week-10-hw-imaging-and-measurement
Homework: Final Project
Please identify at least one (ideally many) aspect(s) of your project that you will measure.
R/: I would like to measure the expression levels of mutant p53 protein across different cancer types. High accumulation of mutant p53 (due to MDM2 being unable to degrade it) would indicate a GOF mutation, making those cancers more susceptible to p53 GOF-driven oncogenesis.
Please describe all of the elements you would like to measure and how you will perform these measurements.
R/:
First, I will perform DNA sequencing to confirm the specific GOF mutation in TP53.
Then, I will use Western Blot (SDS-PAGE + anti-p53 antibody detection) to quantify p53 protein levels across cancer samples.
Additionally, mass spectrometry (LC-MS) will be used to confirm the exact mutant protein sequence and molecular weight.
Homework: Waters Part I — Molecular Weight
We will analyze an eGFP standard on a Waters Xevo G3 QTof MS system.
1. Theoretical Molecular Weight
eGFP Sequence:
MVSKGEELFTG VVPILVELDG DVNGHKFSVS GEGEGDATYG KLTLKFICTT GKLPVPWPTL VTTLTYGVQC FSRYPDHMKQ HDFFKSAMPE GYVQERTIFF KDDGNYKTRA EVKFEGDTLV NRIELKGIDF KEDGNILGHK LEYNYNSHNV YIMADKQKNG IKVNFKIRHN IEDGSVQLAD HYQQNTPIGD GPVLLPDNHY LSTQSALSKD PNEKRDHMVL LEFVTAAGIT LGMDELYKLE HHHHHH
Note: Contains His-tag (HHHHHH) and linker (LE).
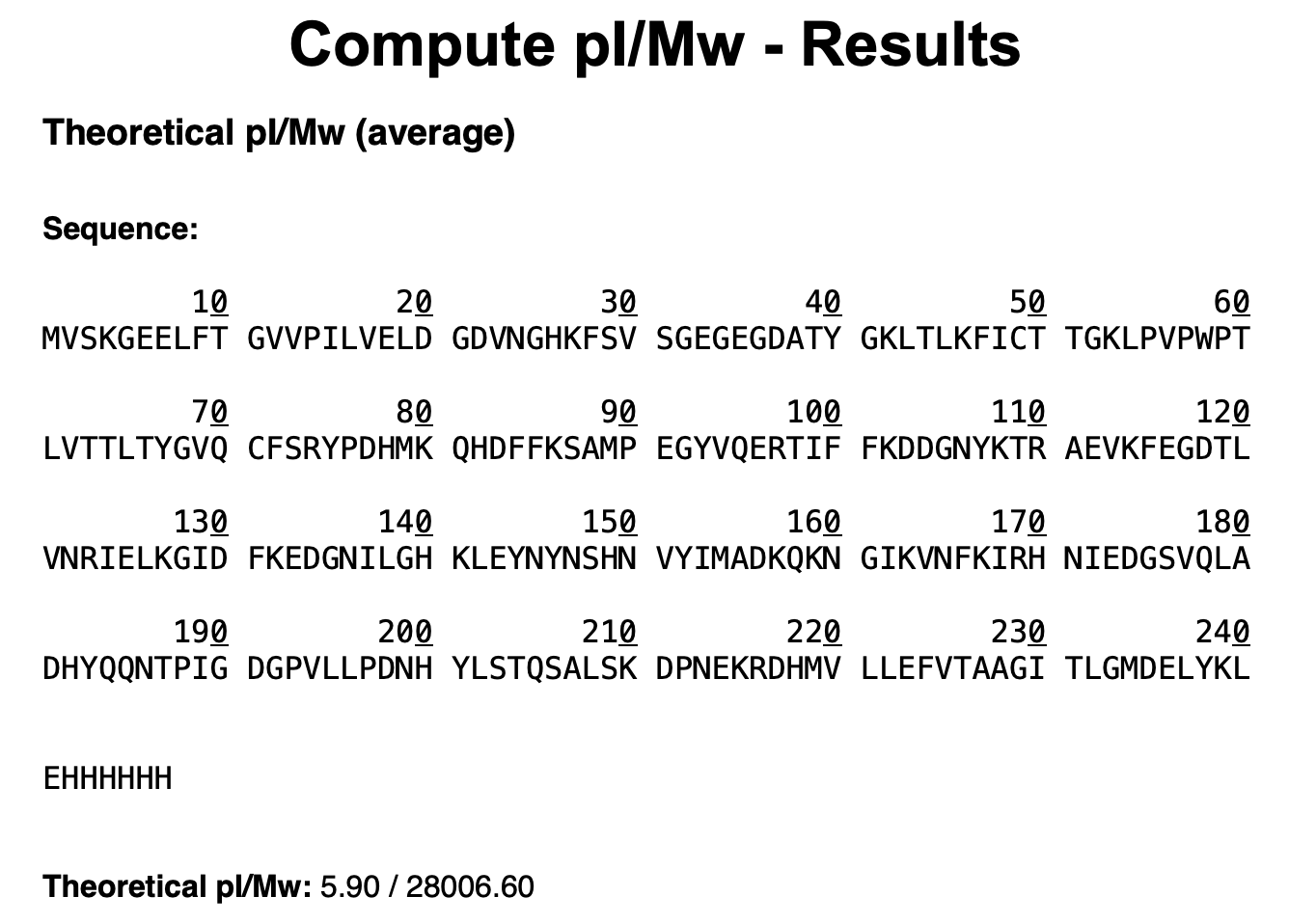
R/: Based on ExPASy Compute pI/Mw, the theoretical molecular weight of eGFP is 28,006.60 Da (~28 kDa).
