Week 2 HW: DNA read, write and edit.
Part I: Benchling & In-silico Gel Art
Begin by importing your DNA sequence and use the Digests tool to test the effects of different restriction enzyme(s). Export your final design as a png and compare with your lab results on your Notion page. See the images below for where to find the Digests tool, selecting the “NEB 2-log” ladder in the Virtual Digest tab, and how to have multiple Digests appear in the same Virtual Digest.
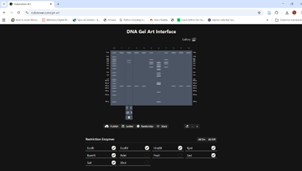
First, I accessed the website https://rcdonovan.com/gel-art to create a sketch of a design based on the lambda sequence with the restriction enzymes:
- EcoRI-HF
- HindIII-HF
- BamHI-HF
- KpnI-HF
- EcoRV-HF
- SacI-HF
- SalI-HF
which I will perform using electrophoresis.
After experimenting a bit with the enzymes and the sequence (lamba), I decided to draw a “sunrise in a forest”.
Electrophoresis Overview
Now let’s talk about electrophoresis. We start with a lambda DNA sequence of about 48,500 base pairs, which we cut into fragments of different sizes using restriction enzymes. During electrophoresis, these DNA fragments are loaded into wells in a gel and move through the gel when an electric current is applied.
The fragments travel at different speeds depending on their size:
- Larger fragments move more slowly and cover shorter distances.
- Smaller fragments move faster and travel farther.
By observing how far each fragment migrates, we can use this pattern to create the chosen visual representation.
Benchling Simulation
Let’s go to the Benchling website (benchling.com) to perform a simulation of electrophoresis using the proposed DNA sequence and restriction enzymes.
To find the GenBank accession number for the lambda sequence (#lambda sequence in GenBank) Google was used. The number for the lambda is J02459.1. Clicking on this number opens the sequence directly on the NIH website (National Library of Medicine, www.ncbi.nlm.nih.gov).
Click on FASTA, which will open a menu. Adjust the options as needed and then download the file.
Now, open https://benchling.com/ create a project and import the sequence.
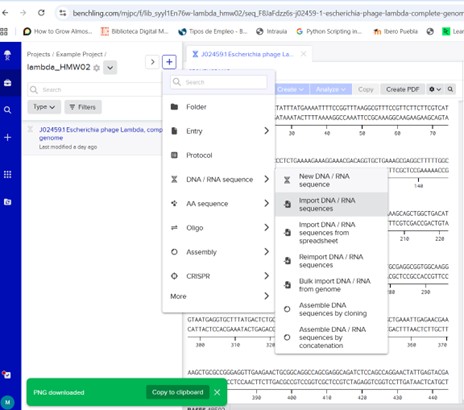

Next, select the scissors icon on the right to perform a digest in the Virtual Digest tab. Choose the restriction enzyme and carry out the digestion.
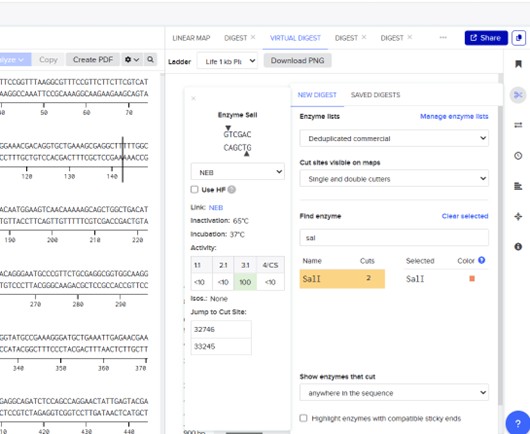

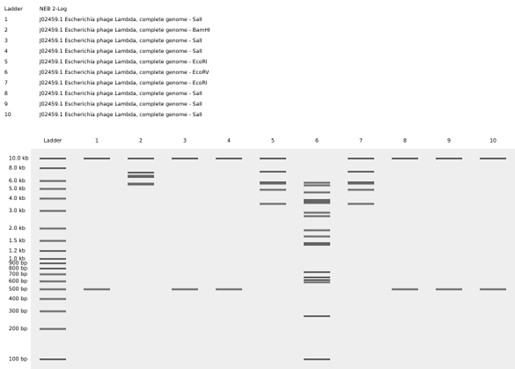
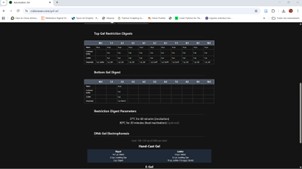
The program will simulate electrophoresis for each well, showing how far the DNA fragments migrate. After performing digests for all nine wells and selecting the ladder, we will have the final proposed image.
