Week 2 HW:DNA read write and edit
Part 1: Benchling & In-silico Gel Art
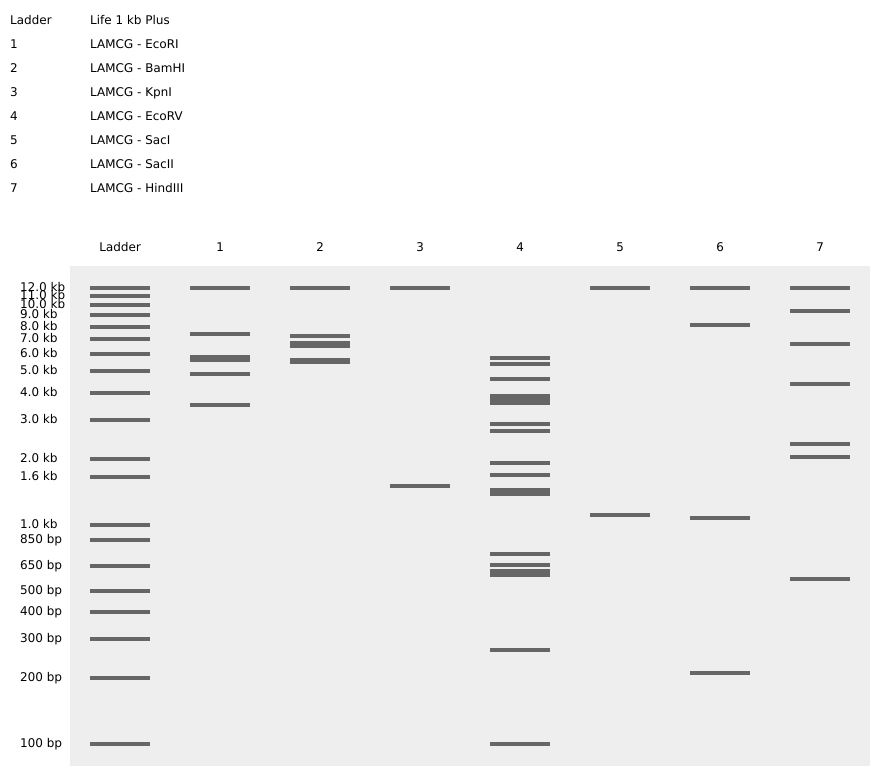

Figure 1. Simulated agarose gel showing the LAMCG plasmid digested with different restriction enzymes.

Figure 2. Screenshot of the online tool used to simulate restriction digests and generate the virtual gel pattern for LAMCG.
Testing update
I was arranging the bands to try to form the letter “M.”
Part 3: DNA Design Challenge
- 3.1. Choose your protein.
I choose protein HSP90
Original sequence: *>sp|P07900|HS90A_HUMAN Heat shock protein HSP 90-alpha OS=Homo sapiens OX=9606 GN=HSP90AA1 PE=1 SV=5 MPEETQTQDQPMEEEEVETFAFQAEIAQLMSLIINTFYSNKEIFLRELISNSSDALDKIRYESLTDPSKLDSGKELHINLIPNKQDRTLTIVDTGIGMTKADLINNLGTIAKSGTKAFMEALQAGADISMIGQFGVGFYSAYLVAEKVTVITKHNDDEQYAWESSAGGSFTVRTDTGEPMGRGTKVILHLKEDQTEYLEERRIKEIVKKHSQFIGYPITLFVEKERDKEVSDDEAEEKEDKEEEKEKEEKESEDKPEIEDVGSDEEEEKKDGDKKKKKKIKEKYIDQEELNKTKPIWTRNPDDITNEEYGEFYKSLTNDWEDHLAVKHFSVEGQLEFRALLFVPRRAPFDLFENRKKKNNIKLYVRRVFIMDNCEELIPEYLNFIRGVVDSEDLPLNISREMLQQSKILKVIRKNLVKKCLELFTELAEDKENYKKFYEQFSKNIKLGIHEDSQNRKKLSELLRYYTSASGDEMVSLKDYCTRMKENQKHIYYITGETKDQVANSAFVERLRKHGLEVIYMIEPIDEYCVQQLKEFEGKTLVSVTKEGLELPEDEEEKKKQEEKKTKFENLCKIMKDILEKKVEKVVVSNRLVTSPCCIVTSTYGWTANMERIMKAQALRDNSTMGYMAAKKHLEINPDHSIIETLRQKAEADKNDKSVK DLVILLYETALLSSGFSLEDPQTHANRIYRMIKLGLGIDEDDPTADDTSAAVTEEMPPLEGDDDTSRMEEVD
- 3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
reverse translation of sp|P07900|HS90A_HUMAN Heat shock protein HSP 90-alpha OS=Homo sapiens OX=9606 GN=HSP90AA1 PE=1 SV=5 to a 2196 base sequence of most likely codons. atgccggaagaaacccagacccaggatcagccgatggaagaagaagaagtggaaacctttgcgtttcaggcggaaattgcgcagctgatgagcctgattattaacaccttttatagcaacaaagaaatttttctgcgcgaactgattagcaacagcagcgatgcgctggataaaattcgctatgaaagcctgaccgatccgagcaaactggatagcggcaaagaactgcatattaacctgattccgaacaaacaggatcgcaccctgaccattgtggataccggcattggcatgaccaaagcggatctgattaacaacctgggcaccattgcgaaaagcggcaccaaagcgtttatggaagcgctgcaggcgggcgcggatattagcatgattggccagtttggcgtgggcttttatagcgcgtatctggtggcggaaaaagtgaccgtgattaccaaacataacgatgatgaacagtatgcgtgggaaagcagcgcgggcggcagctttaccgtgcgcaccgataccggcgaaccgatgggccgcggcaccaaagtgattctgcatctgaaagaagatcagaccgaatatctggaagaacgccgcattaaagaaattgtgaaaaaacatagccagtttattggctatccgattaccctgtttgtggaaaaagaacgcgataaagaagtgagcgatgatgaagcggaagaaaaagaagataaagaagaagaaaaagaaaaagaagaaaaagaaagcgaagataaaccggaaattgaagatgtgggcagcgatgaagaagaagaaaaaaaagatggcgataaaaaaaaaaaaaaaaaaattaaagaaaaatatattgatcaggaagaactgaacaaaaccaaaccgatttggacccgcaacccggatgatattaccaacgaagaatatggcgaattttataaaagcctgaccaacgattgggaagatcatctggcggtgaaacattttagcgtggaaggccagctggaatttcgcgcgctgctgtttgtgccgcgccgcgcgccgtttgatctgtttgaaaaccgcaaaaaaaaaaacaacattaaactgtatgtgcgccgcgtgtttattatggataactgcgaagaactgattccggaatatctgaactttattcgcggcgtggtggatagcgaagatctgccgctgaacattagccgcgaaatgctgcagcagagcaaaattctgaaagtgattcgcaaaaacctggtgaaaaaatgcctggaactgtttaccgaactggcggaagataaagaaaactataaaaaattttatgaacagtttagcaaaaacattaaactgggcattcatgaagatagccagaaccgcaaaaaactgagcgaactgctgcgctattataccagcgcgagcggcgatgaaatggtgagcctgaaagattattgcacccgcatgaaagaaaaccagaaacatatttattatattaccggcgaaaccaaagatcaggtggcgaacagcgcgtttgtggaacgcctgcgcaaacatggcctggaagtgatttatatgattgaaccgattgatgaatattgcgtgcagcagctgaaagaatttgaaggcaaaaccctggtgagcgtgaccaaagaaggcctggaactgccggaagatgaagaagaaaaaaaaaaacaggaagaaaaaaaaaccaaatttgaaaacctgtgcaaaattatgaaagatattctggaaaaaaaagtggaaaaagtggtggtgagcaaccgcctggtgaccagcccgtgctgcattgtgaccagcacctatggctggaccgcgaacatggaacgcattatgaaagcgcaggcgctgcgcgataacagcaccatgggctatatggcggcgaaaaaacatctggaaattaacccggatcatagcattattgaaaccctgcgccagaaagcggaagcggataaaaacgataaaagcgtgaaagatctggtgattctgctgtatgaaaccgcgctgctgagcagcggctttagcctggaagatccgcagacccatgcgaaccgcatttatcgcatgattaaactgggcctgggcattgatgaagatgatccgaccgcggatgataccagcgcggcggtgaccgaagaaatgccgccgctggaaggcgatgatgataccagccgcatggaagaagtggat
- 3.3. Codon optimization.
