Week 5 Homework: Protein Design Part II
Week 5 - Protein Design Part II
Part A: SOD1 Binder Peptide Design (From Pranam)
Part 1: Generate Binders with PepMLM
SOD1 sequence with A4V mutation:
ATKVVCVLKGDGPVQGIINFEQKESNGPVKVWGSIKGLTEGLHGFHVHEFGDNTAGCTSAGPHFNPLSRKHGGPKDEERHVGDLGNVTADKDGVADVSIEDSVISLSGDHCIIGRTLVVHEKADDLGKGGNEESTKTGNAGSRLACGVIGIAQ
| Peptide # | Peptide Sequence | Perplexity Score |
|---|---|---|
| 1 | WHYYVTAIEWKK | 25.725023 |
| 2 | KHYYWVAIRWKK | 27.40476 |
| 3 | HRSPVVGVALKK | 12.718177 |
| 4 | WRYPVAAIELKE | 17.102567 |
| 5 | FLYRWLPSRRGG | N/A |
Part 2: Evaluate Binders with AlphaFold3
| Peptide # | Peptide Sequence | ipTM |
|---|---|---|
| 1 | WHYYVTAIEWKK | 0.31 |
| 2 | KHYYWVAIRWKK | 0.4 |
| 3 | HRSPVVGVALKK | 0.41 |
| 4 | WRYPVAAIELKE | 0.3 |
| 5 | FLYRWLPSRRGG | 0.28 |
| Peptide # | Peptide Sequence | ipTM | Binding Description |
|---|---|---|---|
| 1 | WHYYVTAIEWKK | 0.31 | Binds near the surface of the β-barrel;; mostly surface-exposed. |
| 2 | KHYYWVAIRWKK | 0.40 | Appears bound on the β-barrel and dimer interface, surface-bound |
| 3 | HRSPVVGVALKK | 0.41 | Localized near the β-barrel surface, surface-bound. |
