Week 2 Homework: DNA Read, Write & Edit
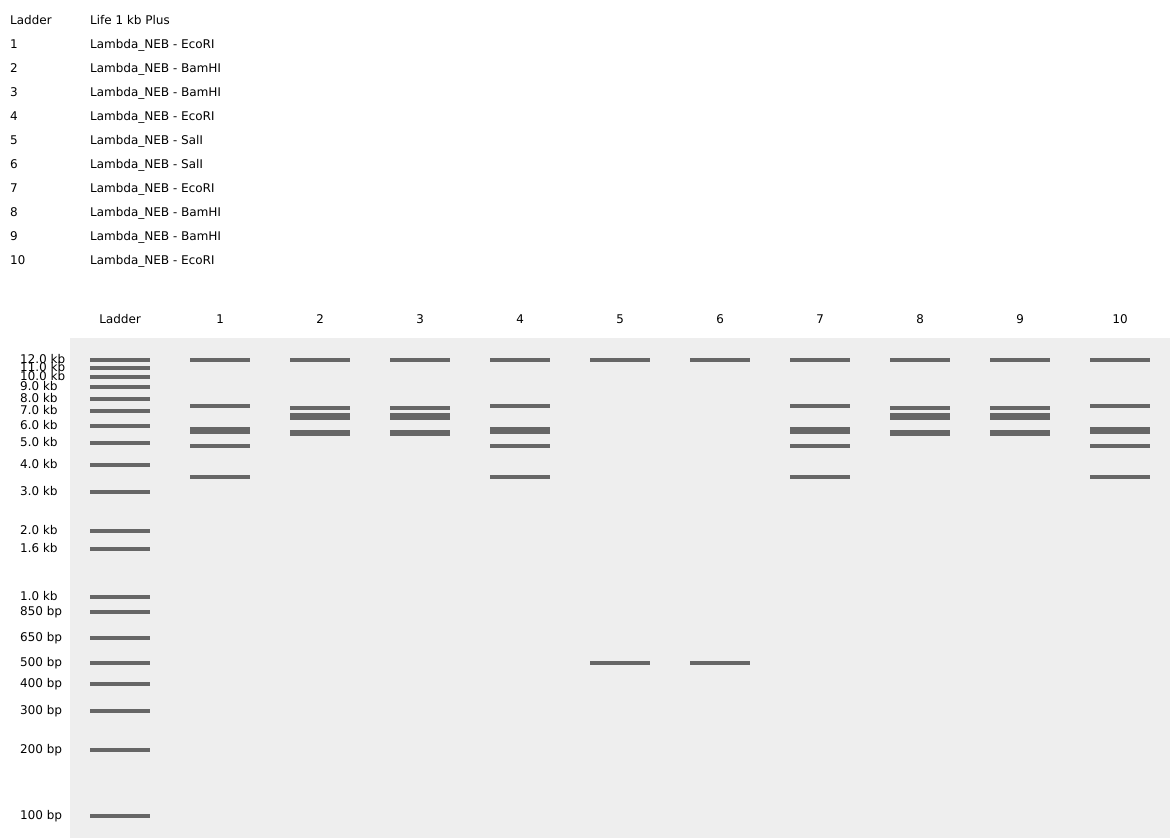
PART 1: Gel Art
For the Gel Art part, I searched Roman’s Gallery and chose the little smiley face :).

PART 3: DNA Design Challenge
3.1 Choose Your Protein
The protein I have chosen for this DNA design challenge is Sonic Hedgehog (Shh), a critical signaling molecule involved in embryonic development, tissue patterning, and organ formation. Shh plays a central role in directing cell fate decisions during development, and mutations in this protein are linked to severe developmental disorders. Its importance in biology, combined with the fact that it is well-characterized and has a known amino acid sequence, makes it an ideal candidate for this exercise.