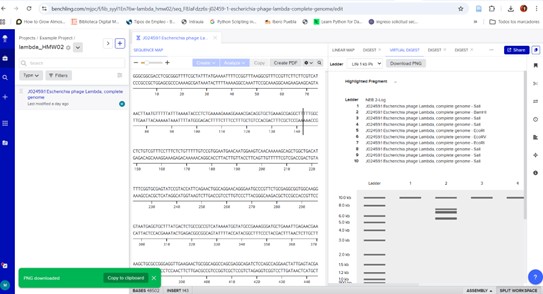

Figure: “Sunrise in the Forest,” sun and tree in the image.
Part 2: Gel Art – Restriction digests and Gel Electrophoresis
Since I did not have access to the lab to perform this section, I will use a gel that I ran a couple of weeks ago:
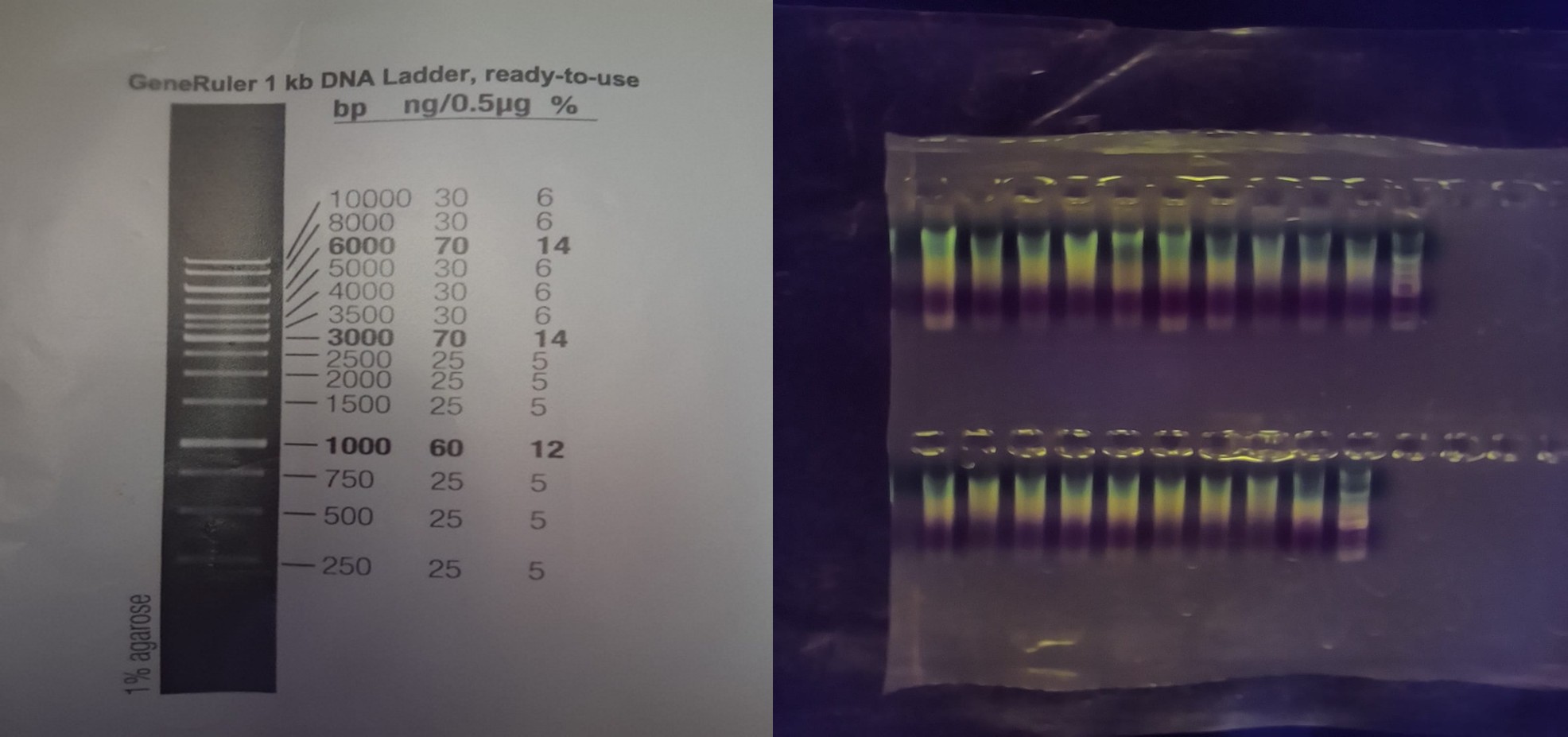

To determine the fragment size, we compared it with the ladder (on the right of each row).