2. Molecular Weight from Charge States
Calculate MW using adjacent charge states from LC-MS (Figure 1).
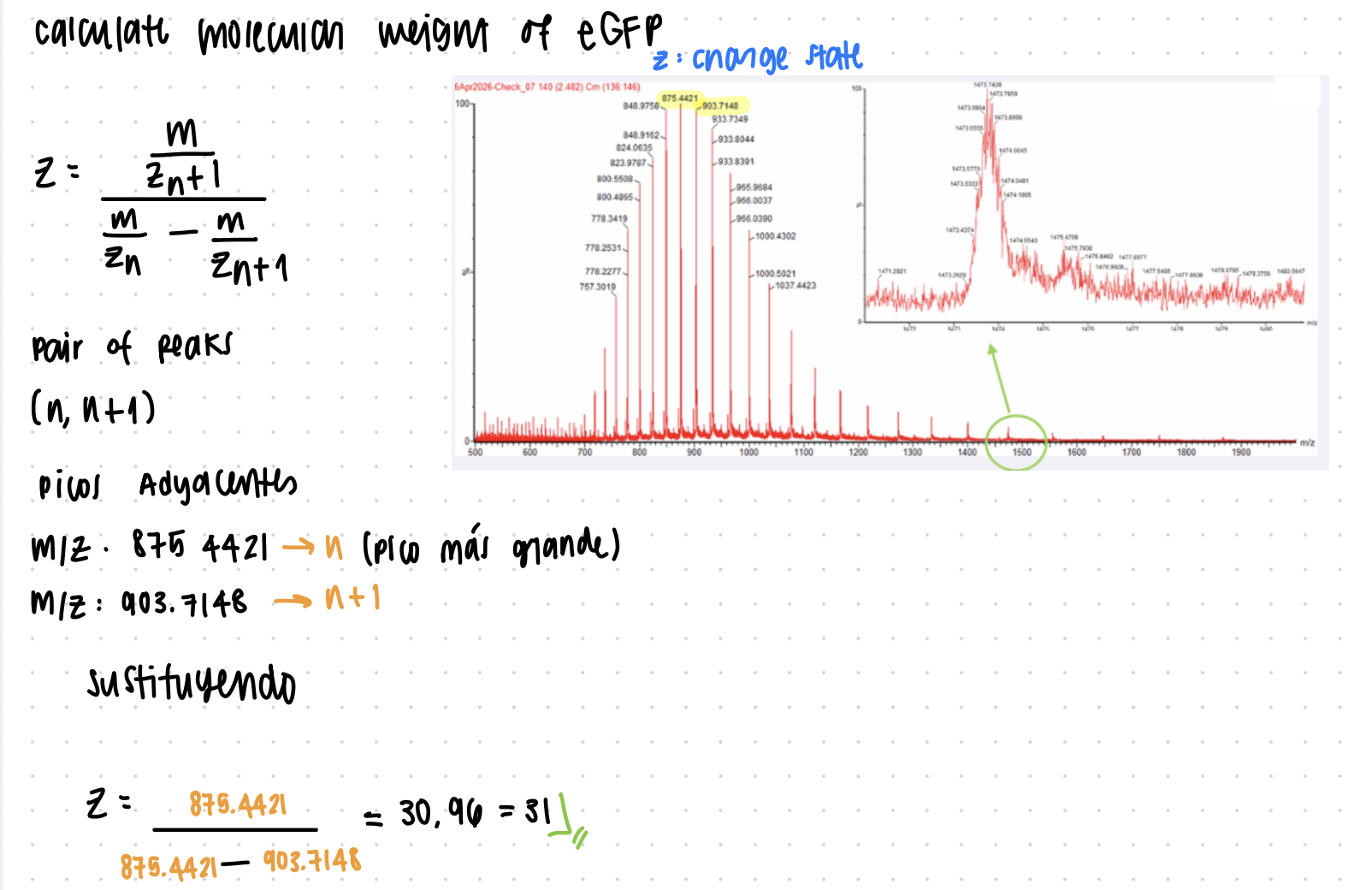
3. Accuracy Calculation
Compare deconvoluted MW vs theoretical MW.