MLPNIILILLIRYCSCGAGSRVYEKYGKQVQLSPATTSSYREWYDSNREHSTRNNTNVDD FQTQLKKSLENTTAAYNATFMQELIEERQRYLEKLNEGQFINDQRRLVEELLDPNYYEKT VHPKRDYTRPTRVNLSMSLYQILDVDEHMQSIEVNVWMVQHWYDEFLDWNPVDYGMINRT IVPYHQIWIPDTYLYNSEELEQKKTESLMNAQLETGHWNQKKDGAKVQLMFPAIYKLSCR MDVRWFPYDRQNCTFIISSWTHDKQTIDYWPLSSTVNLGNMARNDEWEVISFEFVRVEET FKCCTAPWVMLYAHLVIRRKPLYYMINLVVPTSIITIVAVTGFFTPTSSSSERDEKLYLG INTLLTMSVMMLMVCNQMPSTSTYVPLMSWYYIGIIMVIVVGTFLATGVLAIHGQKHYNK PISDRIRKLIYNPVVEFFILSPPTSLIDLWTEFGVISEQRHSTHLDPLLLQHMDPISHTT RADPQHFFGSISSQMCDLQSTYSYTARLATITRQYTQHAKMKALRKNQYRMSMDTSQARG VKKQKMQRRCSLEWEFLANVLDRILLTIFCGFTFAVFIILIGFDSFFTFHTDSPPKTM
3.2 Reverse Translate: Protein → DNA
Using the amino acid sequence from UniProt, I performed a reverse translation to determine the nucleotide sequence that could encode Shh. Reverse translation is the process of inferring a DNA sequence from a protein sequence, based on the codons that encode each amino acid. Because multiple codons can code for the same amino acid, this step may produce several possible DNA sequences. The resulting DNA sequence represents the gene that could be transcribed and translated to produce the Shh protein, reflecting the principles of the Central Dogma, where DNA is transcribed into RNA and RNA is translated into protein.
ATGATGCTGCCGAACATTATTCTGATTCTGCTGATTCGCTATTGCAGCTGCGGCGCGGGCAGCCGCGTGTATGAAAAATATGGCAAACAGGTGCAGCTGAGCCCGGCGACCACCAGCAGCTATCGCGAATGGTATGATAGCAACCGCGAACATAGCACCCGCAACAACACCAACGTGGATGATTTTCAGACCCAGCTGAAAAAAAGCCTGGAAAACACCACCGCGGCGTATAACGCGACCTTTATGCAGGAACTGATTGAAGAACGCCAGCGCTATCTGGAAAAACTGAACGAAGGCCAGTTTATTAACGATCAGCGCCGCCTGGTGGAAGAACTGCTGGATCCGAACTATTATGAAAAAACCGTGCATCCGAAACGCGATTATACCCGCCCGACCCGCGTGAACCTGAGCATGAGCCTGTATCAGATTCTGGATGTGGATGAACATATGCAGAGCATTGAAGTGAACGTGTGGATGGTGCAGCATTGGTATGATGAATTTCTGGATTGGAACCCGGTGGATTATGGCATGATTAACCGCACCATTGTGCCGTATCATCAGATTTGGATTCCGGATACCTATCTGTATAACAGCGAAGAACTGGAACAGAAAAAAACCGAAAGCCTGATGAACGCGCAGCTGGAAACCGGCCATTGGAACCAGAAAAAAGATGGCGCGAAAGTGCAGCTGATGTTTCCGGCGATTTATAAACTGAGCTGCCGCATGGATGTGCGCTGGTTTCCGTATGATCGCCAGAACTGCACCTTTATTATTAGCAGCTGGACCCATGATAAACAGACCATTGATTATTGGCCGCTGAGCAGCACCGTGAACCTGGGCAACATGGCGCGCAACGATGAATGGGAAGTGATTAGCTTTGAATTTGTGCGCGTGGAAGAAACCTTTAAATGCTGCACCGCGCCGTGGGTGATGCTGTATGCGCATCTGGTGATTCGCCGCAAACCGCTGTATTATATGATTAACCTGGTGGTGCCGACCAGCATTATTACCATTGTGGCGGTGACCGGCTTTTTTACCCCGACCAGCAGCAGCAGCGAACGCGATGAAAAACTGTATCTGGGCATTAACACCCTGCTGACCATGAGCGTGATGATGCTGATGGTGTGCAACCAGATGCCGAGCACCAGCACCTATGTGCCGCTGATGAGCTGGTATTATATTGGCATTATTATGGTGATTGTGGTGGGCACCTTTCTGGCGACCGGCGTGCTGGCGATTCATGGCCAGAAACATTATAACAAACCGATTAGCGATCGCATTCGCAAACTGATTTATAACCCGGTGGTGGAATTTTTTATTCTGAGCCCGCCGACCAGCCTGATTGATCTGTGGACCGAATTTGGCGTGATTAGCGAACAGCGCCATAGCACCCATCTGGATCCGCTGCTGCTGCAGCATATGGATCCGATTAGCCATACCACCCGCGCGGATCCGCAGCATTTTTTTGGCAGCATTAGCAGCCAGATGTGCGATCTGCAGAGCACCTATAGCTATACCGCGCGCCTGGCGACCATTACCCGCCAGTATACCCAGCATGCGAAAATGAAAGCGCTGCGCAAAAACCAGTATCGCATGAGCATGGATACCAGCCAGGCGCGCGGCGTGAAAAAACAGAAAATGCAGCGCCGCTGCAGCCTGGAATGGGAATTTCTGGCGAACGTGCTGGATCGCATTCTGCTGACCATTTTTTGCGGCTTTACCTTTGCGGTGTTTATTATTCTGATTGGCTTTGATAGCTTTTTTACCTTTCATACCGATAGCCCGCCGAAAACCATGTAA
3.3 Codon Optimization
Once the DNA sequence is determined, I codon-optimized it for E. coli expression. Codon optimization is important because different organisms prefer certain codons over others for the same amino acid. Using preferred codons in E. coli increases translation efficiency, improves protein yield, and avoids rare codons that can slow ribosomes or create secondary structures. This ensures robust expression of Shh in bacterial systems.