The fragments are approximately 6,000 bp.
The color does not determine the size. The DNA size is determined by the migration distance.
The bands are clear and well separated, indicating that the electrophoresis ran correctly. No lateral smearing or fuzzy bands are observed, which indicates good gel preparation and proper sample loading.
Part 3: DNA Design Challenge (proposal)
Promoter
- Name: BBa_J23106
- Type: Constitutive promoter
- Sequence: TTTACGGCTAGCTCAGTCCTAGGTATAGTGCTAGC
- Purpose: Initiates transcription of PETase.
RBS + Start codon (ATG)
- Sequence: CATTAAAGAGGAGAAAGGTACCATG
- Purpose: Ribosome binding and translation initiation.
CDS (PETase codon-optimized)
- Name: PETase_Ecoli_codopt
- Sequence: tu secuencia codón optimizada aquí
- Purpose: Codes for the PETase protein.
His-tag (optional, C-terminal)
- Sequence: CATCACCATCACCATCATCAC (7x His)
- Purpose: Facilitates purification of PETase protein.
Stop codon
- Sequence: TAA
- Purpose: Terminates translation.
Terminator
- Example: BBa_B0015
- Sequence: CCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATA
- Purpose: Stops transcription and stabilizes mRNA.
Antibiotic resistance / selectable marker
- Example: Ampicillin (AmpR)
- Purpose: Selects for bacteria that successfully took up the plasmid.
3.1. Choose your protein
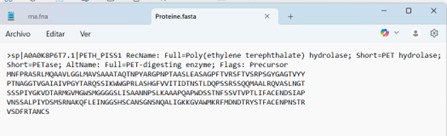
PETase Protein Sequence Description
The protein used in this project is a Poly(ethylene terephthalate) hydrolase (PETase), also known as a PET-digesting enzyme. Its UniProt/NCBI accession is A0A0K8P6T7.1, and it is composed of 290 amino acids. This enzyme originates from a bacterial source and is capable of degrading PET plastic. The sequence has been selected for codon optimization to enable expression in E. coli, facilitating the construction of a plasmid-based reporter system for PET detection in environmental samples.
The PETase protein sequence was obtained from Ideonella sakaiensis by accessing the NIH protein database at https://www.ncbi.nlm.nih.gov/protein and searching for “PETase Ideonella sakaiensis.” The protein entry was selected, and the sequence was downloaded in FASTA format for subsequent use in codon optimization and plasmid design.


I downloaded the Proteine.fasta file.


I examined and visualized the features of the protein sequence through the Features option.
3.2. Reverse Translate: Protein (amino acid) sequence to DNA (nucleotide) sequence.
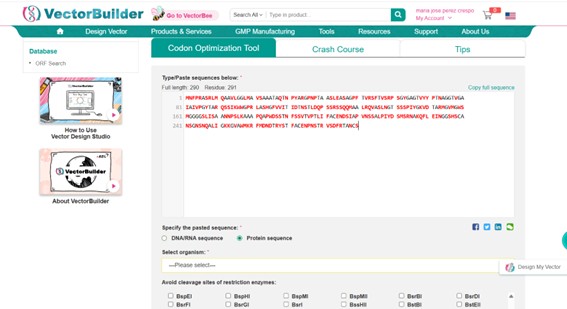
There are available on internet many Codon Optimization Tools (IDT, Twist, GenScript, VectorBuilder, ExoOptimizer…). I chose Vector Builder Optimization Tool (https://en.vectorbuilder.com/).
