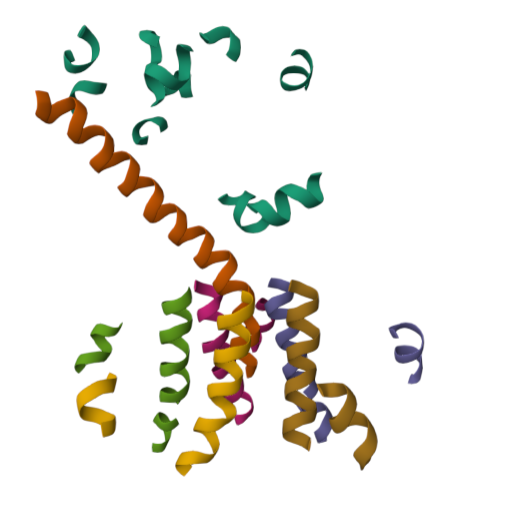hw #4-protein-design-part-i
Part A. Conceptual questions
Questions from Shunguan zhang
- Since meat isnt 100% protein, we have to calculate the porcentage of protein that this piece of meat has. Researching i found that is on average 22%, meaning that of those 500 g of meat, only 110 g are proteins.
Then, dividing the total mass of protein (110 g) for the average molar mass of amino acids (100 g/mol) we find that there is 1.1 moles of amino acids in this piece of meat.
Then, by multiplying by avogadro´s number (6.022 x 1023) we could obtain the aproximatedly number of aminoacids in this piece, that is 6.62 x 1023 aminoacids in a 500 g piece of meat.
That is because when we eat food, we actually break down that complex matrix of cells and molecules in simple ones that we can absorb and distribute to our body, for example, we degrade the proteins in very small chains called polypeptides or even aminoacids, by doing that, the intestinal epitelium can absrob this small molecular by different molecular transport systems. Also, since the aminoacids are essentialy the same by most of living organisms the we can use them to for our proteins.
a
Yes, while mainteining the backbone of the amnioacid you can modify the R group at your will. If you want to design your own aminoacid you will have to think of the functionality that this new group brings into the mix, you will have to consider the interactions that this new functional group has with other aminoacids, you could probably predict them by understandig the caracteristics of this molecule, if it is acid, basic, polar or non polar. I found an intresting example of a synthetic aminoacid, the adition of an azide group to a standard phenylalanine structure, with this new aminoacid you could tag proteins in vivo without killing the cell and you could see the actual interactions of this protein in real time.
It is belived that aminoacids were formed by simple molecules (like water, methane, ammonia and hydrogen) that reacted with the energy provided with electricity in the early ages of the earth. This has been experimented in the famous Stanley Miller experiment, in wich after combining this compunds found in ancient earth with a zap of electricity found a variety of aminoacids.
The chirality of the helix is dictated by the orientation fo the side chains. In a right-handed helix, the side chains point outward and away from the backbone, minimizing steric hindrance. If you tried to force L-amino acids into a left-handed helix, the side chains would bump into the backbone and the structure would be unstable. With D-amino acids, the atoms are arranged in a mirror-image configuration, the geometry is reversed. To keep those side chains pointing safely outward, the entire backbone must twist in the opposite direction, creating a left-handed spiral.
Yes, amino acid chains can twist into several distinct helical shapes depending on how tightly they pack the AA´s.
- The 3(10) helix: this is the second most common helix. It is tighter and thinner than a standard alpha helix. In this one there are exactly 3 AA´s per turn, less than the standar 3.6 in alpha helixes.
- the pi-helix: because of the unique ring shape of Proline forces a different geometry of the helix
This is because almost all life on earth uses L-amino acids, the right handed twists of the helixes allows the side chains to pint outward and interact with another groups of molecules, giving the proteins their functionality. If the side chains were pinting inward, there will be an esteric impediment and probably the proteins will not maintain their biological activity.