3.4 You Have a Sequence! Now What?
With a codon-optimized Shh sequence, several methods can be used to produce the protein:
- Cell-dependent expression: Clone the DNA into a plasmid vector and introduce it into E. coli. The bacterial machinery transcribes the DNA into mRNA and translates it into Shh protein.
- Cell-free systems: Produce Shh protein directly in vitro using transcription-translation mixtures from E. coli lysates.
Both approaches demonstrate how a nucleotide sequence can be converted into a functional protein through natural molecular processes.
3.5 How Does it Work in Nature
In nature, the Shh gene is transcribed into mRNA and translated into a precursor protein, which then undergoes post-translational modifications, including cleavage and lipidation. These modifications are required for proper secretion and signaling activity. This illustrates how a single gene can produce a biologically active protein, highlighting the flow of information from DNA → RNA → protein and the importance of transcriptional and post-translational regulation.
PART 4: Prepare a Twist Synthesis Order
PART 5: DNA Read, Write, and Edit – Microbiome Skincare Applications
5.1 DNA Read – Sequencing
DNA to sequence and why: I would sequence skin microbiome DNA from individuals with different skin types or conditions, such as acne-prone, sensitive, or aged skin. This allows identification of beneficial and harmful microbes, their functional genes, and metabolites affecting skin health. Sequencing this DNA can inform development of probiotic treatments, personalized skincare, and microbial-derived bioactives.
Sequencing technology and details: I would use Oxford Nanopore long-read sequencing, a third-generation method that can read full-length microbial genomes and plasmids. Input DNA would be extracted from skin swabs, lightly sheared, and ligated with nanopore adapters. Essential steps include passing DNA through a protein nanopore, measuring ionic current changes, and decoding bases from these signal variations. The output is long-read sequences of microbial genomes and plasmids, helping resolve strain-level diversity. Limitations include lower per-read accuracy compared with short-read sequencing, but high coverage and error-correction mitigate this.
5.2 DNA Write – Synthesis
DNA to synthesize and why: I would synthesize genes encoding skin-beneficial metabolites, such as bacterial enzymes producing natural moisturizers, antioxidants, or antimicrobial peptides. For example, a sequence coding for a bacterial ceramide-synthesizing enzyme could be used to engineer probiotic strains that restore skin barrier function. This approach could create living or topical treatments that enhance skin health naturally.
Technology for synthesis and details: I would use Twist Bioscience’s array-based high-throughput DNA synthesis. Essential steps include sequential nucleotide addition via phosphoramidite chemistry, oligo cleavage and purification, and assembly into full-length gene constructs. Limitations include sequence length constraints (<200 bases per oligo), potential synthesis errors, and optimization requirements when expressing microbial genes in target probiotic strains.
5.3 DNA Edit – Editing
DNA to edit and why: I would edit the genome of commensal skin bacteria like Staphylococcus epidermidis to enhance production of anti-inflammatory molecules or UV-protective compounds. This could reduce acne, eczema flare-ups, or sun-induced skin damage while maintaining microbiome balance. Edits might include upregulating genes for antimicrobial peptides or protective pigments.
Technology for editing and details: I would use CRISPR-Cas9 coupled with plasmid-based delivery, a precise genome editing tool for bacteria. Essential steps include designing guide RNAs targeting regulatory regions, delivering Cas9 and repair templates into bacterial cells, and screening for successful edits. Inputs include plasmids carrying Cas9, guide RNAs, and donor DNA. Limitations include variable editing efficiency across bacterial strains, potential off-target mutations, and challenges in ensuring stable long-term expression of engineered traits.