Homework: Waters Part II — Secondary/Tertiary Structure
1. Native vs Denatured Proteins
R/:
Native proteins maintain their 3D folded structure, while denatured proteins unfold and expose more basic residues (Lys, Arg), allowing higher protonation.
In MS:
- Denatured proteins: many peaks at low m/z (high charge states)
- Native proteins: fewer peaks at high m/z (low charge states)
2. Charge State Determination
R/:
The peak at ~2800 m/z corresponds to z = 10
Calculation:
z = 28,000 / 2,799.4199 ≈ 10
This low charge state is consistent with a folded (native) protein.
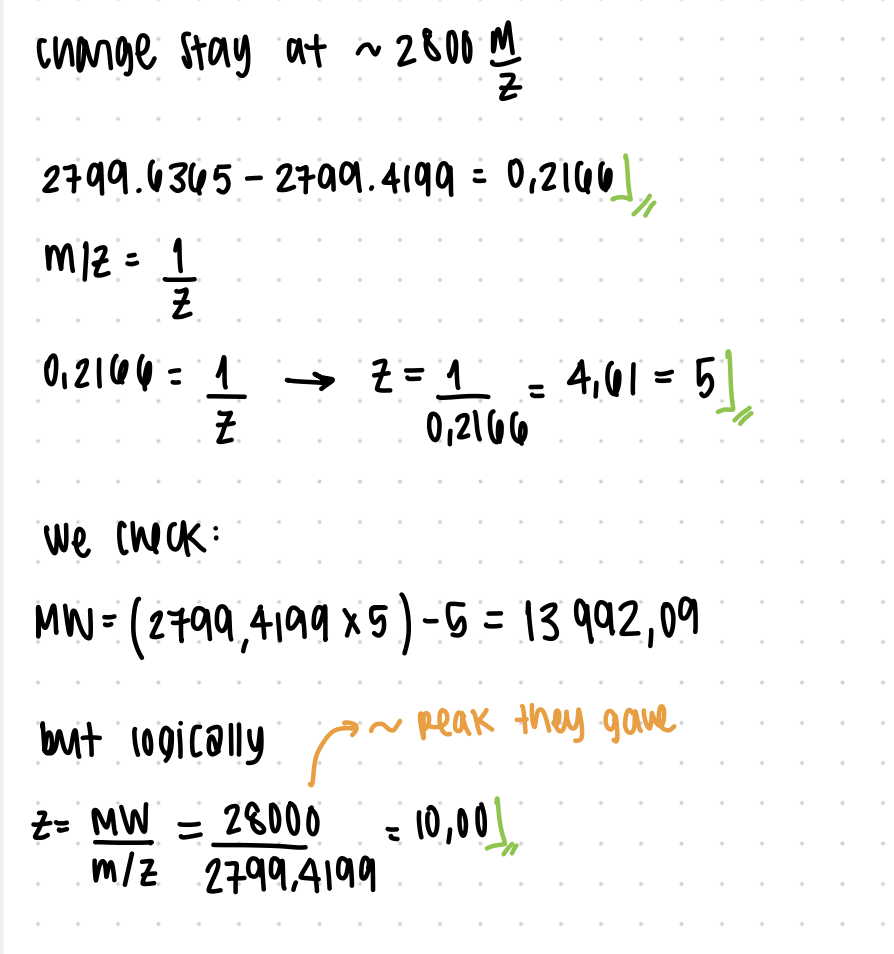
Homework: Waters Part III — Peptide Mapping (Primary Structure)
We digest eGFP with trypsin (cuts after K and R).