That is because in a b-like sheet the chemical groups responsible of hydrogen bonding point outward from the edges, in this way, b-sheets are designed to link up with others.
The reason amyloid diseases are defined by b-sheets is that this is the most thermofynamically stable low energy state of a protein. When a protein misfold, it falls into this lowest state of energy. Differently to alpha sheet (that rely hevily in the side chains for the interactiosn) b-sheets are held together by the protein backbone.
Part B. Analysis and visualization
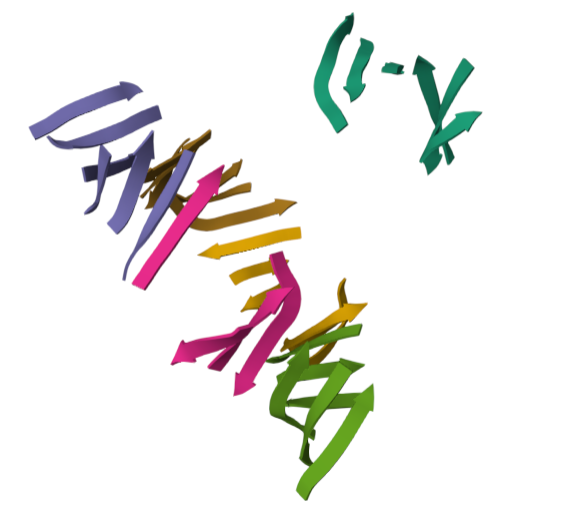
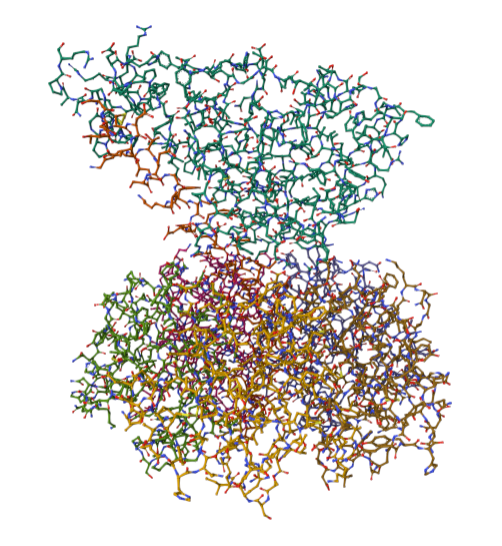
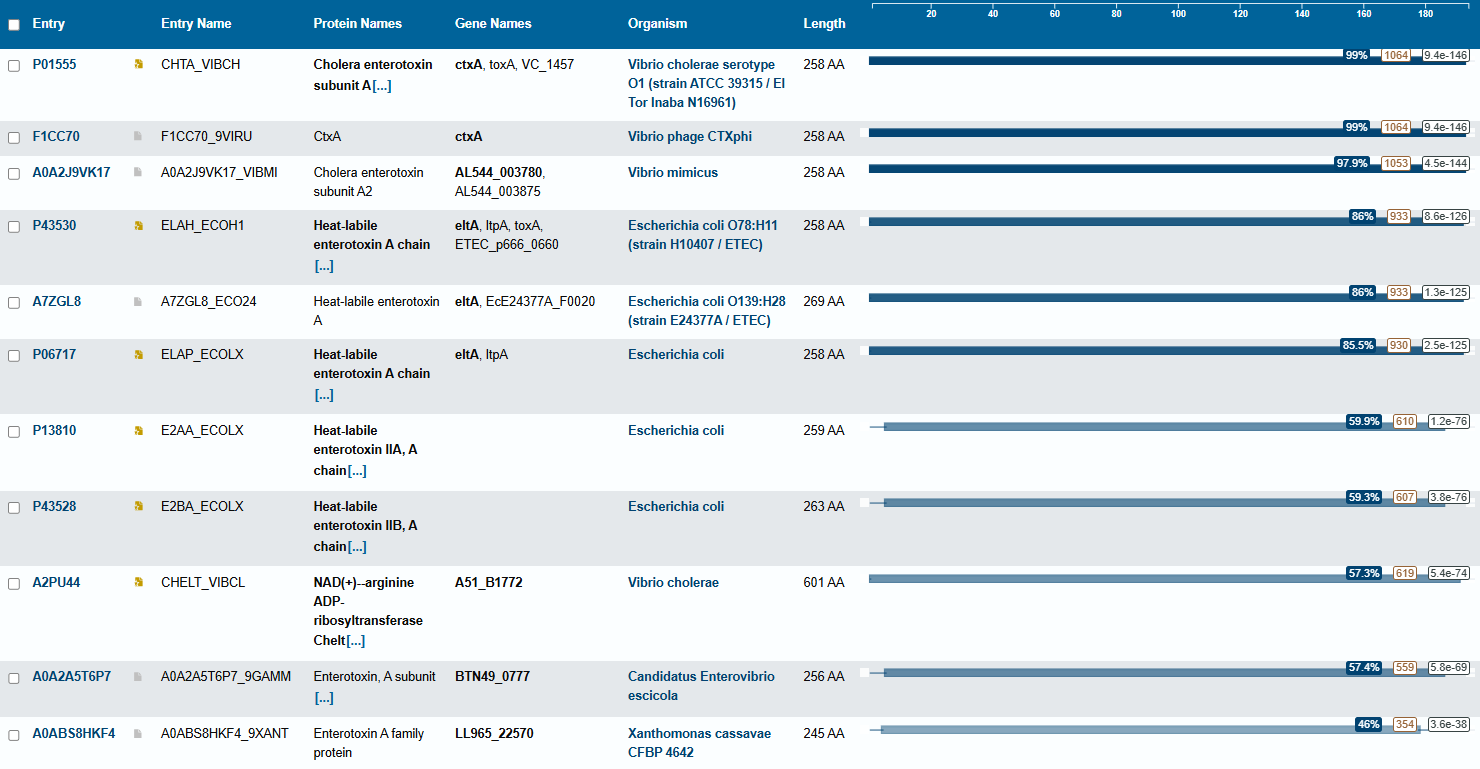
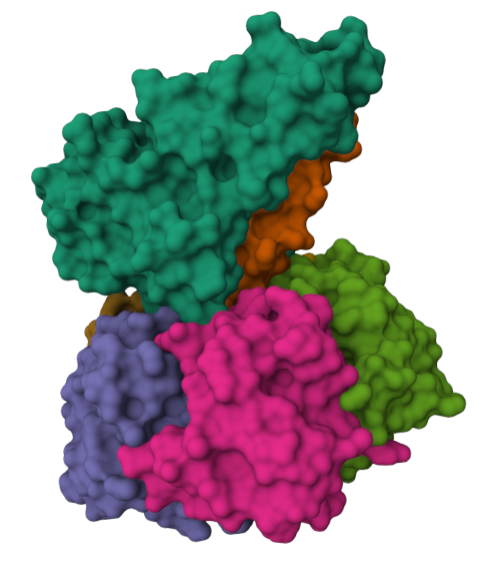
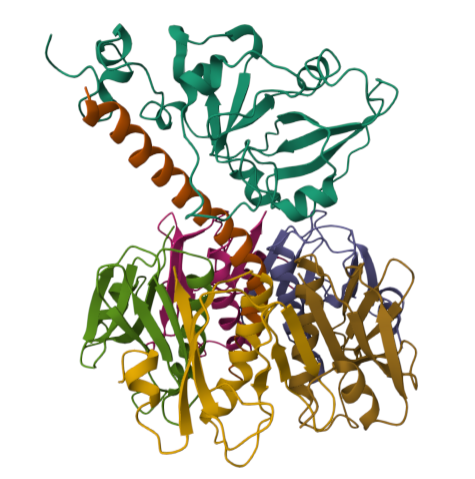
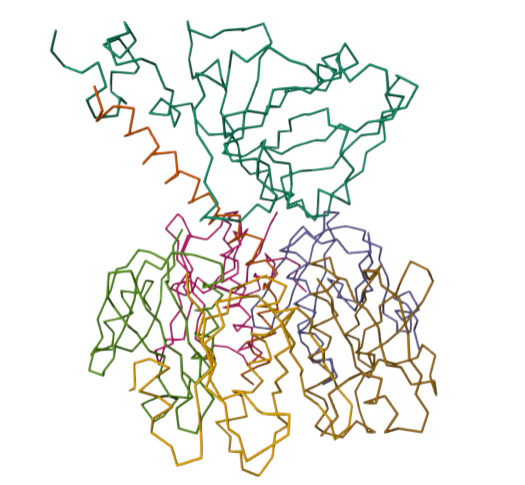
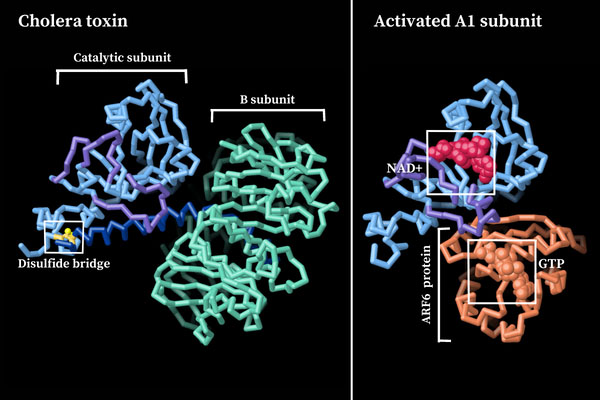