ATGCCGGAAGAAACCCAGACCCAGGATCAGCCGATGGAAGAAGAAGAAGTGGAAACCTTTGCGTTTCAGGCAGAAATTGCGCAGCTGATGTCTCTGATTATTAATACCTTTTATAGCAATAAAGAAATCTTCCTGCGTGAACTGATTAGCAACAGCAGCGATGCACTGGATAAAATTCGCTATGAATCGCTGACCGATCCGAGCAAACTGGATAGCGGCAAAGAACTGCATATTAATCTGATTCCGAACAAACAGGATCGCACCCTGACCATTGTGGATACCGGCATTGGCATGACCAAAGCGGATCTGATTAATAATCTGGGCACCATTGCCAAATCGGGCACCAAAGCCTTTATGGAAGCCCTGCAGGCCGGCGCGGATATTAGCATGATTGGCCAGTTCGGCGTGGGTTTCTATAGCGCCTATCTGGTGGCCGAAAAAGTGACCGTTATCACCAAACATAATGATGATGAACAGTATGCGTGGGAAAGCTCCGCGGGCGGCAGCTTTACCGTGCGCACCGATACCGGCGAACCGATGGGCCGCGGCACGAAAGTTATTCTGCACCTGAAAGAAGATCAGACCGAGTACTTAGAAGAACGTCGTATTAAAGAAATTGTGAAAAAACATAGCCAGTTCATCGGCTATCCGATCACCCTGTTCGTGGAAAAAGAACGCGATAAAGAAGTTAGCGATGATGAAGCGGAAGAAAAAGAAGATAAAGAAGAAGAAAAAGAGAAGGAAGAAAAAGAGAGCGAAGATAAACCGGAAATTGAAGATGTGGGCTCGGATGAAGAAGAAGAAAAAAAAGATGGCGATAAAAAAAAGAAAAAAAAAATTAAAGAAAAATACATTGATCAGGAAGAACTGAATAAGACCAAACCGATTTGGACCCGTAACCCGGATGACATTACCAACGAGGAATATGGCGAATTTTATAAAAGCCTGACCAACGATTGGGAAGATCACCTGGCGGTTAAACATTTTAGCGTGGAAGGCCAGCTGGAATTTCGCGCGCTGCTGTTCGTACCGCGCCGCGCCCCGTTTGATCTGTTTGAAAATCGCAAAAAAAAAAACAATATTAAACTGTATGTTCGCCGCGTCTTCATTATGGATAATTGCGAAGAACTGATTCCGGAATACCTGAACTTTATTCGCGGCGTGGTTGATAGCGAAGATCTGCCGCTGAACATTAGCCGCGAAATGCTGCAGCAGAGCAAAATTCTGAAAGTGATTCGCAAAAACCTGGTAAAAAAATGCCTGGAACTGTTTACCGAACTGGCGGAAGATAAAGAAAATTATAAAAAGTTTTATGAACAGTTTAGCAAAAACATTAAACTGGGCATTCATGAGGATAGCCAAAATCGTAAGAAACTGAGCGAACTGCTGCGCTACTATACCTCGGCGAGCGGCGATGAAATGGTGAGCCTGAAAGATTACTGTACCCGTATGAAAGAGAATCAGAAACATATTTATTACATCACCGGCGAAACTAAAGATCAGGTGGCGAATAGCGCTTTTGTGGAACGTCTGCGTAAACACGGCCTGGAAGTGATTTACATGATTGAACCGATTGATGAATATTGCGTGCAGCAGCTGAAAGAATTTGAAGGTAAAACCCTGGTTAGCGTTACCAAAGAAGGCCTGGAATTACCGGAAGATGAAGAAGAAAAAAAAAAACAGGAAGAAAAAAAAACCAAATTTGAAAATCTGTGCAAAATTATGAAAGATATTCTGGAAAAAAAAGTGGAAAAAGTGGTGGTCAGCAATCGCCTGGTGACCAGCCCGTGCTGCATTGTGACCAGCACCTACGGCTGGACCGCGAATATGGAACGTATTATGAAAGCGCAGGCCCTGCGCGACAATAGCACCATGGGCTACATGGCCGCGAAAAAACACCTGGAAATTAACCCGGATCACAGCATTATTGAAACCCTGCGTCAGAAAGCGGAAGCGGATAAAAACGATAAATCGGTTAAAGATCTGGTGATTCTGCTGTATGAAACCGCGCTGCTGAGCAGCGGCTTTAGCCTGGAAGATCCGCAGACCCATGCGAATCGCATTTATCGCATGATTAAACTGGGACTGGGTATTGATGAAGATGATCCGACCGCGGATGATACCAGTGCGGCGGTTACCGAAGAAATGCCGCCGCTGGAAGGCGATGATGACACCAGCCGCATGGAGGAAGTGGAT
- 3.4. You have a sequence! Now what?
