in Benchling
Finally I obtained the final sequence of the vector with the PhnF insert with the correct essential components of the cloning.
The olny part i couldnt do is adding restriction enzymes for extracting the insert later. I wanted to add EcoRI sites in the sequence but I couldnt in the Gibson assembly system in benchling.
Part 5
DNA read
- What DNA would you want to sequence (e.g., read) and why?
I would like to perform genomic analysis of genetically modified crops. Last year I attended a seminar dictated by Dr. Juan Debernardi where he explained the services that they offer at the plant transformation facility in the UC-Davies. The offer a service where they transform a crop with an Agrobacterium tumefaciens strain and modify it with CrispR. The part that cautivated me the most was the develop of high efficiency CrsipR vector, for wich they performed genomic analysis for evoiding offtargeting of the crops.
- In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why?
I´d like to use a combination of long reads sequencing and short reads for the complete analysis of a genome. I think that this way you could have a bulk of big quality data for performing genomic analysis. For this you could use thecniques like Oxford Nanopore for the long read sequencing (Third generation) and Illumina (Second generation) for the short reads. The difference in the generations of the methods is due to the application of PCR and amplifications techniques used in Illumina where you need to use “connectors” for every individual read.
Specifically in the Illumina method you have to perform a Library preparation, involving the digestion of the DNA fragment into smaller pieces, and bind adapters to all the small molecules of DNA (arround 300 bp). Then you load the DNA into a flow cell and you amplify each single moleculle via bridge PCR to create a cluster of identical DNA strands. Then you sequence by synthesis using fluorescenctly labeled reversible termiantor nucleotides emmiting fluorescence in different wavelenghts so you can identify each base. Then yo can trasnform this fluorescent reads into short reads (about 300 bp) of DNA and assembly using a reference genome.
DNA write
- What DNA would you want to synthesize (e.g., write) and why?
I´d like to synthesize an antifungical gene for its expression in wheat. By designing the DNA in silico withouth the need of a backbone of a PCR, I could optimize the codons for their expression in the wheat specific genetic code. Also I could directly add al the components necessary for the expression like a constitutive plan promoter, a secretory signal and insert it into an agrobacterium based vector.
- What technology or technologies would you use to perform this DNA synthesis and why?
To implement this i would use Silicon based phosphoramidite synthesis to write the DNA molecule with a high fidelity throughput and for a low cost. Once the gene is synthesized and inserted into the wheat crop, I would use Illumina sequencing to verify the result. This assures that te gene is correctly inserted and there are no genetic complications within the genomic DNA of the crop. The limitations of this method are: the speed that requires due to the process of ligating the adapters, and the accuracy of the short reads, due to Wheat being an hexaplid there are many repetitive sequences, it could affect the quality of the short reads, in that case, there could be many alternatives for the genomic analysis of the genetically modified plant.
DNA Edit
- What DNA would you want to edit and why?
I would like to edit the genomic DNA of crops to improve both its nutritional quality and yield. There are many different method for plant transformation or editing, clasically the most direct approach is to transform a plant with and Agrobacterium strain crrrying a Ti derived plasmid that insert into the plant DNA in a random place. One of the most important aspects of plant editing is the design of protocols to circumvent the GMO regultations, like inserting a CrispR vector and eliminating it with plant repoduction thanks to the poliploid caractheristics that modern crops carry. But there are different alternatives to plan editing that do not involve adding a transgene like delivering directly the enzymes for editing in protoplasts or using a viral derived vector. Editing plants to improve their nutritional quality or yield is fundamental for the
- What technology or technologies would you use to perform these DNA edits and why?
To do this I could perform precision Knockouts using CRIPSR to break different genes. I also could use base editors to change a single base in a gene. The future of plant editing is bright, every year, new technologies are being developed, to modify genes without adding a transgene, also, in the developing of new techniques, old ones are not forgotten, like in random mutagenesis using gamma rays or chemical mutagens, the plant phenotipation field is developing quickly.
If i had to choose CrispR to design a knockdown experiment, first I had to get the ORF of the gene from a species specific library, then develop the guide crispr RNA, by using different genomic analysis software to mimimize the off-targeting of the editor, then decide wich Crispr-Cas system i would use, bucause it determines the PAM region, and different CAS enzymes have different functions and effectivness. Then, once the CrispR vector is designed, i would have to build and transfrom it into an Agrobacterium tumefaciens strain and transform it into the crop by floral dipping or plant regeneration, the select the transformants by antibiotic selection and generate the offspring of three or four generations. The most important limitations of a CrispR editing protocol is that it is time consuming, it can take several months to grow a fully developed crop from a graph, and somateimes it could take some several generations for the mutation to stabilize. Another aspect to consider is the natural defense mechanisms that plants have against viruses, involving the DICER protein that degrades single stranded mRNA, if you express your crispR vector in large quantities, using a constitutive vector for example, the plant would recognize the mRNA as foreign and degrade it, expanding the resistance to all the plant and evading the transation of the Cas system. Also, some crispR systems have a very low editing efficiency and even if the transgene is present, the cas system does not edit the gene, in those cases you have to develop the crispR vector all over again or chose a different Cas system.