The PETase protein sequence in FASTA format was used as input in
VectorBuilder → Tools → Codon Optimization Tool.
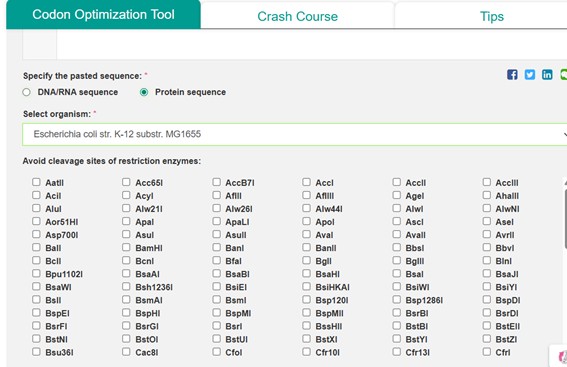
- The amino acid sequence was pasted directly, specifying it as a protein sequence.
- The host organism was set to Escherichia coli str. K-12 substr. MG1655 to optimize codon usage for this bacterium.
- No restriction sites were selected for avoidance.
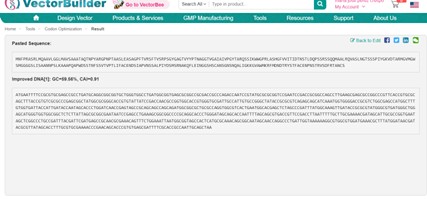
The tool generated a DNA sequence (ORF) optimized for E. coli, with the following characteristics:
- Start codon: ATG and stop codon: TAA
- GC content: 59.56%
- Codon Adaptation Index (CAI): 0.91, indicating high compatibility with E. coli codon preferences.
- The sequence is now ready to be inserted into a plasmid for expression.
3.3. Codon optimization
NOTE: Codon optimization was performed along with reverse translation in Vector Builder (VB).
Codon optimization is important because, although multiple codons can encode the same amino acid, different organisms preferentially use certain codons more frequently. If a gene contains codons that are rarely used in the host organism, translation may be inefficient, leading to low protein expression. By optimizing codon usage, I adapted the PETase nucleotide sequence to match the codon preferences of Escherichia coli without altering the amino acid sequence of the protein.
For this project, I chose to optimize PETase for E. coli because it is the host organism where the plasmid will be expressed. Since PETase originates from the bacterium Ideonella sakaiensis, its native codon usage differs from E. coli, so codon optimization improves expression efficiency in this bacterial system.
During codon optimization, restriction enzyme recognition sites were not specifically removed because the sequence was designed for direct insertion into a plasmid using VectorBuilder, which ensures proper assembly. This optimization ensures high expression of PETase in E. coli, allowing the gene to be used effectively in the plasmid as a biological marker for PET detection.
3.4. You have a sequence! Now what?
Producing PETase from the Optimized DNA Sequence
The codon-optimized DNA sequence for PETase can be expressed using cell-dependent or cell-free systems. In bacteria like E. coli, the plasmid is transcribed into mRNA and translated into protein using the host’s machinery, with selection markers ensuring only cells carrying the gene produce PETase. Alternatively, cell-free systems synthesize protein directly from the DNA in vitro, without living cells. Optimizing the sequence ensures efficient production of functional PETase for detection or degradation of PET.
Both approaches follow the central dogma of molecular biology: DNA is transcribed into RNA, and RNA is translated into protein.
3.5. How does it work in nature/biological systems?
In nature, a single gene contains the information to produce a protein through transcription and translation. The DNA sequence of the PETase gene is first transcribed into messenger RNA (mRNA) by RNA polymerase. This mRNA serves as a template for translation, where ribosomes read the codons in sets of three nucleotides to assemble the corresponding amino acids into the PETase protein.
Some genes in biological systems can produce multiple protein variants through mechanisms such as alternative splicing, alternative start codons, or frameshifting, but PETase from Ideonella sakaiensis produces a single functional protein. Aligning the DNA sequence, the transcribed RNA, and the translated protein shows how the nucleotide code directly determines the amino acid sequence of the protein.