What technologies could be used to produce this protein from your DNA? Describe in your words the DNA sequence can be transcribed and translated into your protein. You may describe either cell-dependent or cell-free methods, or both.
R/:Once I have the DNA sequence, I would use a cell-free expression system to produce the protein. In this approach, the DNA is added directly to a reaction mixture that contains all the molecular components required for transcription and translation, such as RNA polymerase, ribosomes, nucleotides, amino acids, tRNAs, and necessary enzymes.
First, the DNA sequence is transcribed into mRNA by RNA polymerase within the reaction mixture. Then, ribosomes bind to the mRNA and translate it into the corresponding amino acid sequence, forming the protein. Because this process occurs in vitro (outside of living cells), it allows rapid protein production without the need for cell transformation, growth, or cloning into a host organism.
Cell-free systems are particularly useful for fast prototyping, testing gene constructs, or producing proteins that may be toxic to living cells. The entire process still follows the central dogma (DNA → RNA → Protein), but it happens in a controlled biochemical environment rather than inside a living cell.
- 3.5. How does it work in nature/biological systems?
R/:In nature, gene expression follows the central dogma of molecular biology. First, the DNA sequence of a gene is transcribed into messenger RNA (mRNA) by RNA polymerase. In eukaryotic cells, the initial transcript (pre-mRNA) undergoes processing, including 5’ capping, polyadenylation, and splicing to remove introns. Once mature mRNA is formed, it is transported to the cytoplasm, where ribosomes bind to it and translate the nucleotide sequence into an amino acid chain. Transfer RNAs (tRNAs) bring the appropriate amino acids according to codon–anticodon pairing. The resulting polypeptide then folds into its functional three-dimensional structure, sometimes with the help of molecular chaperones. This tightly regulated process ensures that proteins are produced at the right time, in the right amount, and in the appropriate cellular context.
- Describe how a single gene codes for multiple proteins at the transcriptional level.
R/:A single gene can produce multiple proteins at the transcriptional level mainly through alternative splicing. In eukaryotic cells, genes are composed of exons and introns. After transcription, the initial RNA transcript (pre-mRNA) contains both regions. During RNA processing, introns are removed, and exons are joined together. However, the cell does not always combine exons in the same way. Different exons can be included or excluded, generating distinct mRNA transcripts from the same gene. These different mRNAs are then translated into different protein isoforms. This mechanism increases protein diversity without increasing the total number of genes in the genome.
- Try aligning the DNA sequence, the transcribed RNA, and also the resulting translated Protein.