For this part of the homework i actually choose one of the repertory of “the protein of the month” on PDB. This molecule is the cholera toxin. this protein attacks intesitanl cells to cause life-threatening dehydration. It contains two sub units A (shown in blue) includes the toxic enzyme portion (A1 chain) and the linker (A2 chain), and subunit B (shown in turquoise) includes the carbohydrate-binding portion. When the toxin binds to a cell, the disulfide bridge (shown in yellow) that connects the two chains of the A subunit is then broken, releasing the toxic portion (shown in light blue) into the cell. The structure on the right (PDB entry 2a5f) shows that when the A1 chain binds to the ARF6 protein (shown in orange, with a bound GTP), the toxin’s catalytic loops (shown in yellow) undergo conformational changes and NAD+ (shown in green) binds to the active site. The activated A1 subunit can then attach an ADP group to permanently activate a G-protein.

Protein sequence
For the simplicity of this part of the assignmen i will only use the Chain A of the toxin.
1XTC_1|Chain A|CHOLERA TOXIN|Vibrio cholerae (44104) NDDKLYRADSRPPDEIKQSGGLMPRGQSEYFDRGTQMNINLYDHARGTQTGFVRHDDGYVSTSISLRSAHLVGQTILSGHSTYYLYVLATAPNMFNVNDVLGAYSPHPDEQEVSALGGIPYSQIYGWYRVHFGVLDEQLHRNRGYRDRYYSNLDIAPAADGYGLAGFPPEHRAWREEPWIHHAPPGCGNAPRSS
The length of the protein is: 240 aminoacids. The most common amino acid is: G, which appears 22 times.
Sequence Length: 240 amino acids
Amino Acid Frequencies: G: 22 (9.17%) S: 21 (8.75%) D: 19 (7.92%) Y: 19 (7.92%) R: 17 (7.08%) L: 16 (6.67%) A: 15 (6.25%) P: 14 (5.83%) I: 12 (5.00%) Q: 12 (5.00%) N: 11 (4.58%) E: 11 (4.58%) V: 10 (4.17%) T: 9 (3.75%) H: 9 (3.75%) K: 7 (2.92%) F: 7 (2.92%) M: 4 (1.67%) W: 3 (1.25%) C: 2 (0.83%)