| 4 | WRYPVAAIELKE | 0.30 | Surface-bound,minimal contact with N-terminal residues, approaches dimer interface. |
| 5 | FLYRWLPSRRGG | 0.28 | Surface-bound, does not penetrate deeply into β-barrel. |
The ipTM scores for the PepMLM-generated peptides range from 0.30 to 0.41, slightly higher than the known binder (0.28). Peptides 2 and 3 show the highest ipTM scores, indicating stronger predicted structural confidence in the SOD1–peptide complex.
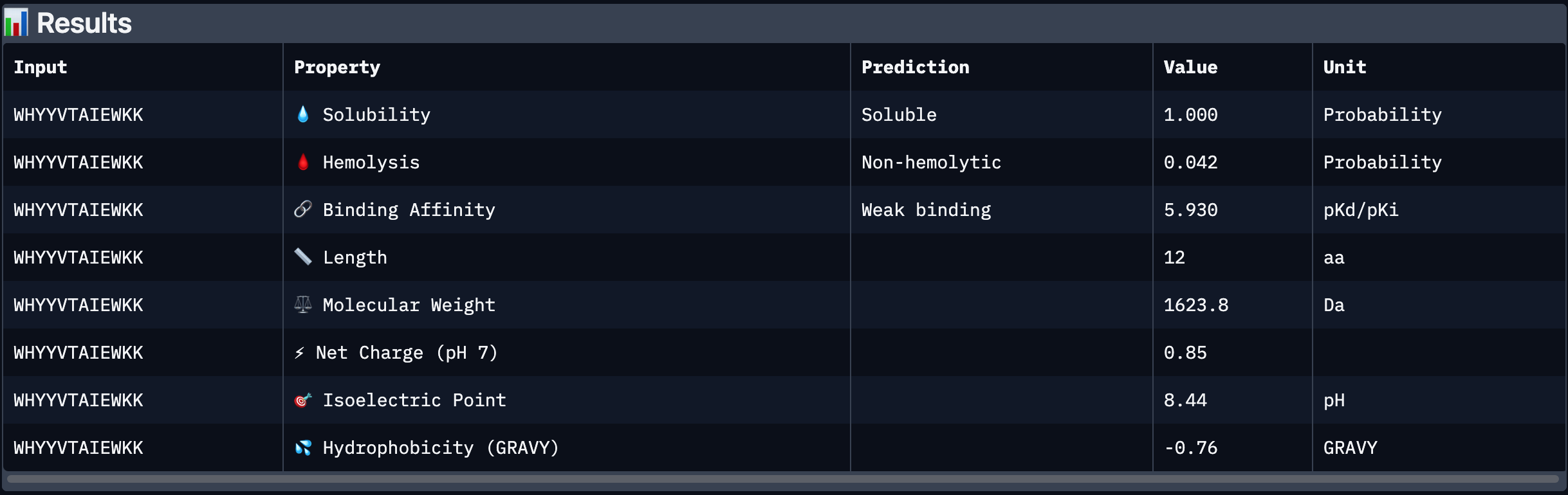