Part 4: Prepare a Twist DNA Synthesis Order
4.2. Build Your DNA Insert Sequence
Benchling ink: https://benchling.com/s/seq-52XZoIYMGmiV8rSoyWAZ?m=slm-6exDNpn6ZWIEA6nOots7
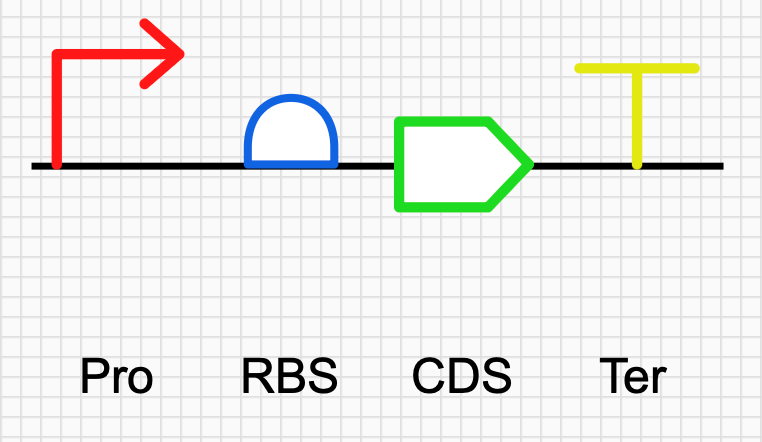
4.3 to 4.6

Part 5: DNA Read/Write/Edit
- 5.1 DNA Read
(i) What DNA would you want to sequence (e.g., read) and why? This could be DNA related to human health (e.g. genes related to disease research), environmental monitoring (e.g., sewage waste water, biodiversity analysis), and beyond (e.g. DNA data storage, biobank).
R/:I would like to sequence DNA from gastrointestinal stromal tumor (GIST) samples, specifically focusing on mutations in the KIT and PDGFRA genes. Mainly because KIT mutations cause the receptor to be constitutively activated, meaning it continuously signals cell proliferation even without an external signal. This is why sequencing this gene is important, understanding the specific mutation helps select the most effective existing treatment, such as tyrosine kinase inhibitors like imatinib, for each specific patient. Additionally, since mutations can vary between patients, sequencing improves the precision of medicine by allowing more personalized and targeted therapies.
(ii) In lecture, a variety of sequencing technologies were mentioned. What technology or technologies would you use to perform sequencing on your DNA and why? R/: I would use Illumina Also answer the following questions:
Is your method first-, second- or third-generation or other? How so? R/: It’s second generation because it uses massively parallel sequencing, reading millions of fragments simultaneously, unlike Sanger which sequences one fragment at a time. What is your input? How do you prepare your input (e.g. fragmentation, adapter ligation, PCR)? List the essential steps. R/: The input it’s DNA from GIST samples.The indispensable steps are first fragmentation because we have a long DNA and illumin reads short fragments, after we do adapter ligation so the machine recognize each fragment and lastly a PCR amplification so we have enough material to read. What are the essential steps of your chosen sequencing technology, how does it decode the bases of your DNA sample (base calling)? R/:After preparation, the fragments are attached to a flow cell where cluster generation occurs each fragment is copied thousands of times to amplify the signal. Then, sequencing by synthesis begins: fluorescently labeled nucleotides are added one at a time, each base (A, T, C, G) emitting a different color. A camera captures an image after each incorporation, and the computer translates each color into a base this is called base calling. This process repeats for each position along the fragment What is the output of your chosen sequencing technology? R/:The output of Illumina sequencing is millions of short reads sequences of bases (A, T, C, G) typically 150-300 base pairs long. These reads are then aligned to a reference genome using bioinformatics tools to identify mutations in the KIT gene, such as substitutions or insertions/deletions, that may be driving GIST.
- 5.2 DNA Write
(i) What DNA would you want to synthesize (e.g., write) and why? These could be individual genes, clusters of genes or genetic circuits, whole genomes, and beyond. As described in class thus far, applications could range from therapeutics and drug discovery (e.g., mRNA vaccines and therapies) to novel biomaterials (e.g. structural proteins), to sensors (e.g., genetic circuits for sensing and responding to inflammation, environmental stimuli, etc.), to art (DNA origamis). If possible, include the specific genetic sequence(s) of what you would like to synthesize! You will have the opportunity to actually have Twist synthesize these DNA constructs! :)
R/: I would like to synthesize a genetic biosensor circuit designed to detect early increases in HSP90 dependence in KIT-mutant GIST cells. The biosensor would consist of a synthetic DNA sequence that, when inserted into tumor cells, monitors HSP90 activity levels and produces a detectable signal (such as a fluorescent protein) when levels rise above a threshold. This is clinically relevant because increased HSP90 dependence is an early indicator of cellular adaptation preceding imatinib resistance, meaning the biosensor could alert clinicians before full resistance develops, allowing earlier treatment adjustments. The specific sequence would include a HSP90-responsive promoter driving a reporter gene, designed to activate transcription proportionally to HSP90 activity levels in KIT-mutant cells.