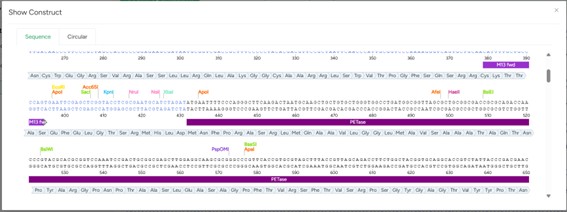

The construct of DNA and amino acids was obtained in genscript during codon optimization sequence.
Part 4: Prepare a Twist DNA Synthesis Order
4.1. Create a Twist account and a Benchling account
Twist account doesn’t work properly. I used instead VectorBuilder and Benchling.
4.2. Build a DNA Insert Sequence
I adjusted the proposed PETase plasmid design as follows:
I have to build a DNA Insert Sequence from the Promoter to Terminator, which includes:
- Promoter – initiates transcription of the gene. (BBa_J23106 TTTACGGCTAGCTCAGTCCTAGGTATAGTGCTAGC, position 1-35).
- Ribosome Binding Site (RBS / Kozak) – ensures proper ribosome binding and efficient translation initiation (Kozak GCCACC, position 36 - 41).
- Start Codon – defines the beginning of the open reading frame (ATG, position 42 - 44).
- Codon-Optimized PETase ORF – DNA codon optimized sequence for E. coli (position 45 – 911).
- Stop Codon – properly terminates translation (TAA, position 912 - 914).
- Transcription Terminator – ensures transcription stops correctly (BBa_B0015 CCAGGCATCAAATAAAACGAAAGGCTCAGTCGAAAGACTGGGCCTTTCGTTTTATCTGTTGTTTGTCGGTGAACGCTCTCTACTAGAGTCACACTGGCTCACCTTCGGGTGGGCCTTTCTGCGTTTATA).
- Antibiotic Selection Marker – allows selection of bacteria carrying the plasmid (Ampicillin - Amp).
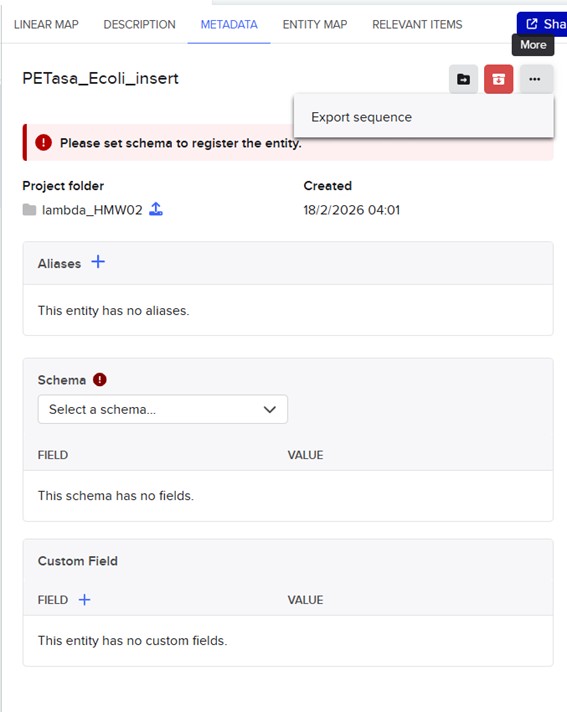
In Benchling, I create a new sequence that will result from sequentially adding the DNA corresponding to the proposed plasmid design. I am going to create an insert sequence manually.
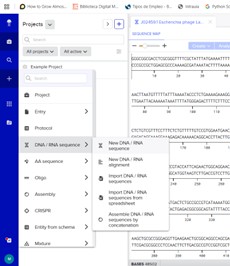

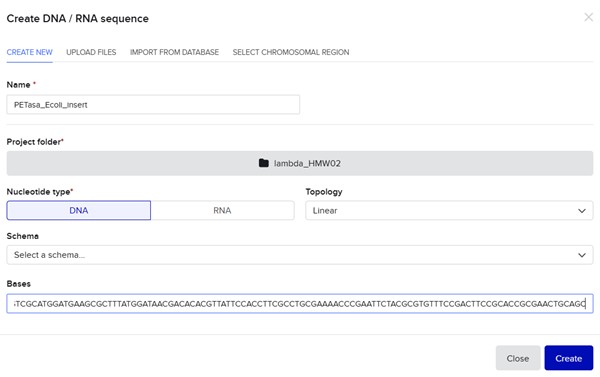

Then click Sequence Map and right-clicking over the sequence and create an annotation that describes what each piece is.