Part 3: Evaluate Properties of Generated Peptides in the PeptiVerse
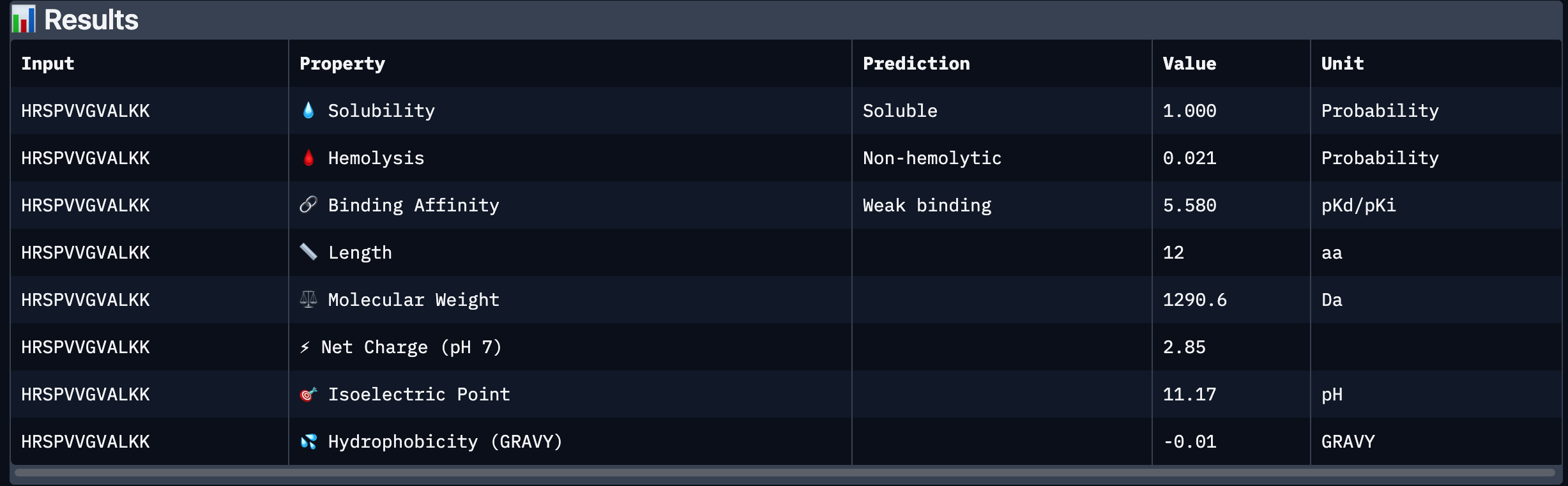
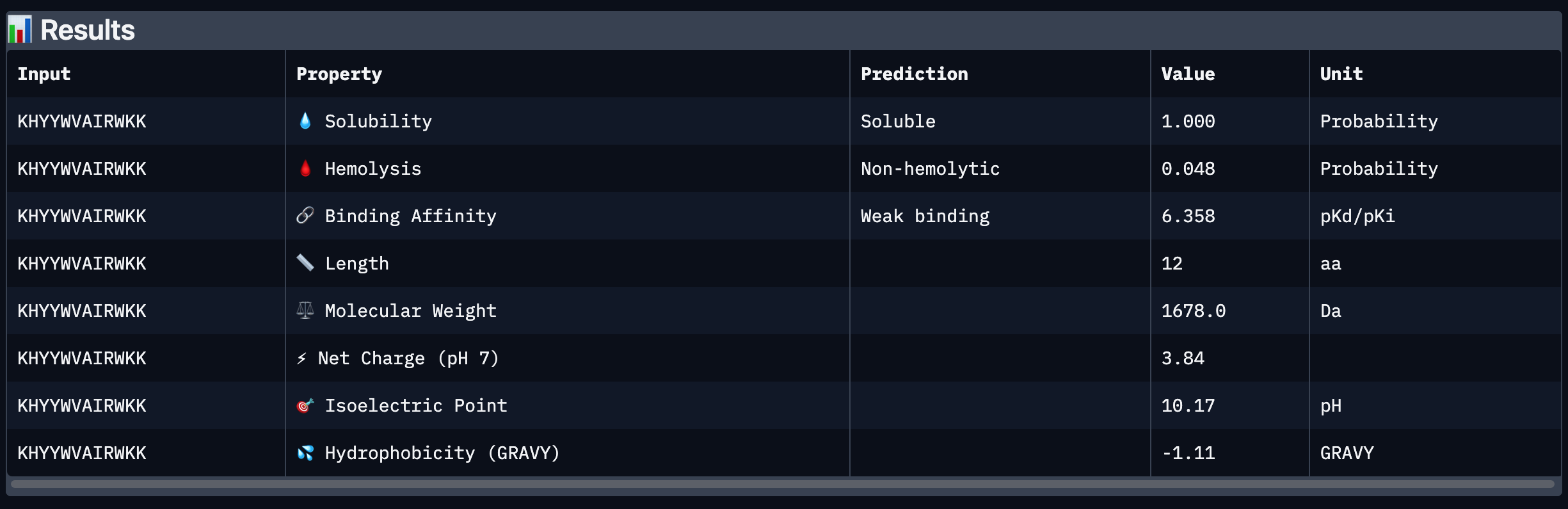
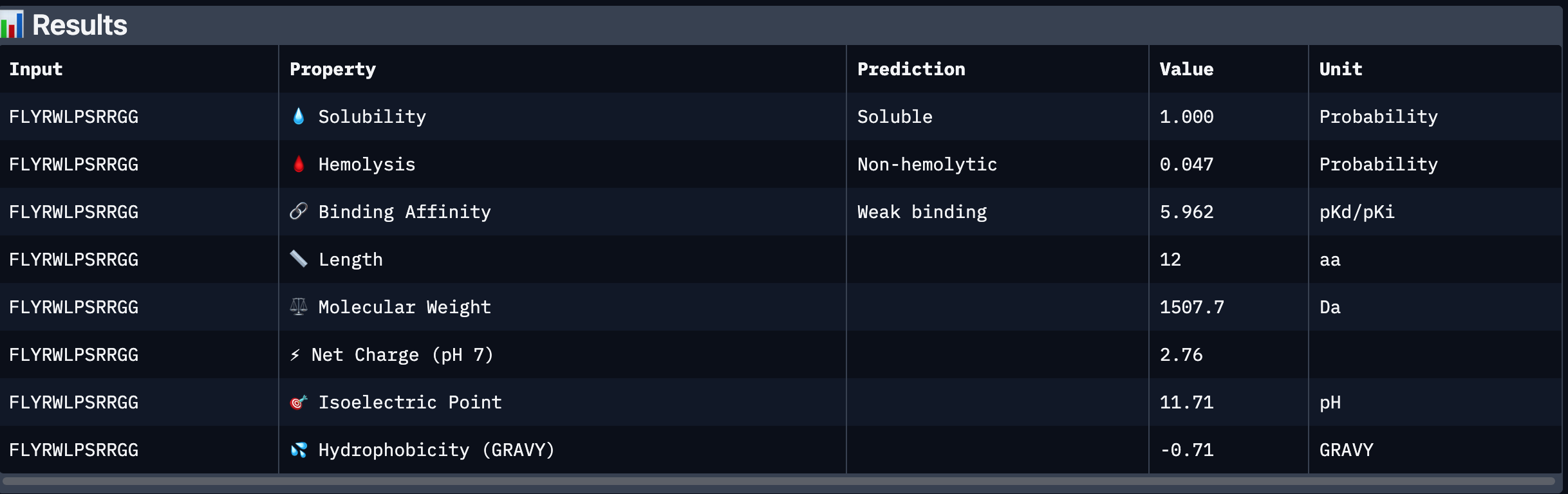
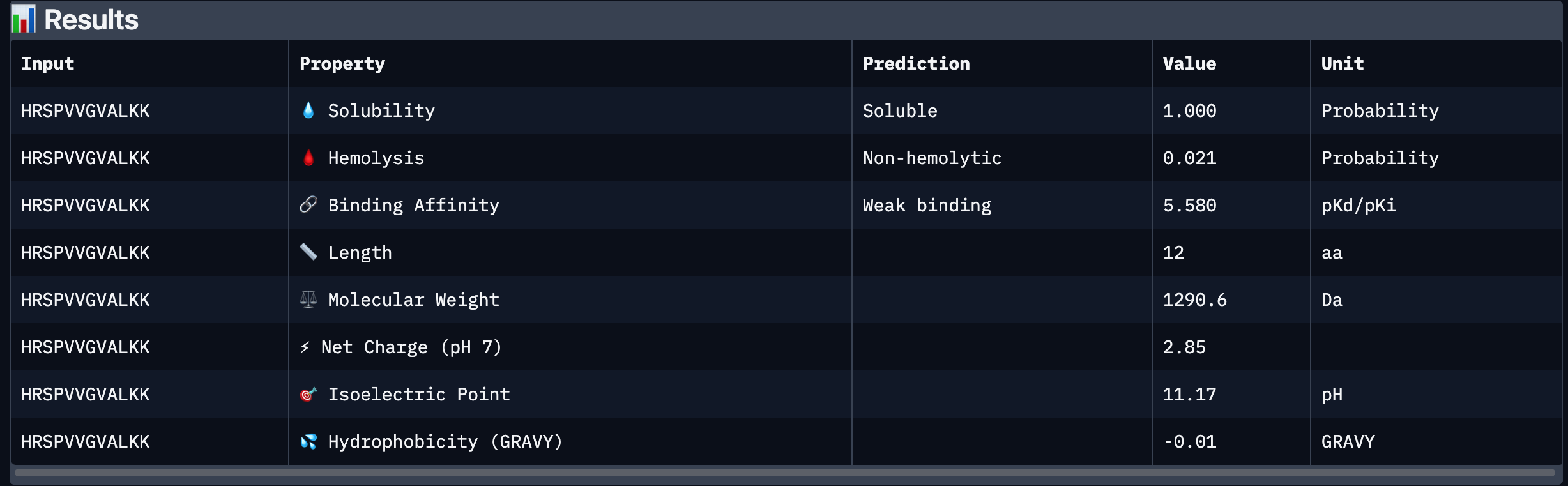
After analysing all the peptides and their predicted binding and therapeutic properties I concluded that their affinity binding does not rise with the ipTM index, all of my peptides are soluble( constant index of 1) and are non-hemolytic.
I chose peptide 2 (HRSPVVGVALKK), because it has one of the highest ipTM (0.40)and the highest binding affinity(6.35).