1. Lysine and Arginine Count
R/:
eGFP contains:
- 20 Lysines (K)
- 6 Arginines (R)
- Total = 26 cleavage sites
2. Number of Peptides
Steps:
- Use ExPASy PeptideMass
- Input sequence
- Perform trypsin digestion
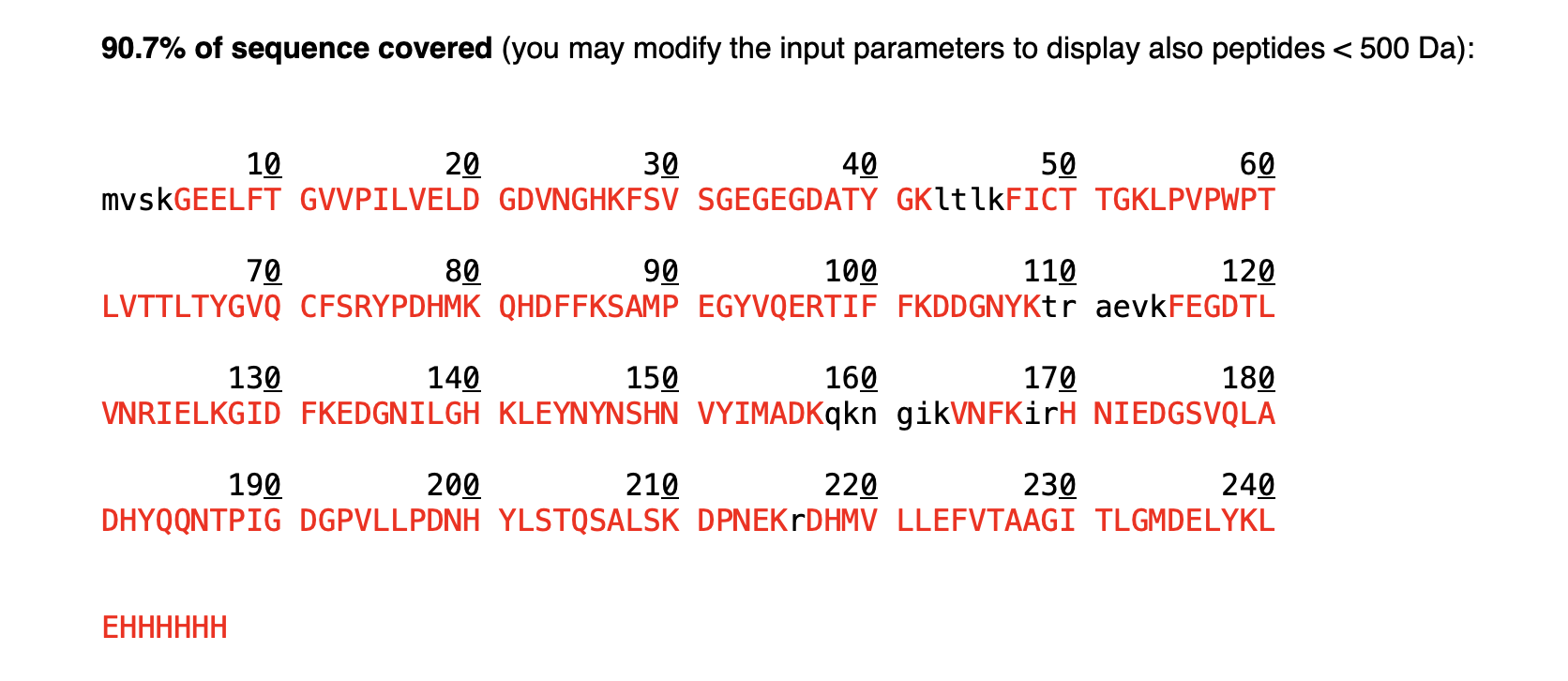

R/:
Tryptic digestion of eGFP generates 25 peptides.
Based on the LC-MS data for the Peptide Map data generated in lab (please use Figure 5a as a reference) how many chromatographic peaks do you see in the eGFP peptide map between 0.5 and 6 minutes? You may count all peaks that are >10% relative abundance. Figure 5a. Total ion chromatogram (TIC) of the eGFP peptide map. The peak at 2.78 minutes is circled, and its MS data is shown in the mass spectrum in Figure 5b, below. Figure 5a. Total ion chromatogram (TIC) of the eGFP peptide map. The peak at 2.78 minutes is circled, and its MS data is shown in the mass spectrum in Figure 5b, below.
Assuming all the peaks are peptides, does the number of peaks match the number of peptides predicted from question 2 above? Are there more peaks in the chromatogram or fewer?
R/: The number of peaks does not match. There are fewer peaks (22) in the chromatogram than peptides predicted (25). This could be explained by the size of some peptides — very small peptides (<500 Da) are not retained well in the column and therefore are not visible as distinct peaks in the chromatogram.
HASTA AQUI QUEDE 5. Identify the mass-to-charge ( ) of the peptide shown in Figure 5b. What is the charge () of the most abundant charge state of the peptide (use the separation of the isotopes to determine the charge state). Calculate the mass of the singly charged form of the peptide () based on its and .