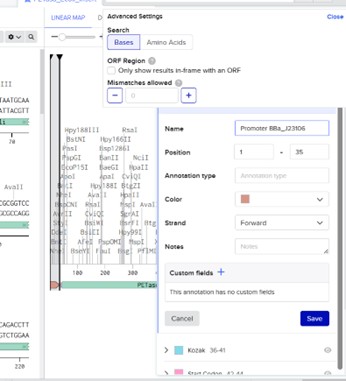

This final construct ensures correct transcription and translation of the PETase protein in a bacterial expression system.
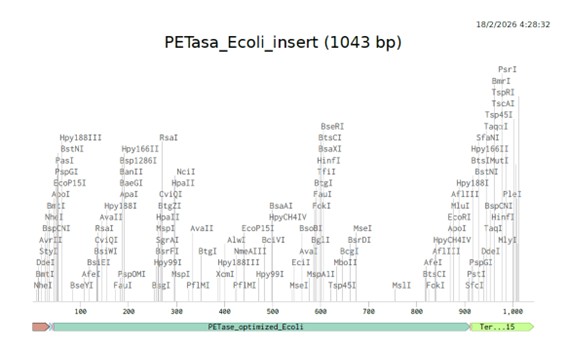
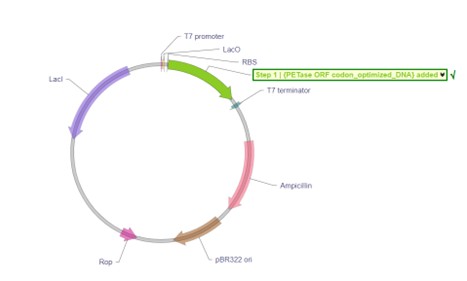
I clicked on Linear Map, to preview the sequence (https://benchling.com/s/seq-XuMlxoCIaSAHse8uxNbh?m=slm-UieHiRHS78CKGHaLm4cL).
I downloaded both files:
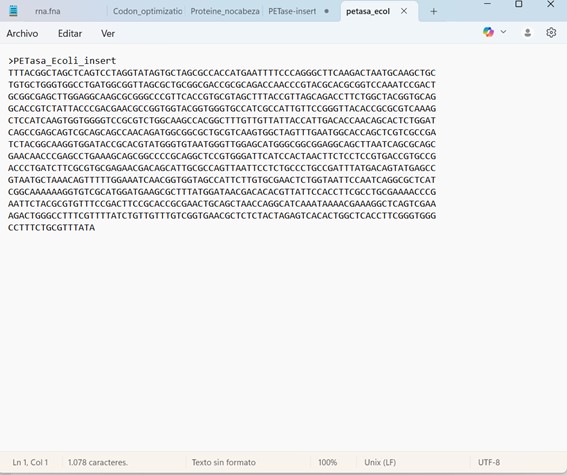
And I also exported and downloaded the FASTA file for the constructed sequence.
The final assembled sequence is known as an expression cassette, since it contains all the necessary elements required for gene expression, including the promoter, RBS, start codon, codon-optimized PETase coding sequence, stop codon, and terminator.
This cassette can be downloaded as a FASTA file for further use. In this format, the sequence can be shared, analyzed, synthesized, or inserted into a circular plasmid backbone for recombinant protein expression.