(ii) What technology or technologies would you use to perform this DNA synthesis and why? R/:For synthesizing my biosensor genetic circuit, I would use array-based phosphoramidite synthesis, as offered by Twist Bioscience. This technology is ideal because it allows precise, high-throughput synthesis of specific designed sequences at relatively low cost, without needing a biological template.
Also answer the following questions:
What are the essential steps of your chosen sequencing methods? R/:The essential steps are: (1) the DNA sequence is designed computationally; (2) synthesis occurs on a chip where nucleotides are added one at a time chemically to growing DNA chains; (3) each nucleotide is protected by a chemical group that is removed before the next base is added, allowing controlled sequential addition; (4) the completed sequences are cleaved from the chip and assembled into longer constructs if needed through Gibson assembly or similar methods.
What are the limitations of your sequencing method (if any) in terms of speed, accuracy, scalability? R/:The main limitations are accuracy, errors can accumulate as the sequence gets longer, making synthesis of very long sequences challenging. Additionally, longer sequences are more expensive and slower to produce. Array-based synthesis improves scalability but error rates remain a concern, often requiring sequence verification after synthesis using Sanger or Illumina sequencing.
- 5.3 DNA Edit
(i) What DNA would you want to edit and why? In class, George shared a variety of ways to edit the genes and genomes of humans and other organisms. Such DNA editing technologies have profound implications for human health, development, and even human longevity and human augmentation. DNA editing is also already commonly leveraged for flora and fauna, for example in nature conservation efforts, (animal/plant restoration, de-extinction), or in agriculture (e.g. plant breeding, nitrogen fixation). What kinds of edits might you want to make to DNA (e.g., human genomes and beyond) and why?
R/:I would want to edit the KIT gene in GIST tumor cells using CRISPR-Cas9 technology. Specifically, I would correct the activating mutations in KIT that cause constitutive activation of the tyrosine kinase receptor, driving uncontrolled cell proliferation. By editing the mutated sequence back to its wildtype form, the receptor would only signal when appropriate, stopping uncontrolled tumor growth at its source rather than just blocking it pharmacologically as imatinib does. This approach is particularly compelling because imatinib resistance is a major clinical challenge in GIST patients eventually develop secondary mutations that render the drug ineffective. A CRISPR-based correction of the primary KIT mutation could offer a more permanent solution, potentially eliminating the tumor’s ability to develop resistance. Additionally, since different patients carry different KIT mutations, CRISPR could be personalized to target each patient’s specific mutation, aligning with the precision medicine approach.
(ii) What technology or technologies would you use to perform these DNA edits and why? Also answer the following questions:
How does your technology of choice edit DNA? What are the essential steps?
R/:I would use CRISPR-Cas9 to correct the activating KIT mutations in GIST tumor cells, as it allows precise, targeted editing of specific DNA sequences and can be personalized for each patient’s mutation. CRISPR-Cas9 edits DNA through the following steps: (1) the guide RNA (gRNA) directs the Cas9 protein to the specific mutated KIT sequence; (2) Cas9 creates a double-strand break at that exact location; (3) a correct DNA template is provided and the cell repairs the break using homology-directed repair (HDR), replacing the mutated sequence with the corrected wildtype sequence.
What preparation do you need to do (e.g. design steps) and what is the input (e.g. DNA template, enzymes, plasmids, primers, guides, cells) for the editing?
R/:The preparation steps include: (1) designing a gRNA that matches the specific KIT mutation of the patient — this is done computationally; (2) preparing the Cas9 protein or encoding it in a plasmid; (3) preparing a HDR template containing the correct wildtype KIT sequence; (4) delivering all components into the tumor cells via a viral vector or nanoparticles. The inputs are therefore: the gRNA, Cas9 enzyme, HDR DNA template, and the target tumor cells.
What are the limitations of your editing methods (if any) in terms of efficiency or precision?
R/:The main limitations are: (1) off-target edits Cas9 may cut unintended genomic locations causing unwanted mutations; (2) low HDR efficiency, especially in non-dividing cells; (3) delivery challenges getting CRISPR machinery efficiently into tumor cells in vivo remains technically difficult; (4) since KIT mutations vary between patients, a new gRNA must be designed for each case, making large-scale application complex.