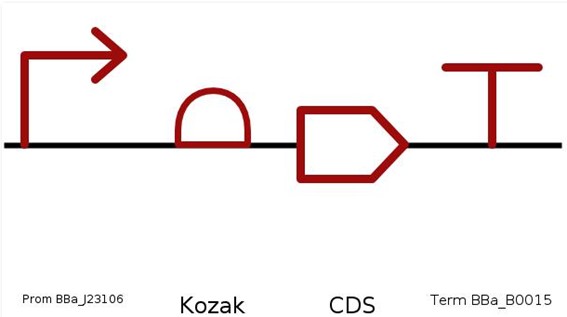
To visualize the genetic construct, I used SBOL Canvas to create a standardized graphical representation of the expression cassette. This tool allowed me to clearly illustrate the organization of the promoter, Kozak, codon-optimized PETase coding sequence, and terminator.
4.3. VectorBuilder: clonal genes
For this project, we selected the Clonal Genes option because it provides the DNA sequence cloned into a circular plasmid backbone, ready for direct transformation into E. coli.
Since the Twist Bioscience platform was not accessible, VectorBuilder was used as an alternative to insert the cassette into the plasmid backbone. VectorBuilder offers similar gene synthesis and cloning services, enabling the insertion of the designed expression cassette into a suitable plasmid backbone for bacterial expression. This alternative allows the construct to be obtained as a ready-to-use circular DNA molecule for transformation and recombinant protein production.
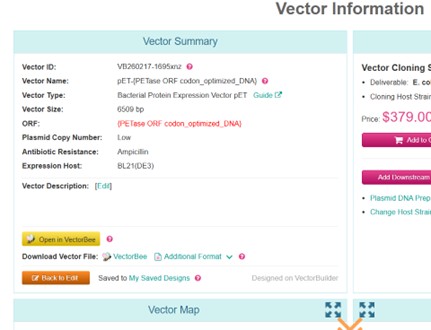
Open VectorBuilder and choose a vector system. I have chosen a bacterial recombinant protein expression vector (pET Guide) to express the codon-optimized PETase in E. coli. This vector is designed for high-level protein expression, supports all essential elements (RBS, start/stop codons, optional tags, terminator), and allows selection with antibiotics. It provides a reliable and compatible platform for producing functional PETase in a bacterial system.
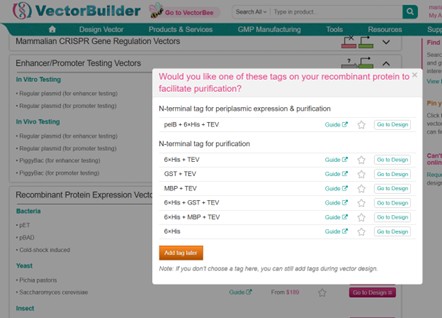

I did not add a purification tag to PETase at this stage. The protein can be expressed and remain functional without a tag, and a His-tag or other purification tag can be added later if needed for downstream applications.
The vector was constructed using the codon-optimized DNA of the PETase protein. A PDF file (VB260217-1695xnz(pET-{PETase ORF codon_optimized_DNA}).pdf) containing the vector information, a FASTA file (VB260217-1695xnz.fasta) with the sequence, and a GenBank file (VB260217-1695xnz.gb) were downloaded.
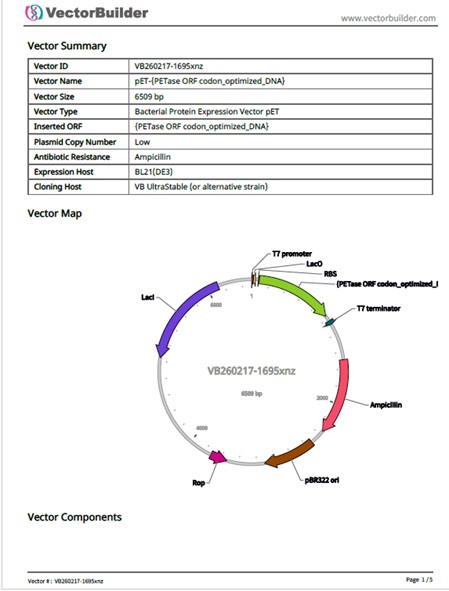

When selecting the vector, I was presented with other options that I was not certain were appropriate. It is important to carefully evaluate which vector is most suitable and to ensure that the promoters, RBS, and other components included in the vector are optimal for this project.
Part 5: DNA Read/Write/Edit
5.1 DNA Read
1. What DNA would you want to sequence (e.g., read) and why?
I would sequence DNA from soil or ocean microbial communities exposed to plastic, aiming to detect PETase or other plastic-degrading genes. This would help identify microorganisms capable of PET degradation and monitor their presence in different environments. I would also like to work with fossil and isolated organisms to better understand evolutionary adaptations.
2. What technology or technologies would you use to perform sequencing on your DNA and why?
I would use Illumina NGS (Sequencing by Synthesis) for high-throughput and accurate sequencing of environmental DNA. For fossil or isolated organisms, long-read technologies such as PacBio or Oxford Nanopore could help assemble fragmented genomes.
3. Is your method first-, second-, or third-generation? How so?
For detecting PETase genes in environmental samples, Illumina NGS (Sequencing by Synthesis) is a second-generation sequencing method. It produces millions of short reads in parallel, offering high throughput and accuracy, which makes it ideal for complex samples.
4. What is your input? How do you prepare your input? List the essential steps.
The input for Illumina NGS is extracted DNA from environmental samples. Essential steps include: DNA extraction, fragmentation, adapter ligation, PCR amplification, and library quality control before sequencing.
5. What are the essential steps of your chosen sequencing technology? How does it decode the bases of your DNA sample (base calling)?
Illumina Sequencing by Synthesis works by attaching DNA fragments to a flow cell and amplifying them into clusters. Fluorescently labeled nucleotides are incorporated one at a time, and after each incorporation, imaging captures the fluorescent signal. Software then converts these signals into base calls (A, C, G, T).
6. What is the output of your chosen sequencing technology?
The output of Illumina sequencing consists of FASTQ files containing DNA sequences and quality scores for each base. If multiple samples are sequenced together, the reads are demultiplexed using sample-specific barcodes.
5.2 DNA Write
1. What DNA would you want to synthesize and why?
I would like to synthesize genes related to environmental rescue, including biosensors, plastic-degrading enzymes such as PETase, and biomaterials for bioremediation. These DNA constructs could allow microorganisms to detect pollutants, respond to environmental stimuli, and break down plastics. For example, I would include the codon-optimized PETase ORF used in this project, along with regulatory elements such as promoters and ribosome binding sites to ensure proper expression.
2. What technology or technologies would you use to perform this DNA synthesis and why?
I would use commercial DNA synthesis platforms such as Twist Bioscience or enzymatic DNA synthesis methods. These technologies allow accurate and customizable synthesis of codon-optimized genes and regulatory elements. They are scalable, precise, and suitable for constructing complete expression cassettes.
3. What are the essential steps of your chosen DNA synthesis method?
DNA synthesis typically involves chemically or enzymatically synthesizing short oligonucleotides, assembling them into longer DNA fragments, verifying sequence accuracy, and cloning the construct into a plasmid backbone. The final product is then amplified and quality-checked before delivery.
4. What are the limitations of your DNA synthesis method in terms of speed, accuracy, and scalability?
DNA synthesis can be limited by cost, turnaround time, and potential synthesis errors in long or repetitive sequences. Very large constructs may require hierarchical assembly, increasing complexity and time.
5.3 DNA Edit
1. What DNA would you want to edit and why?
I would like to edit genes in microorganisms involved in plastic degradation to improve their efficiency, stability, or environmental tolerance. For example, I could modify the PETase gene to enhance its catalytic activity or thermal stability. Editing could also be applied to environmental bacteria to optimize metabolic pathways for bioremediation.
2. What technology or technologies would you use to perform these DNA edits and why?
I would use CRISPR-Cas systems because they provide precise, efficient, and versatile genome editing capabilities in microorganisms, plants, or animals.
3. How does your technology of choice edit DNA? What are the essential steps?
CRISPR-Cas uses a guide RNA designed to match a specific DNA target sequence. The Cas nuclease creates a double-strand break at the target site. The cell then repairs the break either through non-homologous end joining (causing insertions or deletions) or homology-directed repair (allowing precise edits if a repair template is provided).
4. What are the limitations of your editing method in terms of efficiency or precision?
CRISPR-Cas systems may have off-target effects, variable editing efficiency depending on the target sequence, and limitations related to delivery methods. Additionally, precise edits require efficient homology-directed repair, which may not occur at high frequency in all cell types.